Next-generation neoantigen mRNA vaccines: Immuno-engineering strategies for precision cancer immunotherapy
Published in Bioengineering & Biotechnology, Cancer, and Immunology
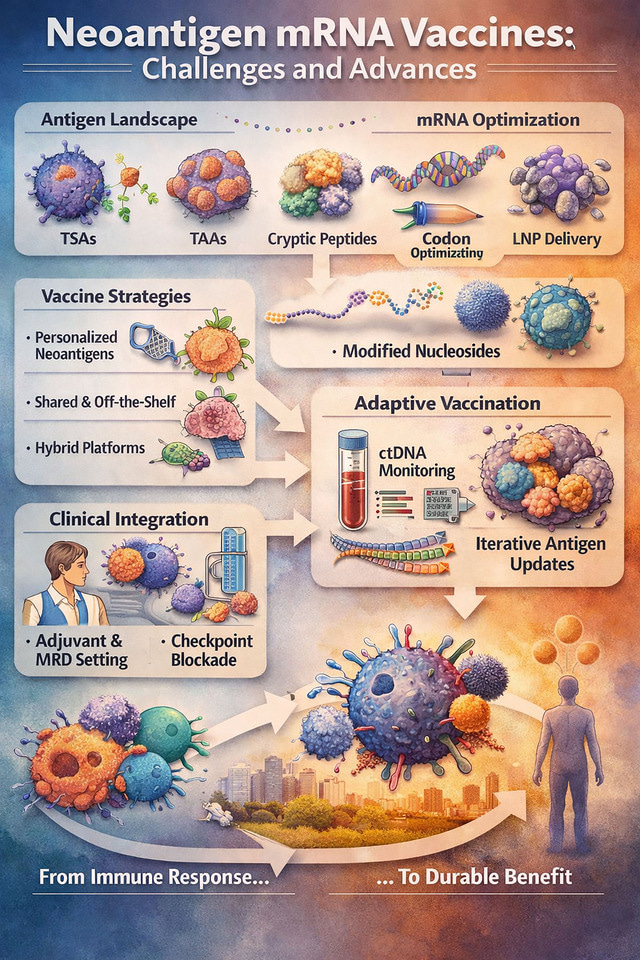
This work 1 was motivated by a persistent gap in the field: strong immunogenicity does not consistently translate into durable clinical benefit. Early clinical studies show that neoantigen mRNA vaccines can induce CD8⁺ and CD4⁺ T-cell responses and expand tumor-reactive clones, particularly in melanoma and NSCLC, yet clinical efficacy remains heterogeneous 2,3. This disconnect between immune activation and sustained tumor control defined the central focus of the paper. Importantly, the work also highlights that the antigen landscape itself is broader than previously assumed, extending beyond mutation-derived epitopes to include cryptic peptides generated from non-canonical translation events.
-
A continuum of antigen and vaccine strategies
A central contribution of this review is the structured classification of antigen and vaccine strategies. Tumor-specific antigens (TSAs), tumor-associated antigens (TAAs), shared neoantigens, and hybrid constructs represent distinct but interconnected approaches 4. Personalized TSA-based vaccines provide maximal specificity but are constrained by manufacturing timelines (6–8 weeks), cost, and tumor heterogeneity 5. Shared and off-the-shelf vaccines targeting recurrent driver mutations such as KRAS and TERT enable scalable deployment, while hybrid designs combine individualized and recurrent antigens to increase epitope diversity and reduce immune escape 6,7. Notably, the inclusion of cryptic neoantigens, derived from non-canonical ORFs, intronic regions, and lncRNAs, expands the targetable repertoire, particularly in tumors with low mutational burden 8,9.
-
Immunoengineering and functional optimization
A key advance highlighted in this work is the shift toward functional immunogenic optimization. Improvements in antigen prioritization, through WES, RNA sequencing, and immunopeptidomics, have enhanced neoepitope identification, although validation remains essential due to prediction inaccuracies 10. At the molecular level, mRNA engineering strategies, including codon optimization, nucleoside modification (e.g., m¹Ψ), UTR tuning, and multi-epitope construct design, have improved antigen expression and presentation. Parallel advances in LNP delivery systems and targeting strategies have enhanced dendritic cell uptake and T-cell priming, contributing to more consistent immune responses across diverse HLA backgrounds 11,12.
-
Adaptive vaccination and dynamic antigen targeting
A major forward-looking aspect of the paper is the concept of adaptive vaccination. Tumor evolution under immune pressure frequently results in antigen loss and clonal escape, particularly when targeting subclonal mutations. The integration of ctDNA monitoring enables real-time tracking of tumor dynamics and creates the possibility of iterative vaccine redesign during treatment 13,14. This approach shifts the paradigm from static vaccine constructs to dynamic, evolution-informed immunotherapy strategies capable of adjusting to tumor heterogeneity over time.
-
Future direction
The next phase of neoantigen mRNA vaccination will be defined by adaptability and integration. Tumor evolution and immune editing continue to limit the durability of fixed vaccine designs, making dynamic strategies increasingly important 15. Approaches incorporating ctDNA monitoring may enable real-time tracking of clonal changes and iterative updating of antigen composition during treatment. At the same time, hybrid and modular vaccine platforms that combine TSAs, shared neoantigens, and cryptic peptides from non-canonical translation offer a practical path to balance precision with scalability 16,17. Advances in delivery systems and immune modulation will further refine how these vaccines engage both innate and adaptive immunity 18. Clinically, their impact will depend on rational positioning, particularly in adjuvant and minimal residual disease settings, and integration with checkpoint blockade to overcome immune suppression 19,20.
-
Summar
Overall, this work positions neoantigen mRNA vaccination as a flexible and evolving therapeutic framework rather than a single platform. By expanding the antigen landscape to include diverse classes: TSAs, TAAs, shared, hybrid, and cryptic neoantigens, and integrating advances in antigen prioritization, mRNA engineering, and delivery, the field is moving toward more functionally optimized and clinically adaptable strategies. The central insight is that durable clinical benefit will depend not only on antigen specificity, but on aligning molecular design, immune activation, and clinical context within a coordinated translational approach.
- About Authors
Saber Imani is an independent Assistant Professor focused on mRNA-based cancer vaccines and translational oncology, with research centered on elucidating the molecular mechanisms underlying cancer and viral diseases, particularly in the context of immune checkpoint inhibitors (ICIs) and immunotherapy-driven treatment strategies. His work integrates mechanistic biology with therapeutic development, aiming to reprogram the tumor microenvironment to enhance anti-tumor immune responses and advance next-generation cancer therapies, including ICI-based interventions and mRNA vaccine platforms, with the broader goal of improving clinical outcomes and addressing critical challenges in oncology.
Parham Jabbarzadeh Kaboli is a cancer researcher based in the United States, specializing in translational pharmacology and tumor biology. He earned his Ph.D. in Pharmacology from Universiti Putra Malaysia in 2018 and has since developed a strong research profile, with an h-index of 30 and over 2,600 citations. His work focuses on the development of targeted therapeutic strategies and the mechanistic understanding of drug resistance in cancer, particularly in breast cancer. Dr. Kaboli has contributed to more than 40 peer-reviewed publications and has mentored emerging scientists across diverse academic environments. His research journey spans multiple international institutions, including Malaysia, China, Taiwan, and the United States, providing him with a broad and integrative scientific perspective. During his postdoctoral training at China Medical University in Taiwan, he conducted advanced studies on the c-MET receptor tyrosine kinase, exploring its role in tumor progression and therapeutic resistance. He is currently a Visiting Scientist at Aptegen LLC in York, Pennsylvania, where he is engaged in research at the interface of molecular targeting and translational innovation. Looking forward, Dr. Kaboli aims to establish an independent research program focused on overcoming therapeutic resistance in solid tumors through the integration of immunotherapy and clinically relevant strategies. He remains deeply committed to scientific mentorship, interdisciplinary collaboration, and advancing cancer research across global boundaries.
-
References
-
-
Li, X., et al. Next-generation neoantigen mRNA vaccines: Immuno-engineering strategies for precision cancer immunotherapy. Cell Oncol (Dordr) 49(2026).
-
Sahin, U., et al. An RNA vaccine drives immunity in checkpoint-inhibitor-treated melanoma. Nature585, 107-112 (2020).
-
Yang, J., et al. A pan-KRAS degrader for the treatment of KRAS-mutant cancers. Cell Discov 10, 70 (2024).
-
Jabbarzadeh Kaboli, P. & Proestos, C. Editorial: Glycobiology and glycosylation: deciphering the secrets of glycans in humans and pathogens. Front Immunol 16, 1560135 (2025).
-
Pao, S.C., Chu, M.T. & Hung, S.I. Therapeutic Vaccines Targeting Neoantigens to Induce T-Cell Immunity against Cancers. Pharmaceutics 14(2022).
-
Bond, M.J., et al. Rewiring the fusion oncoprotein EWS/FLI1 in Ewing sarcoma with bivalent small molecules. bioRxiv (2025).
-
Goncalves, G., Dolcetti, R., Ooi, J.D. & Faridi, P. Cryptic but critical: non-canonical antigens in cancer immunotherapy. Trends Immunol 46, 499-501 (2025).
-
Ely, Z.A., et al. Pancreatic cancer-restricted cryptic antigens are targets for T cell recognition. Science388, eadk3487 (2025).
-
Gao, Y., et al. ETV6 dependency in Ewing sarcoma by antagonism of EWS-FLI1-mediated enhancer activation. Nat Cell Biol 25, 298-308 (2023).
-
Ramadan, E., Ahmed, A. & Naguib, Y.W. Advances in mRNA LNP-Based Cancer Vaccines: Mechanisms, Formulation Aspects, Challenges, and Future Directions. J Pers Med 14(2024).
-
Han, X., et al. Plug-and-play assembly of biodegradable ionizable lipids for potent mRNA delivery and gene editing in vivo. bioRxiv (2025).
-
Wu, S., et al. Paracyclophane-based ionizable lipids for efficient mRNA delivery in vivo. Journal of controlled release : official journal of the Controlled Release Society 376, 395-401 (2024).
-
Bittla, P., et al. Exploring Circulating Tumor DNA (CtDNA) and Its Role in Early Detection of Cancer: A Systematic Review. Cureus 15, e45784 (2023).
-
Schenk, D., et al. Comprehensive Longitudinal ctDNA Monitoring in Metastatic Cancer Patients Treated with an Individualized Neoantigen-directed Vaccine. bioRxiv, 2024.2012.2004.626817 (2024).
-
Supabphol, S., Li, L., Goedegebuure, S.P. & Gillanders, W.E. Neoantigen vaccine platforms in clinical development: understanding the future of personalized immunotherapy. Expert Opin Investig Drugs 30, 529-541 (2021).
-
Rojas, L.A., et al. Personalized RNA neoantigen vaccines stimulate T cells in pancreatic cancer. Nature618, 144-150 (2023).
-
Laumont, C.M., et al. Noncoding regions are the main source of targetable tumor-specific antigens. Science translational medicine 10(2018).
-
Singh, P., et al. Advancements and challenges in personalized neoantigen-based cancer vaccines. Oncology reviews 19, 1541326 (2025).
-
Fatima, M., An, T. & Hong, K.J. Revolutionizing mRNA Vaccines Through Innovative Formulation and Delivery Strategies. Biomolecules 15(2025).
-
Blass, E. & Ott, P.A. Advances in the development of personalized neoantigen-based therapeutic cancer vaccines. Nat Rev Clin Oncol 18, 215-229 (2021).
-
Follow the Topic
Related Collections
With Collections, you can get published faster and increase your visibility.
Onco-proteogenomics
With the advancement of sequencing technologies, whole genome sequence information can be obtained robustly in a fairly short time and at low cost. Yet, currently there is often a lack of connection of genomic data to proteomic data, probably due to the fact that standard proteomics workflows rely on databases of only known wild-type/canonical proteins, but these databases generally do not integrate the data of cancer-associated proteins with countless amino acid sequence alterations due to gene mutations, post-translational modifications, and aberrant chromosome rearrangement. Therefore, in the next phase, it becomes extremely important to integrate experimentally-generated Mass Spectrometric data accordingly with the genomic information to identify tumor-specific proteins and isoforms.
Onco-proteogenomics has great promise and holds the key in improving our understanding of cancer biology and it is anticipated that the routine utilization of this integrated omics approach would become one of the gold standards of precision oncology. In this collection, we welcome original articles focusing on cancer proteogenomics research.
Publishing Model: Open Access
Deadline: Ongoing
Cancer-Associated Fibroblasts
Studies have shown that cancer-associated fibroblasts (CAFs) play a crucial role in the tumor microenvironment. Despite numerous research efforts in this area, targeting CAFs remains challenging. In this collection, Cellular Oncology welcomes submissions focusing on the recent developments in cancer-associated fibroblasts research. View our webinar recording on the topic of cancer-associated fibroblasts, with expert panelists discussing this exciting research field.
Publishing Model: Open Access
Deadline: Ongoing

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in