Deep learning for the detection of benign and malignant pulmonary nodules in non-screening chest CT scans
Published in Cancer

The challenge of lung nodule detection in routine clinical practice
Lung cancer is worldwide the leading cause of cancer death1. It is often diagnosed in a late stage as symptoms usually do not appear until the cancer has progressed into a higher stage2. For this reason, nationwide lung cancer screening programs have been initiated in Korea, China, and USA that aim to detect lung cancer as early as possible. In a screening program, high-risk patients (older smokers) are invited for an examination of their lungs with a low-dose CT scan. Lung cancer may appear as a small spot or “nodule” in the lungs. While trials have provided evidence that low-dose CT screening reduces lung cancer mortality3,4, its adoption into clinical practice has been slow in Europe and other countries.
As a result, early-stage lung cancer is generally diagnosed after incidentally detecting nodules in CT scans that were ordered for other medical reasons5,6. Detecting lung nodules is challenging in this setting. Unlike lung cancer screening, the focus of a CT examination is not necessarily on lung cancer detection and the scans are highly heterogenous due to more diverse scanning protocols and patients. Efficient nodule detection and management will become increasingly important in the near future: the workload of radiologists has drastically increased in the last 15 years, mainly due to the increased demand for CT imaging7.
Artificial Intelligence for radiological support
Software using AI technology can potentially support radiologists in detecting lung nodules in CT scans. Previous studies have shown that when AI software is used as an extra reader, radiologists obtain a higher detection sensitivity, improve the uniformity of their management recommendations, and reduce their reading time8-11. However, only a small number of studies have investigated the performance of AI solutions in a non-screening setting. In our study, we wanted to extend this line of research by addressing three common limitations: first, we evaluated the generalization performance of the AI software by opting for a multi-center study design instead of a single center design. Second, we invited a panel of five expert thoracic radiologists for determining the reference standard instead of one or two radiologists, because large interobserver variability exists in nodule detection. Third and most importantly, we investigated whether an AI system can find the nodules that really matter by using reliable, nodule-level malignancy labels. AI studies have either investigated the detection performance for all nodules (regardless of malignancy status) or cancers on scan-level. Therefore, this study aims to bridge the gap between lung cancer and nodule detection AI studies.
Bridging the gap between lung cancer and nodule detection AI studies
We developed a deep learning-based system and retrospectively assessed its performance for the detection of actionable benign nodules (requiring follow-up), small lung cancers, and metastases in CT scans acquired from a routine clinical setting in two Dutch hospitals (internal and external validation). Our nodule detection pipeline has three components: first, the system takes a CT scan and detects the lungs slice-by-slice to obtain a region of interest. Second, nodule candidates are generated by analyzing overlapping CT volumes of five slices each. Finally, nine different slices from a 3D patch around each nodule candidate are analyzed to determine whether a nodule is present. By employing this 2.5D detection approach, the system can quickly analyze the CT scan while preserving robustness to reliably distinguish nodules from lung vessels and other structures.
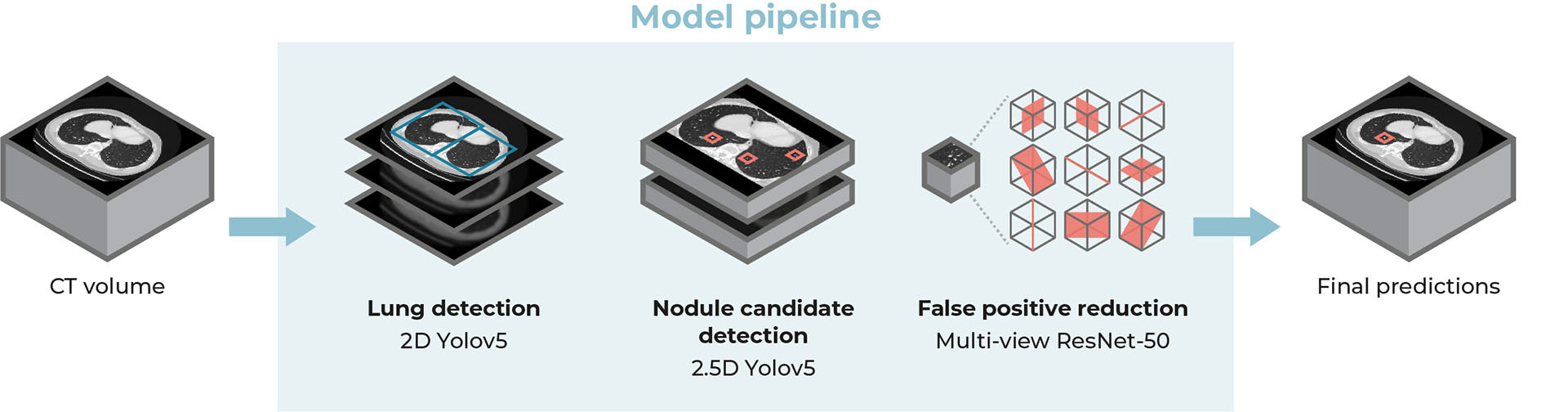
Overview of the proposed lung nodule detection system.
To evaluate the performance of the AI system, we asked a panel of five thoracic radiologists to independently label all nodules in the test datasets. Two additional radiologists verified the malignancy status of each nodule (with histopathological proof or follow-up imaging) and searched for any missed cancers by using data from the national Netherlands Cancer Registry. The AI detection performance was evaluated by measuring the sensitivity at predefined false positive rates on a free receiver operating characteristic (FROC) curve and was then compared with the panel of radiologists.
Performance of the AI system
On the external test set (100 scans from 100 patients), the sensitivity of the AI system for detecting benign nodules, primary lung cancers, and metastases was respectively 94.3%, 96.9%, and 92.0% at a clinically acceptable threshold of 1 false positive per scan (FP/s) on average. These sensitivities were found to be comparable to or higher than the radiologists (average difference of 14, 2, and 17 percentage points for the aforementioned categories) at the cost of a slightly higher FP/s (average difference of 0.6). We found that the AI system could locate most lung metastases (78%) that were missed by the majority of radiologists. These were usually located in known blind spots (e.g., base of the lower lobes and the right sided azygo-esophageal recess12,13) and our results suggest that these could be overcome with the help of AI.
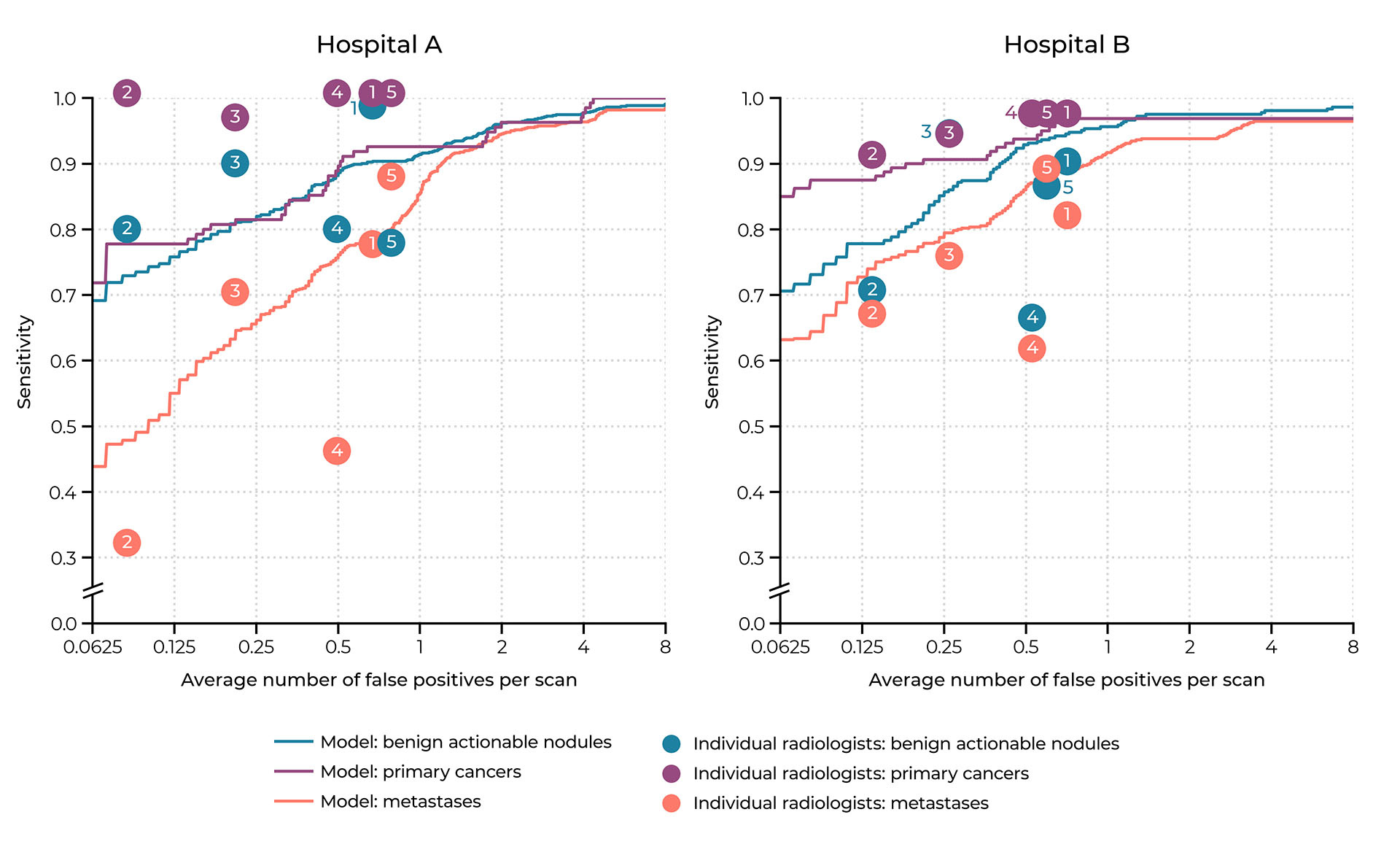
Operating points of the individual radiologists of the panel compared to the free response receiver operating characteristic (FROC) curves of the AI system on the internal test set (hospital A) and external test set (hospital B).
Take-home message
Our findings demonstrate that AI can accurately detect benign and malignant lung nodules in heterogenous CT scans from a non-screening setting. A comparison with thoracic radiologists shows that it has potential to aid radiologists in finding all clinically relevant nodules in routine clinical practice. We hope to facilitate future lung cancer research by providing access to our algorithm on the platform Grand Challenge.
Algorithm availability
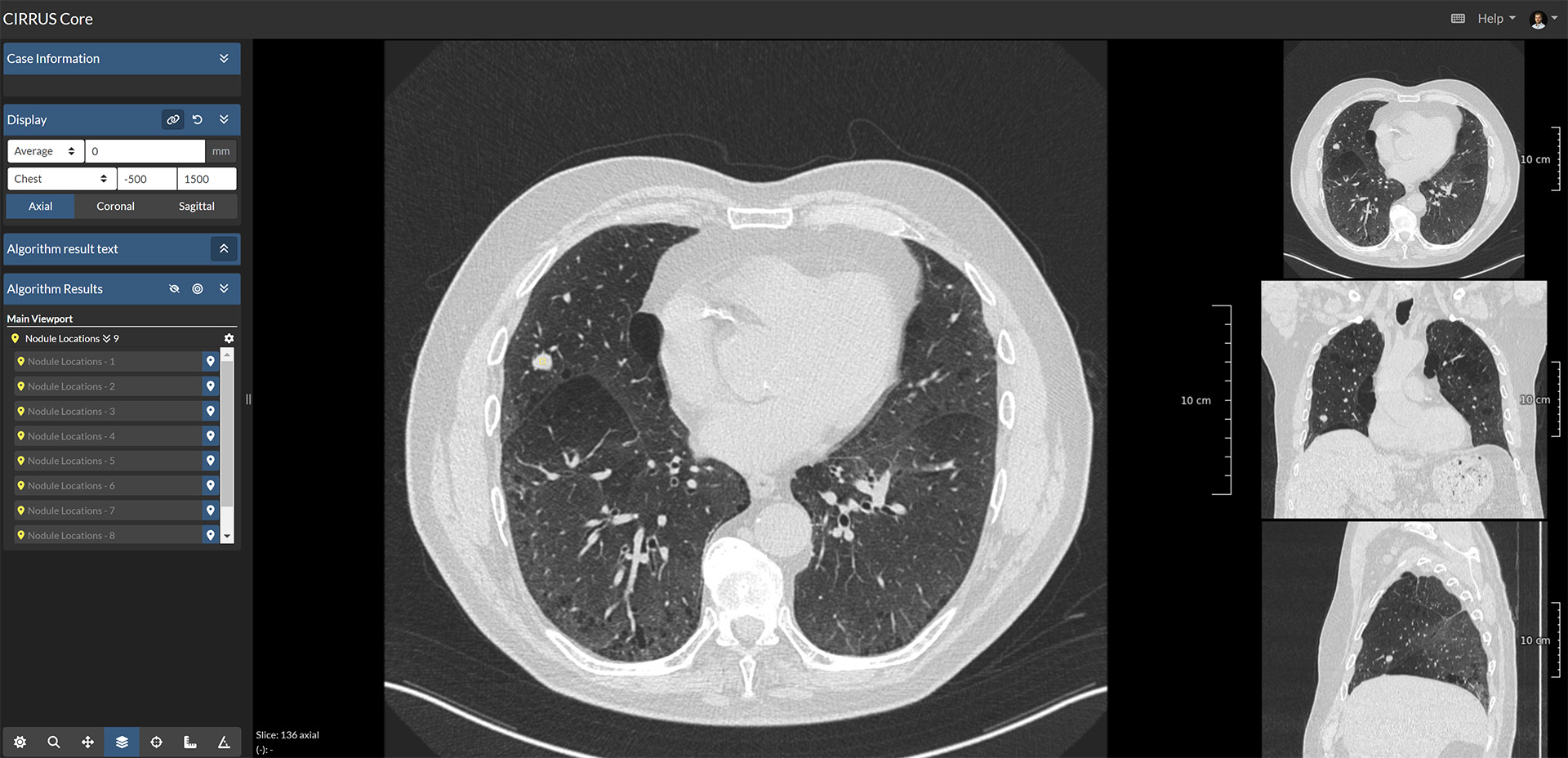
We made our AI system publicly available for research purposes on the web platform Grand Challenge. On this platform, the algorithm can be readily executed in the cloud and does not require the installation of any software. The algorithm takes a thorax or thorax-abdomen CT(A) scan as input, which can be provided as either a DICOM or MHA file. Then, it returns the location and corresponding likelihood score of any lung nodule. The nodules are highlighted as points in the scan, which can be inspected in a professional radiology workstation. Researchers can also download the complete algorithm outputs via the GC-API when many scans need to be analyzed. More detailed background and usage information can be found on the algorithm information page.
References
- Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA. Cancer J. Clin. 71, 209–249 (2021).
- Birring, S. S. & Peake, M. D. Symptoms and the early diagnosis of lung cancer. Thorax 60, 268–269 (2005).
- The National Lung Screening Trial Research Team. Reduced Lung-Cancer Mortality with Low-Dose Computed Tomographic Screening. N. Engl. J. Med. 365, 395–409 (2011).
- de Koning, H. J. et al. Reduced Lung-Cancer Mortality with Volume CT Screening in a Randomized Trial. N. Engl. J. Med. 382, 503–513 (2020).
- Gould, M. K. et al. Recent Trends in the Identification of Incidental Pulmonary Nodules. Am. J. Respir. Crit. Care Med. 192, 1208–1214 (2015).
- Hendrix, W. et al. Trends in the incidence of pulmonary nodules in chest computed tomography: 10-year results from two Dutch hospitals. Eur. Radiol. (2023) doi:10.1007/s00330-023-09826-3.
- Bruls, R. J. M. & Kwee, R. M. Workload for radiologists during on-call hours: dramatic increase in the past 15 years. Insights Imaging 11, 1–7 (2020).
- Murchison, J. T. et al. Validation of a deep learning computer aided system for CT based lung nodule detection, classification, and growth rate estimation in a routine clinical population. PLoS One 17, e0266799 (2022).
- Jacobs, C. et al. Assisted versus Manual Interpretation of Low-Dose CT Scans for Lung Cancer Screening: Impact on Lung-RADS Agreement. Radiol. Imaging Cancer 3, e200160 (2021).
- Hempel, H. L., Engbersen, M. P., Wakkie, J., Kelckhoven, B. J. van & Monyé, W. de. Higher agreement between readers with deep learning CAD software for reporting pulmonary nodules on CT. Eur. J. Radiol. Open 9, (2022).
- Kozuka, T. et al. Efficiency of a computer-aided diagnosis (CAD) system with deep learning in detection of pulmonary nodules on 1-mm-thick images of computed tomography. Jpn. J. Radiol. 38, 1052–1061 (2020).
- Miki, S. et al. Prospective Study of Spatial Distribution of Missed Lung Nodules by Readers in CT Lung Screening Using Computer-assisted Detection. Acad. Radiol. 28, 647–654 (2021).
- Chen, H. et al. A retrospective study analyzing missed diagnosis of lung metastases at their early stages on computed tomography. J. Thorac. Dis. 11, 3360–3368 (2019).
Follow the Topic
-
Communications Medicine
A selective open access journal from Nature Portfolio publishing high-quality research, reviews and commentary across all clinical, translational, and public health research fields.
Related Collections
With Collections, you can get published faster and increase your visibility.
Healthy Aging
Publishing Model: Open Access
Deadline: Jun 01, 2026
Public health and health governance in China
Publishing Model: Open Access
Deadline: Jul 31, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in