A bacterial epoxide hydrolase might broaden the host range of bacteria through oxylipin signalling hijack
Published in Microbiology and Plant Science
All we had known about the possible functions of a protein called Cif (CFTR inhibitory factor) was studied by only a few research groups around the world. We had known that it is an epoxide hydrolase — an enzyme that cleaves various compounds having an epoxide moiety. We had known that it is present in two notorious nosocomial bacteria — Pseudomonas aeruginosa and Acinetobacter nosocomialis. And that it downregulates CFTR (cystic fibrosis transmembrane conductance regulator) through the ubiquitination system — thus phenocopying cystic fibrosis.
I don’t remember why I started constructing a phylogenetic tree for it some years ago — I believe this was just an exploratory study. Using the Cif of Pseudomonas aeruginosa as a BLAST search query, I obtained the list of putative homologues and constructed a tree. Cif’s of Pseudomonas aeruginosa and Acinetobacter nosocomialis appeared to belong to the same cluster — indicating that all this cluster is represented by functional Cif’s, bacterial enzymes with the same function (Fig. 1).
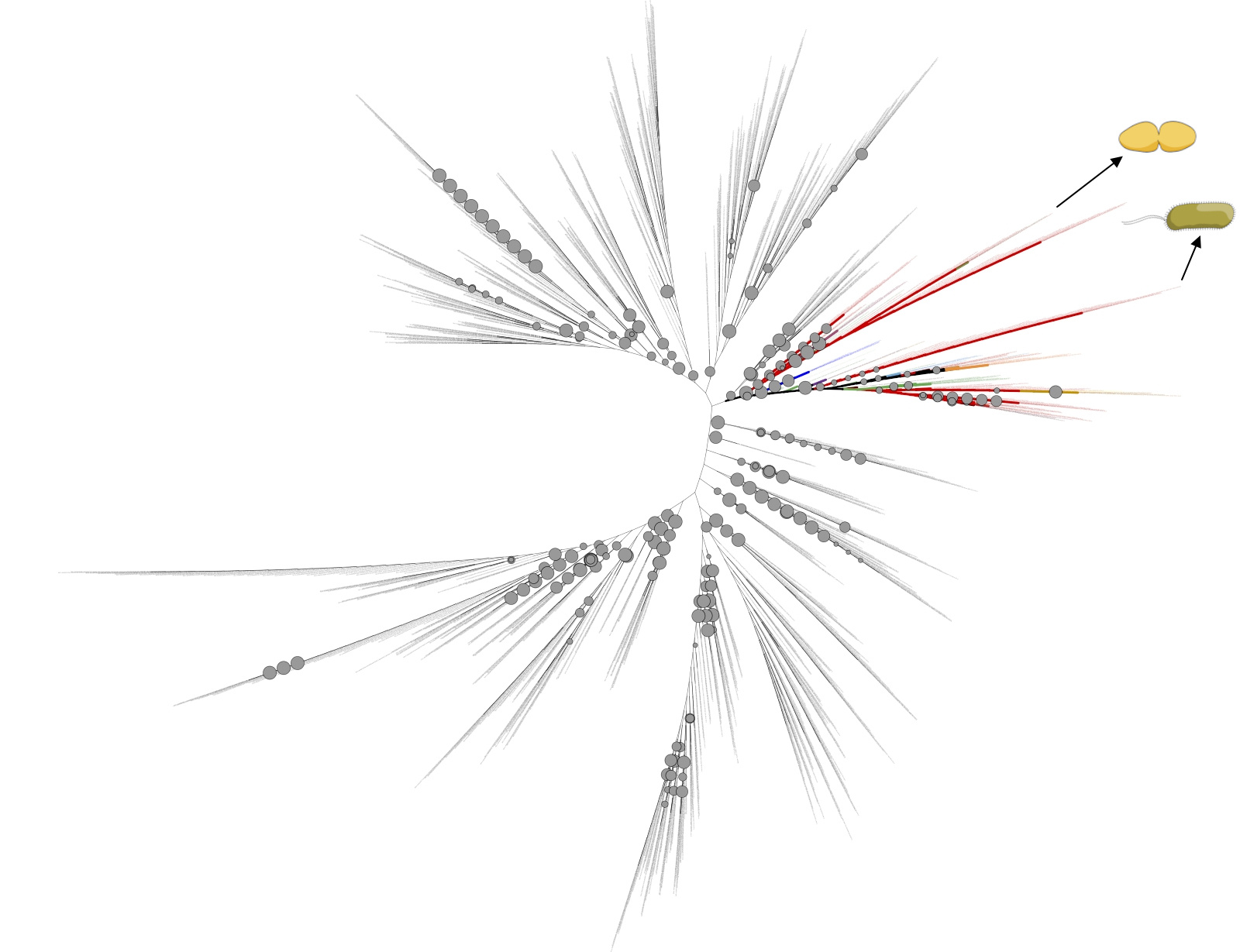

In this cluster, there were Cif’s of several tens of bacteria whose ecological niche had been already described in the literature. And their ecological niches were not restricted to humans and nosocomial environments — instead, this was the patchwork of nosocomial pathogens, plant growth-promoting rhizobacteria, plant pathogens, insect-associated bacteria, colonizers of mollusks and shrimp… and a plenty of bacteria in which these functions were overlapped. This was highly reminiscent of another case — the case of lipoxygenase.
I covered this enzyme in detail in my previous posts on this blog and the respective papers. Besides multicellularity, lipoxygenase also appears to be linked to host-microbe interactions. And the ecophysiological profile of its carriers is strikingly similar: extremely broad and varying host ranges, overlap and coincidence of pathogenic and symbiotic functions, specific ecological associations (such as plant rhizosphere and lung infections in humans). At that time, my main question was “Why?” Why do these enzymes look so similar in terms of the ecological distribution of their carriers, if they are completely different activity and molecular targets?
Insight came much later, when colleagues showed me a recent experimental article on Cif [1] while writing a collaborative review. In this article, the same team that once described Cif, characterized its ability to cleave fatty acid epoxides and this influence the immune signalling. This was the moment of “Eureka!”: oxylipin signalling is the same system that is presumably targeted by bacterial lipoxygenases. This could be the common targeted pathway that might be an explanation of the similarity between the ecological profiles of the lipoxygenase carriers and Cif carriers. But I needed to prove the similarity first.
In the previous preprint, I used the complicated methodology of network text analysis while I extracted entities from PubMed texts on the lipoxygenase-carrying bacteria manually and tried to identify the overall pattern. This method raised some questions in the editorial offices where I managed to obtain the reviews, and I chose the simpler methodology for a new paper. The resulting paper is now available as a preprint on Preprints.org server [2].
For this paper, I made it simpler. I took the list of carriers of putative Cif’s (from the phylogenetic tree cluster mentioned above) and the list of carriers of putative lipoxygenases (from the previous preprint). Then I searched articles for each species for which ecological data were available and compared the “lipoxygenase list” and the “Cif list” statistically by the list of ecological traits.
These traits were selected by their overall occurrence in the literature on these species, but the traits list was collected to cover such crucial ecological traits as the host’s taxonomic group and habitat, effect on the host, localization within the host’s organism etc. Thus, I obtained the list of 36 traits for comparison such as “insect-pathogen”, “plant-endophytic”, or “hydrocarbon-deg”.
Two-proportion Z-test showed the significant difference only by 6 of these 36 traits, if we used p-value threshold = 0.05. However, I dealt with multiple comparisons by 36 traits, and this result could not be used for any conclusion without Bonferroni correction. With Bonferroni correction, we should have p-value threshold = 0.05/36 = 0.001388(8)... This barrier has not been crossed by any or the traits used.
Here, we might conclude that the "lipoxygenase list" and "Cif list" of bacteria are completely similar by their ecological profile, and both LOX and Cif have the similar ecological effect... if Z-test were good enough in confirming null hypotheses! But this is not the case.
To check this, I used the correlation analysis where the occurrence of a trait in LOX carriers was x, its occurrence in Cif carriers was y, and each trait was treated like a dot on the correlation field. Thus, we avoided the problems of multiple comparison, because all traits were treated as an entire set compared by two values (you can say that I inverted the problem a little bit).
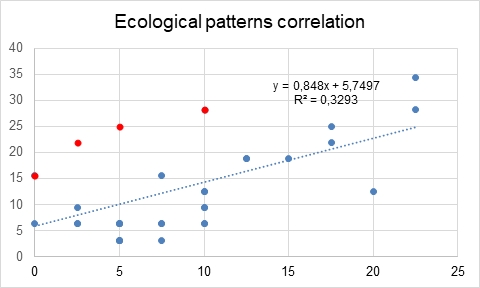
The correlation appeared to be relatively weak, with R-square of only 0.33. However, if you look at the correlation plot (Fig. 2), you can see the set of 5 dots forming an unusual “upward arc” on the plot and thus spoil the whole picture. I have marked them in red — and they represent the 6 traits with notable differences I mentioned above (one of the dots represents 2 traits due to the same occurrences). I decided to temporarily remove these 6 traits from the analysis and calculate the correlation without them — and then look at these 6 traits separately.
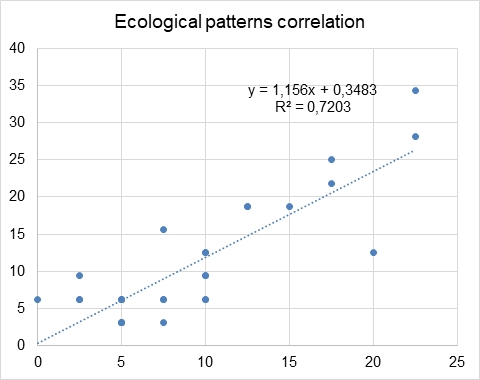
Without these traits, correlation improved and became stronger, with the R-square of 0.72 (Fig. 3). Moreover, the regression equation changed to y = 1.156x + 0.3483. On the scale of our plot, it is almost the same as y = x, and the coefficients could be legitimately rounded to 1 and 0, respectively. It means that the occurrence of 30 traits (of total 36) is almost the same from the statistical point of view. They do not differ significantly by these 30 traits and are a kind of the same statistical universe.
These traits form a collective “ecological portrait” of oxylipin spoofers which resembles the “ecological portrait” of lipoxygenase-carrying bacteria I outlined in the previous blog post on my preprint (Fig. 4). Moreover, even the remaining 6 traits which are statistically different between lipoxygenase carriers and Cif carriers indirectly confirm my hypothesis on bacterial lipoxygenases in cross-kingdom host jumps.
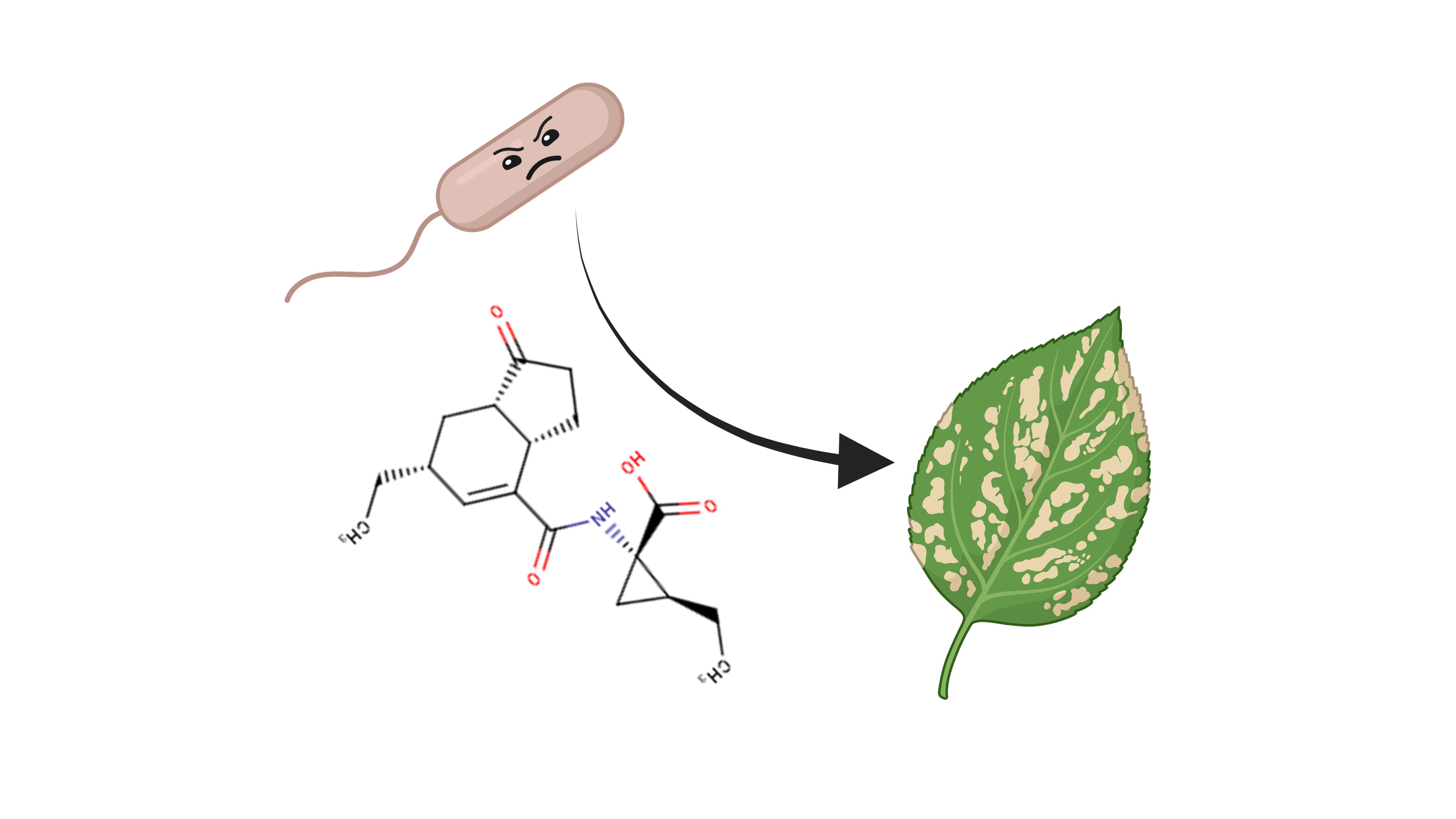
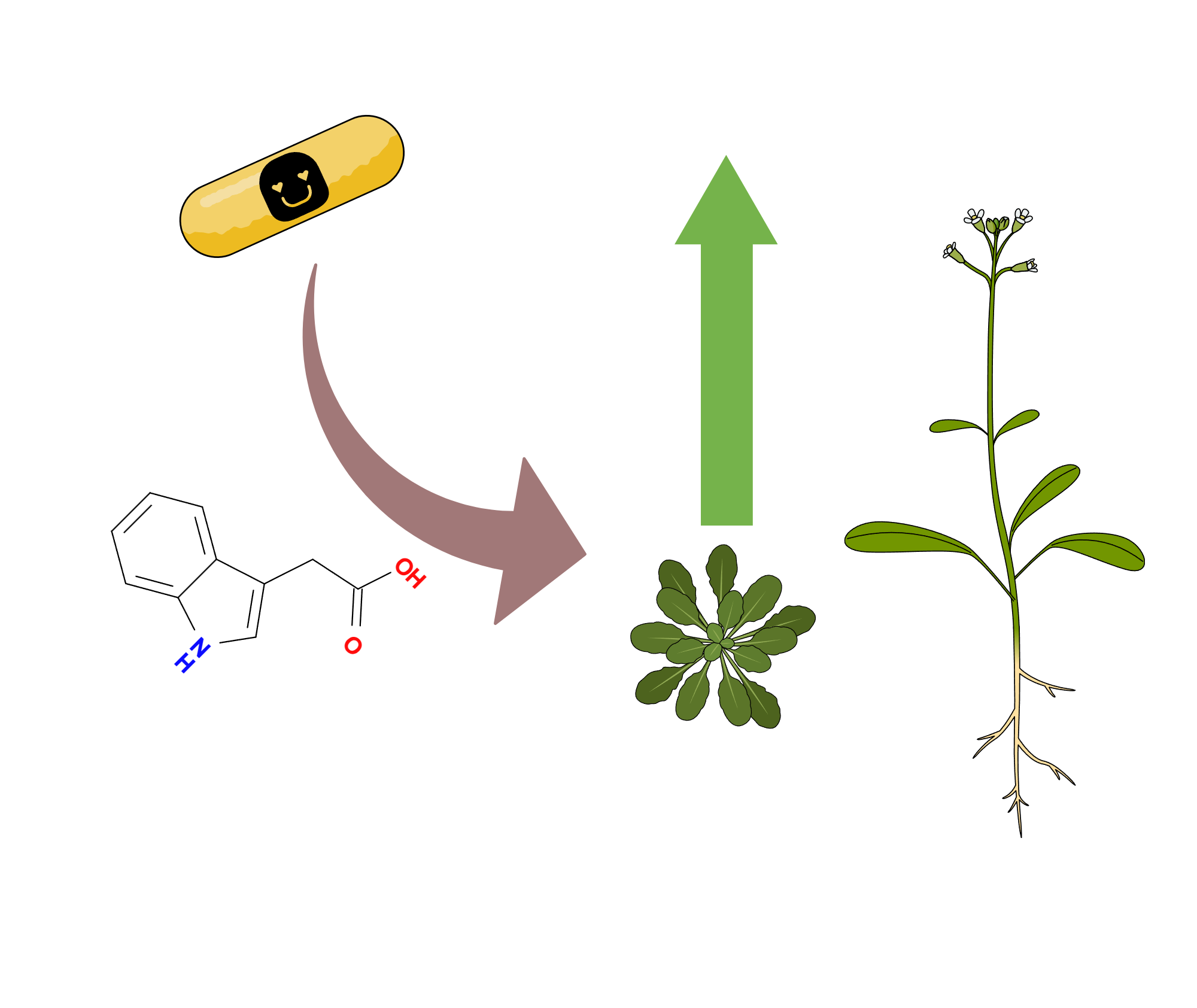
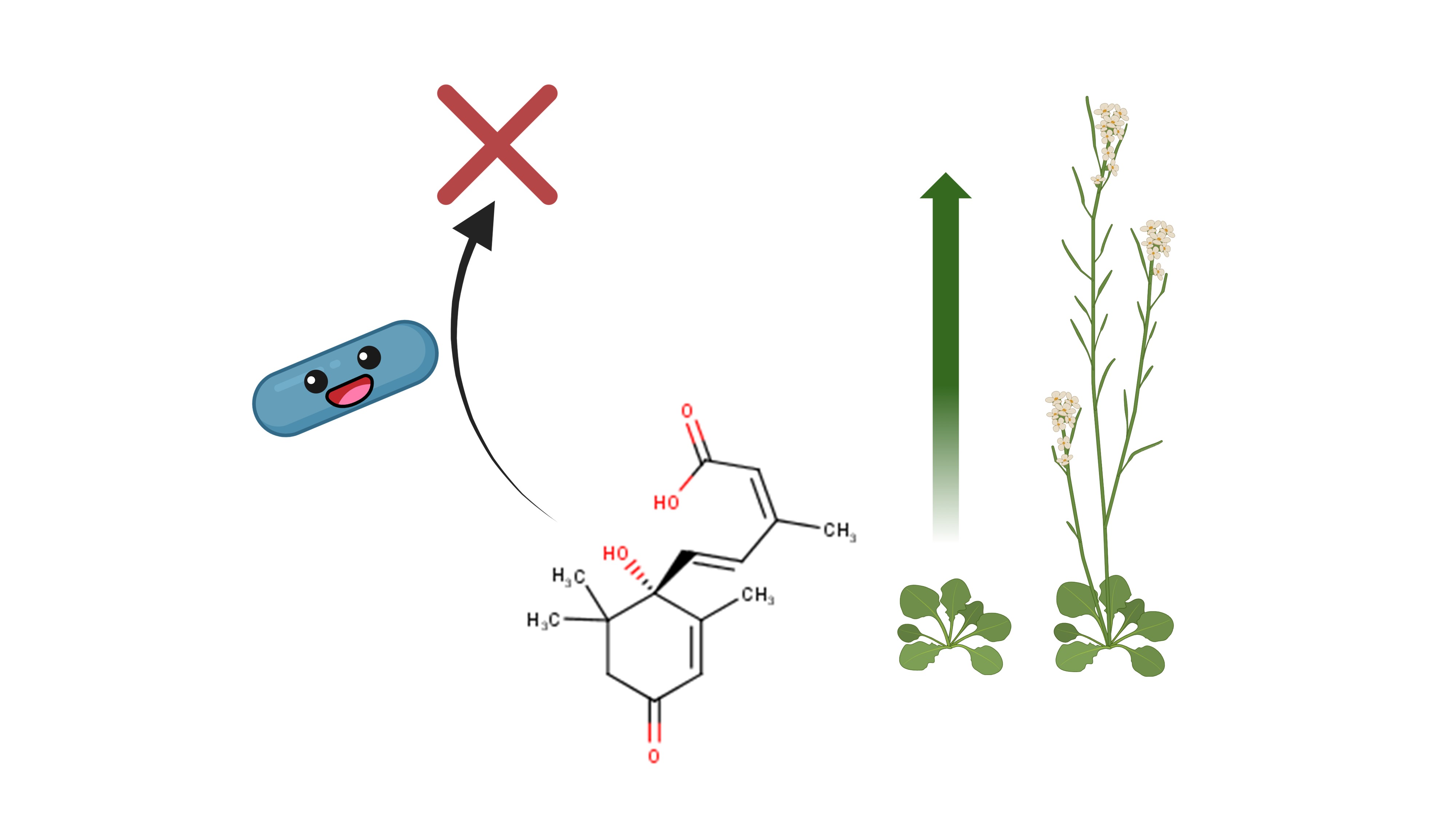
These 6 traits include the ability to cause bacteremia/sepsis, plant growth promotion, plant hormone production, xenobiotic degradation, hydrocarbon degradation, PAH degradation (Fig. 5). By all of these points Cif-carrying bacteria are more successful than the LOX-carrying bacteria — they occur more often in the “Cif list”. The latest 3 traits are related to bioremediation and are not directly linked to host-microbe relationships. Regarding the plant hormone production and plant hormone production, it seems that there is a causal relationship between them. The most notorious hormone producer in my “Cif list” was Pseudomonas syringae — a plant pathogen which produces a jasmonate mimic coronatine to suppress the plant’s immune response (Fig. 6). But most plant hormone manipulators exert a growth-promoting effect — for example, producing the plant growth hormone auxin (Fig. 7) or degrading abscisic acid, a plant quiescence hormone which suppresses growth (Fig. 8).
.png)
Thus, Cif-carrying bacteria seem to be better plant hormone manipulators (by our statistics!) and are therefore better plant growth promoters. But why? The answer lies within the hypothesis I have put forward in the previous preprint.
In the previous study, I proposed the mechanism which might mediate plant-human host jumps in LOX-carrying bacteria (Fig. 9). According to this hypothesis, they carry (ω-6)-lipoxygenase which produces (ω-6)-fatty acid hydroperoxides from the host’s lipids. Then, the host’s enzymes convert these intermediate products to potent oxylipins: they are lipoxin A4 in humans and jasmonate in plants. Both of them can suppress the host’s immune response when they are overproduced and the equilibrium of defense response mediators is outbalanced. But this also implies that LOX-carrying bacteria are indirect jasmonate producers by design! They just are not described in this regard because they do not produce jasmonates themselves in vitro — and thus their hormone-hijacking activity is not included in my statistics.
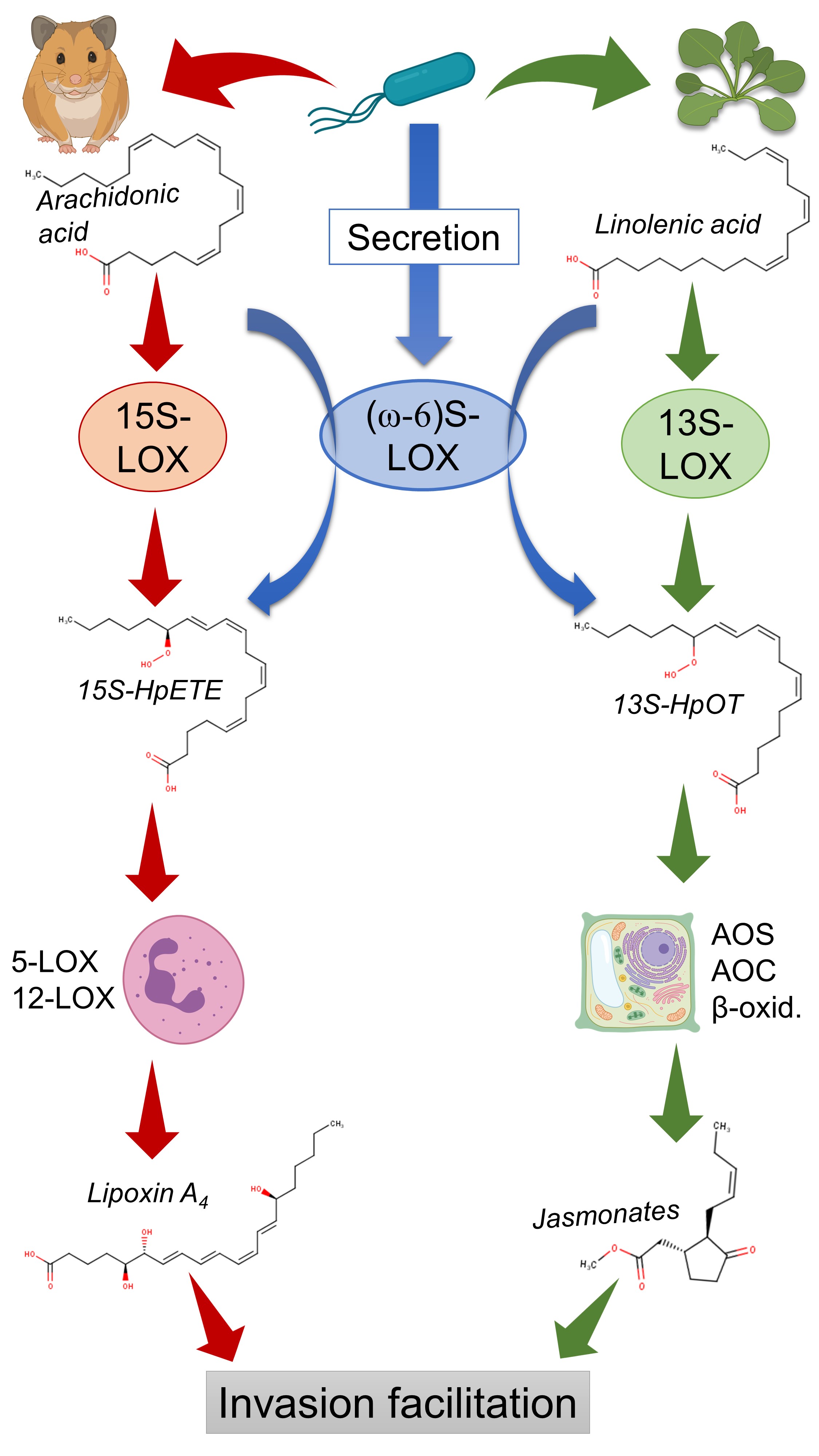
This explains why Pseudomonas syringae produces coronatine: it has Cif to manipulate plant oxylipins but does not have lipoxygenase to trigger jasmonate overproduction — thus it uses an alternative jasmonate mimic. All other hormone-hijackers from the “Cif list” are the same case: they need additional self-sustained hormone-producing activity, because Cif does not yield hormones like lipoxygenase.
This is an additional piece of puzzle in terms of plant-microbe interactions. The higher activity towards causing septic infections and degrading xenobiotics are still missing pieces of the puzzle and new challenges for the research project. But, unfortunately, they are not the only challenges of this paper.
This paper was written for the conference proceedings issue but was rejected by peer review after two revisions. After this, I tried to post it on bioRxiv where it was also rejected upon screening. The decision letter read:
We regret to inform you that your manuscript is inappropriate for bioRxiv. All submissions are screened to ensure they match bioRxiv's posting criteria and scope. Simple molecular modeling, sequence alignments and results of facile database searches are generally not sufficient and therefore, your submission was considered out of scope.
Please be assured that this conclusion simply refers to the manuscript’s appropriateness for bioRxiv and is not a judgment on the merits of the work described (check out our FAQs for more information).
This statement shows that the paper has a significant issue: using only open data without mathematically complicated methods makes the research seem relatively “simple” — and not a research in the strict sense. Its formal publication might be difficult due to the expectations of larger scale and complexity of the work. Despite this, I consider this paper to be important and worth publishing — because it addresses some questions to the previous controversial preprint and might lead to new understanding of nosocomial pathogenicity mechanisms.
I will keep submitting it to different journals — and if it eventually gets published, I will add an update. If it meets the same fate as the previous preprint, it will remain an “open science” project forever and will be still available on Preprints.org and other open science platforms. On this blog, for example.
References
- Hvorecny K.L., Bahl C.D., Kitamura S., Lee K.S.S., Hammock B.D., Morisseau C., Madden D.R. (2017) Active-Site Flexibility and Substrate Specificity in a Bacterial Virulence Factor: Crystallographic Snapshots of an Epoxide Hydrolase. Structure, 25(5):697-707.e4. https://doi.org/10.1016/j.str.2017.03.002
- Kurakin, G. (2026) Lipoxygenases and CFTR Inhibitory Factors Might Share the Same Role in Host-Microbe Interactions. Preprints, 202602.1829.v.1. https://doi.org/10.20944/preprints202602.1829.v1
Other references are presented as direct hypertext links to avoid the post overload and altimetric inflation.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in