Protected by the peroxisome
Published in Bioengineering & Biotechnology
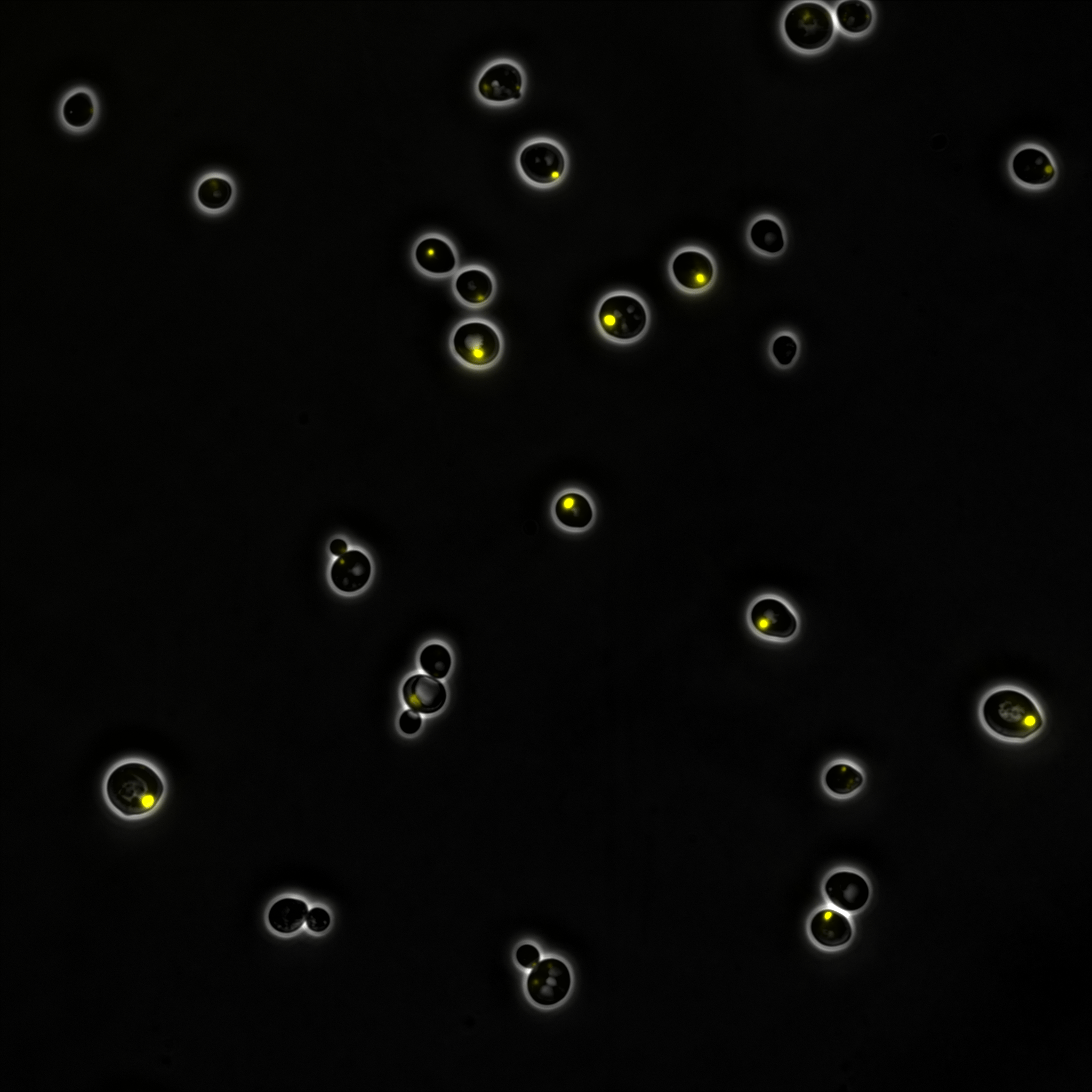

Plants produce a wide variety of metabolites that are used as medicines. Unfortunately, plant-based production requires a lot of land, is susceptible to adverse weather and climate change, and is difficult to reengineer for production of alternative metabolites with desired medicinal properties. Production of these plant metabolites in a fermentation process, using a production host such as yeast, is a promising strategy to improve efficiency and enable production of novel medicines.
While working on a specific plant metabolic family, the benzylisoquinoline alkaloids (BIAs), we discovered an interesting phenomenon. I had been producing variants of a key enzyme in the BIA pathway, norcoclaurine synthase (NCS), as part of my PhD work. Intriguingly, I found that when my best NCS variant was expressed in yeast, the yeast grew slower than normal. This outcome appeared to be linked to the NCS enzyme itself, because the toxicity was present even in the absence of substrates required for the reaction catalyzed by NCS.
As we began to think about ways to alleviate the observed toxicity, one approach that seemed promising was to target NCS to a subcellular compartment to protect the rest of the cell from NCS toxicity. We thought the peroxisome could be an ideal compartment because it has high capacity for protein and is permeable to small molecules, suggesting that a peroxisome-compartmentalized NCS would retain access to its small molecule substrates.
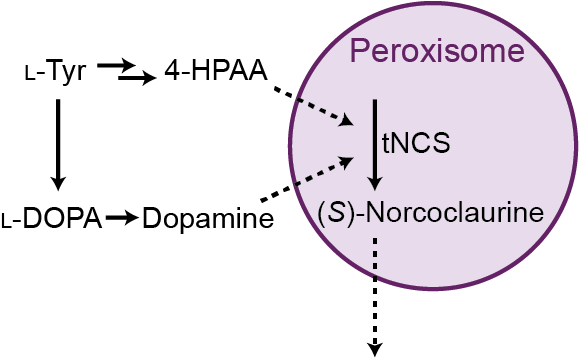
Of course, it helped that postdoc Jen Samson was our lab’s resident peroxisome expert and that Professor John Dueber had directed previous research on peroxisome engineering. Jen provided guidance when I initially tried expressing NCS in the peroxisome with undergrad Brian Choi. This preliminary experiment looked promising, so Jen and I teamed up to work on the project, in what became an invaluable collaboration for everyone involved. We demonstrated that NCS could be efficiently compartmentalized in the peroxisome, that compartmentalization improved growth of NCS-expressing yeast, and importantly, that compartmentalization increased production levels of key BIA molecules such as (S)-norcoclaurine and (S)-reticuline.
Because we were expressing NCS at high levels to produce more BIAs, we wondered how much NCS we could fit into the peroxisome. As we were testing the limits of this compartmentalization strategy by expressing toxic NCS at very high levels, graduate student Jordan Baker was working on an approach to increase the protein cargo capacity of peroxisomes. Proliferation of peroxisomes in terms of size and number naturally occurs when both glucose is absent and fatty acids are present in the environment. This proliferation is controlled by transcription factors that respond to environmental cues. Jordan engineered three transcription factors in the peroxisome proliferation network to be always “on” and independent of glucose and fatty acids, so that they would consistently induce peroxisome proliferation and increase peroxisome protein capacity. We utilized this innovative approach to compartmentalize toxic NCS and demonstrated further improved BIA production levels.
This project is an excellent example of how collaboration can bring together different perspectives and expertises when viewing the same problem. We anticipate that engineered compartmentalization will be used to address other problems in metabolic engineering, including the use of peroxisomes to sequester toxic proteins as described here.
Follow the Topic
-
Nature Chemical Biology
An international monthly journal that provides a high-visibility forum for the chemical biology community, combining the scientific ideas and approaches of chemistry, biology and allied disciplines to understand and manipulate biological systems with molecular precision.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in