How to select the best candidate or the key factors? Hierarchical topological clustering can help
Published in Research Data
Often, we find ourselves needing to choose the best candidates or key factors from a long list of possibilities or datasets. For example, we may wish to select genes that appear distinguished from the rest based on their expression in cancerous tissue, identify the countries with which trade relations are dominant on an international scale, or select the best candidate for a job. At another level, we may be interested in characterizing multicluster dynamics in datasets with a hierarchical structure. For example, we may be wish to study the fragmentation of population groups according to different criteria and their evolution according to circumstances. With the development of machine learning techniques, it is tempting to resort to clustering methods to reach conclusions. Standard clustering techniques focus on finding a definite group structure and tend to eliminate outliers, either by considering them as noise or by forcing their membership in groups. We present a hierarchical topological clustering algorithm (HTC) that automatically identifies the presence of outliers and multicluster structures at different scales, being robust to noise.
Topological methods have the potential of exploring the shape of datasets across different scales by analyzing topological features (connected components, holes) that persist at different spatial resolutions. Once the distance measuring the similarity of the elements in the dataset is chosen, we construct a nested one-parameter set of simplicial complexes through a Vietoris-Rips filtration. We then use this simplicial complex to define a clustering hierarchy spanning from small to large scales. For each value of the filtration parameter, the resulting disjoint components of the simplicial complex define clusters of an arbitrary shape. These clusters merge into larger clusters as we vary the filtration parameter and evolve to a different simplex of the family. We progress from small scale clusters to cluster structures at larger scales, keeping track of their relationships and the individual elements within each cluster throughout the process. Cluster configurations that persist for broad filtration intervals (across scales or within scales of interest) are considered dominant. We introduce the concept of persistence gap to identify these configurations and their posible multicluster structure. Persistent outliers are candidates to represent meanigful rare elements, that is, elements with a particular significance for the dataset under study (such as key genes or trade partners), rather than noise. This procedure can be applied with any distance, though some distances are known to be more adequate to compare elements in specific applications. For instance, Wasserstein distances tend to be more adequate than Euclidean distances to compare images. Moreover, the stability of the clusters under noise is quantifiable through Bottleneck distances defined on persistence diagrams, a property not enjoyed by standard hierarchical clustering (HC).
We demonstrate the potential of our algorithm with different distances (Euclidean distance, Wasserstein distance, Fermat distance) on datasets of moderate size extracted from real life applications in which outliers and/or multiclustering are relevant:
- Experimental images of mutated cells invading healthy populations: The interface between the two populations typically consists of a main barrier and the contours of a multitude of islands of one population within the other, located at varying distances from the main interface. While HTC captures the resulting geometric island structure, standard clustering algorithms such as HC, Kmeans, and DBSCAN fail.
- Swarm dynamic structure: In the studies of swarm dynamics, recording the positions of each member of the swarm over time we obtain a time series of datasets displaying time dependent multicluster structure. HTC identifies automatically the dominant clusters and the main subclusters within them, as well as their time evolution, by analyzing the persistence gaps.
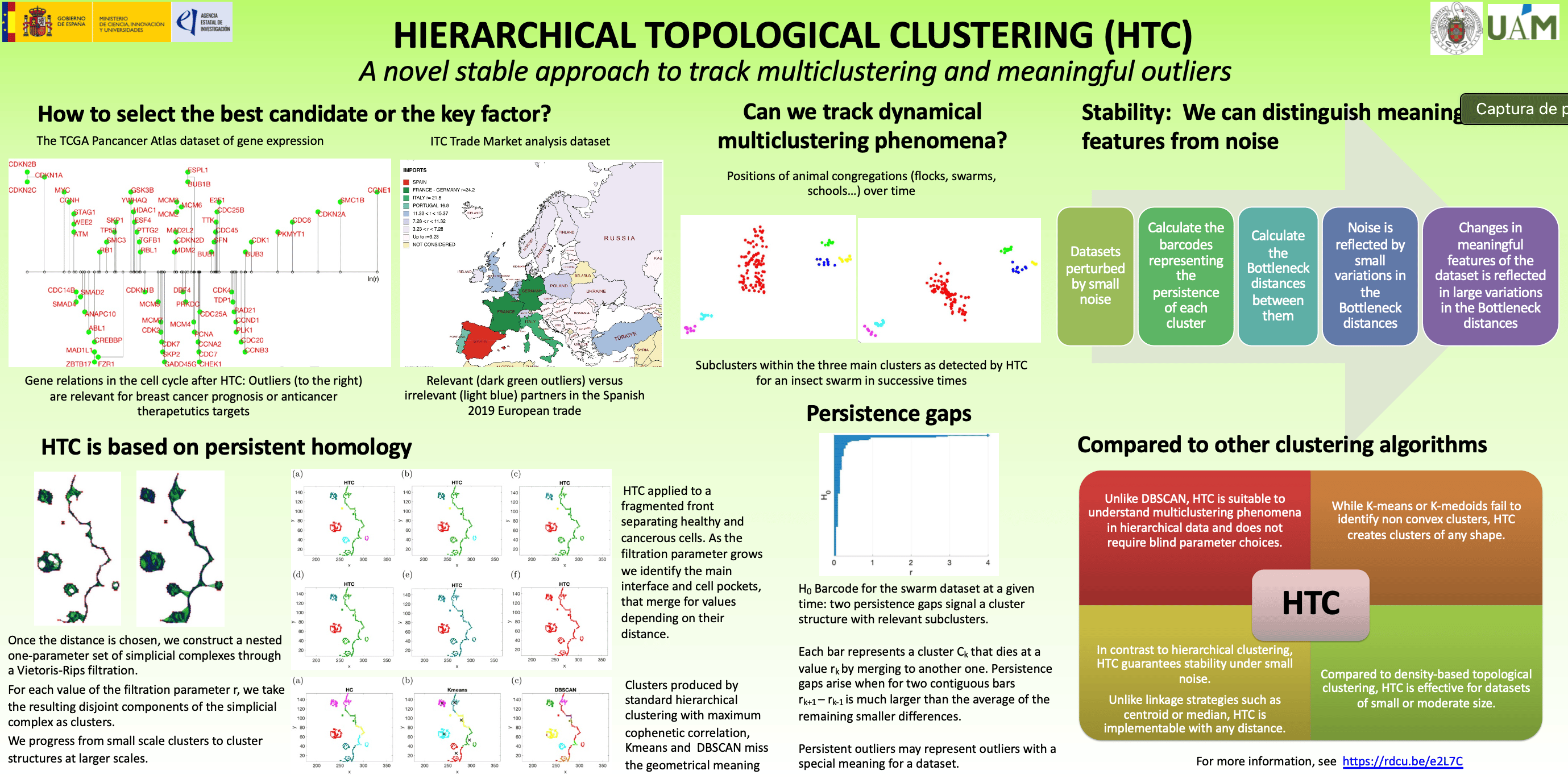
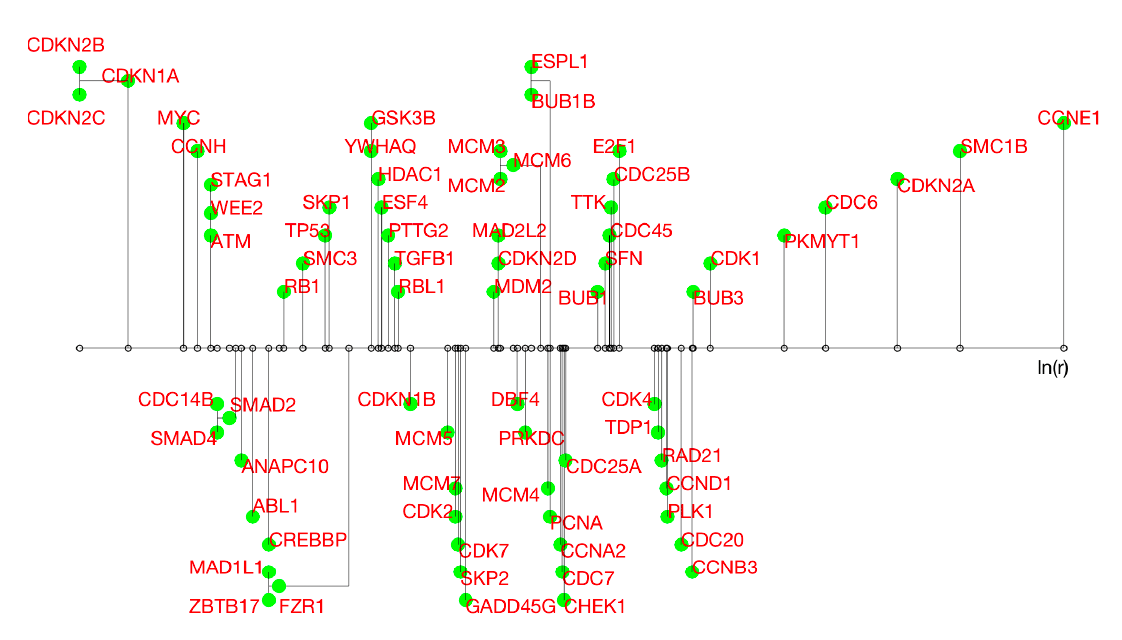
- Gene expression data for cancer studies: Programs such as the TCGA Pan-Cancer Atlas measure the expression of human genes in cancer tissue and healthy tissue in order to identify genes that play an important role in different types of cancer. These datasets are then conveniently renormalized and analyzed trying to identify relevant cycles. Considering only genes involved in the cell cycle from the breast cancer dataset, HTC identifies a structure with distinguished outliers, all of which have been proven to be relevant for breast cancer prognosis or singled out as anti-cancer therapeutic targets. The attached image represents how the genes are grouped. The genes isolated on the far right are the dominant outliers.
- Import-export trade volume between European countries: Trade datasets for international business development collect the collect the amount of purchases or sales of all types of goods from any country to the others. HTC identifies a structure with distinguished outliers: the relevant trade partners for each country.
- Image quality: Images can be corrupted by excessive compression and/or by the presence of streaks or strange dots. Given a dataset made up of the same image with different degrees of compression and occasional presence of streaks, HTC is able to distinguish the two types of disturbances when classifying.
In these tests, we compare the performance of the algorithm against a known ground truth. For more sistematic performance analyses, we resort to the standard accuracy, precission, recall and F1-score metrics. Tests are simple enough to be run in a laptop with MATLAB.
Unlike popular clustering methods like DBSCAN, our strategy is suitable to understand multiclustering phenomena in hierarchical data and does not require blind parameter choices. While popular methods such as K-means or K-medoids fail to identify non convex clusters, the proposed algorithm creates clusters of any shape. In contrast to hierarchical clustering, we can guarantee stability under small noise. On the other hand, unlike linkage strategies such as centroid or median, this algorithm can be implemented with any distance choice, not just Euclidean distances. The complexity of the algorithm is similar, bounded from above by MN2, N being the amount of data and M the resolution levels. As the size of the dataset grows, other density-based topological clustering techniques may be more effective.
Published in https://doi.org/10.1007/s00500-025-11028-6 https://rdcu.be/e2L7C
Follow the Topic
-
Soft Computing
This journal is a hub for system solutions based on unique soft computing techniques.
Related Collections
With Collections, you can get published faster and increase your visibility.
Advances in Large-Scale and Intelligent Optimization: Algorithms, Models, and Applications
Extended Papers from the International Workshop on Big Optimization 2025 WBO25 that was held by the University of Catania, Italy, 31 July – 1 August 2025
This collection aims to gather cutting-edge research on large-scale and intelligent optimization, covering both theoretical developments and practical applications. Contributions are welcome on novel algorithms, hybrid and memory-enhanced metaheuristics, optimization under uncertainty, and AI-integrated approaches.
This collection is open to all contributors, not limited to invited authors.
We especially encourage submissions addressing complex, real-world problems in diverse domains such as transportation, telecommunications and energy systems. What makes this collection unique is its focus on scalability, robustness, and adaptability in modern optimization, with special attention to interdisciplinary and impactful applications.
Publishing Model: Hybrid
Deadline: Jul 31, 2026
Intelligent Systems & Decision Making: Human Insights in the Era of AI
Extended Papers from the International Summer Conference ISC2025 that was held by the University of Catania, Italy, June 19 – 20, 2025
This collection is dedicated to the best accepted papers on the International Summer Conference – IDS 2025 (https://decisionsciencealliance.org/isc-2025/), properly revised and considerably extended with respect the conference version. In addition, to guarantee as high as possible quality of manuscripts, the collection is also opened to all researchers interested in the topics even if not attending the conference.
We especially encourage submissions that explore the trends and latest advancements in decision science and technology and understand how these innovations can influence industries and practices.
Publishing Model: Hybrid
Deadline: Sep 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in