Uncovering complex microbiome activities via metatranscriptomics during 24 hours of oral biofilm assembly and maturation
Published in Microbiology

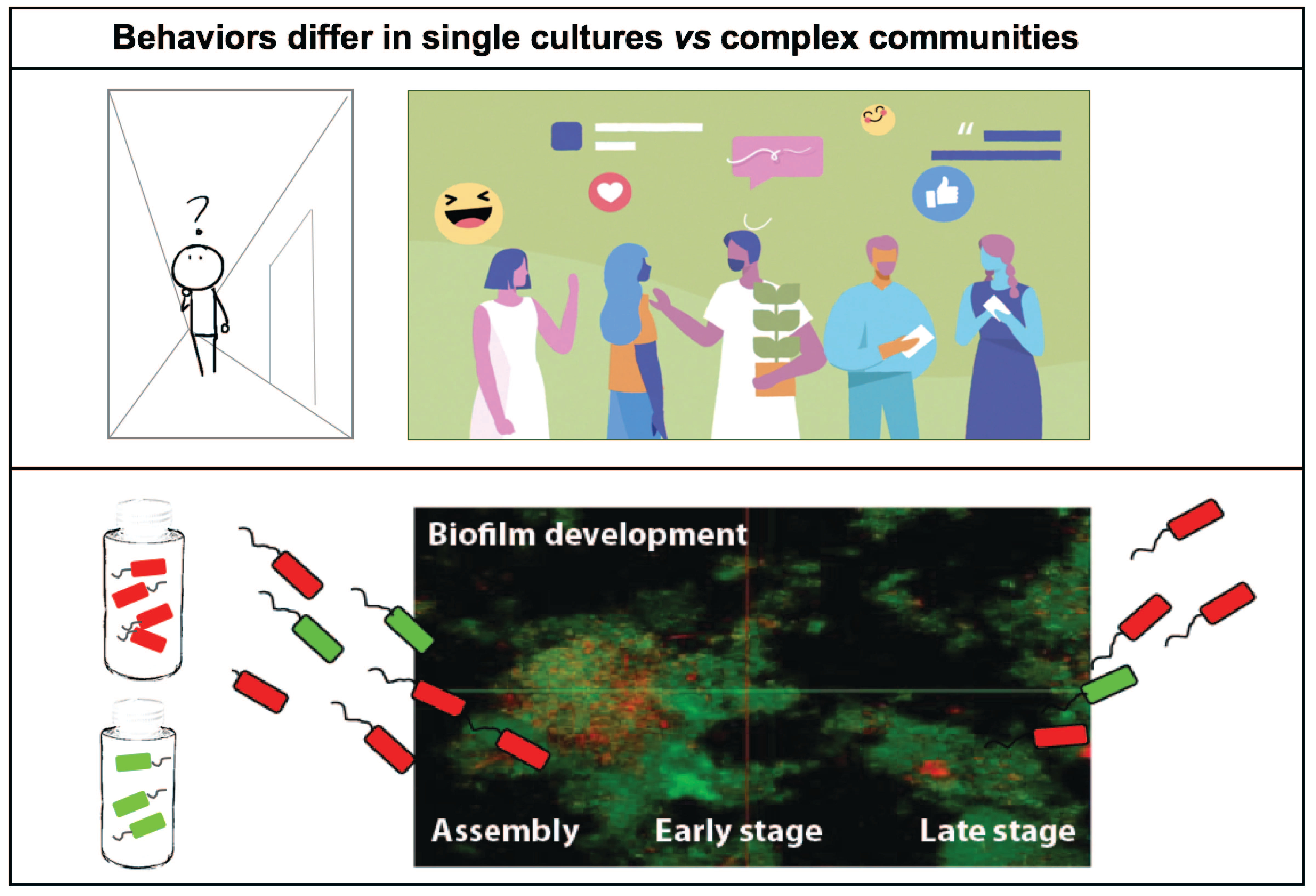
Oral bacteria growing as dental plaque have been known for several decades to catabolize dietary sugars and convert them into highly acidic metabolites (e.g. lactic acid), which cause tooth enamel erosion and eventually caries disease. With frequent consumption of carbohydrates, particularly when concurrent with a lack of oral hygiene, increased bacterial production of a sticky glucan matrix is favored, enmeshing cells and preventing diffusion of acidic metabolites. This results in a feed forward selection of acidogenic (acid producing) and aciduric (acid thriving) bacteria such as those belonging to mutant streptococci and lactobacilli. These bacteria dominate the dental plaque by producing copious amounts of lactic acid, delivered directly on the tooth enamel, which worsens the disease state and prevents reestablishment of health-associated community members.
Our first challenge was to develop a growth medium which captured growth of hundreds of bacterial plaque-growing species simultaneously. After iterations of media optimization, we succeeded in developing a novel growth medium that allows us to obtain a mixture of over a hundred of plaque bacteria by seeding as little as ten microliters of saliva in a total volume of one milliliter of growth medium. By using live/dead staining, confocal imaging and deep sequencing technologies (16S rRNA gene fragment sequencing and metagenomics) we were excited to discover that ~130 oral bacterial species were thriving in the sticky biofilm communities. To confirm our observations, we repeated the same experiments several times, seeded biofilm at different days in different 24-well plates, and confirmed there were no batch effects or biological biases – we realized it was possible to obtain the same community over and over again – like a perpetual machine!
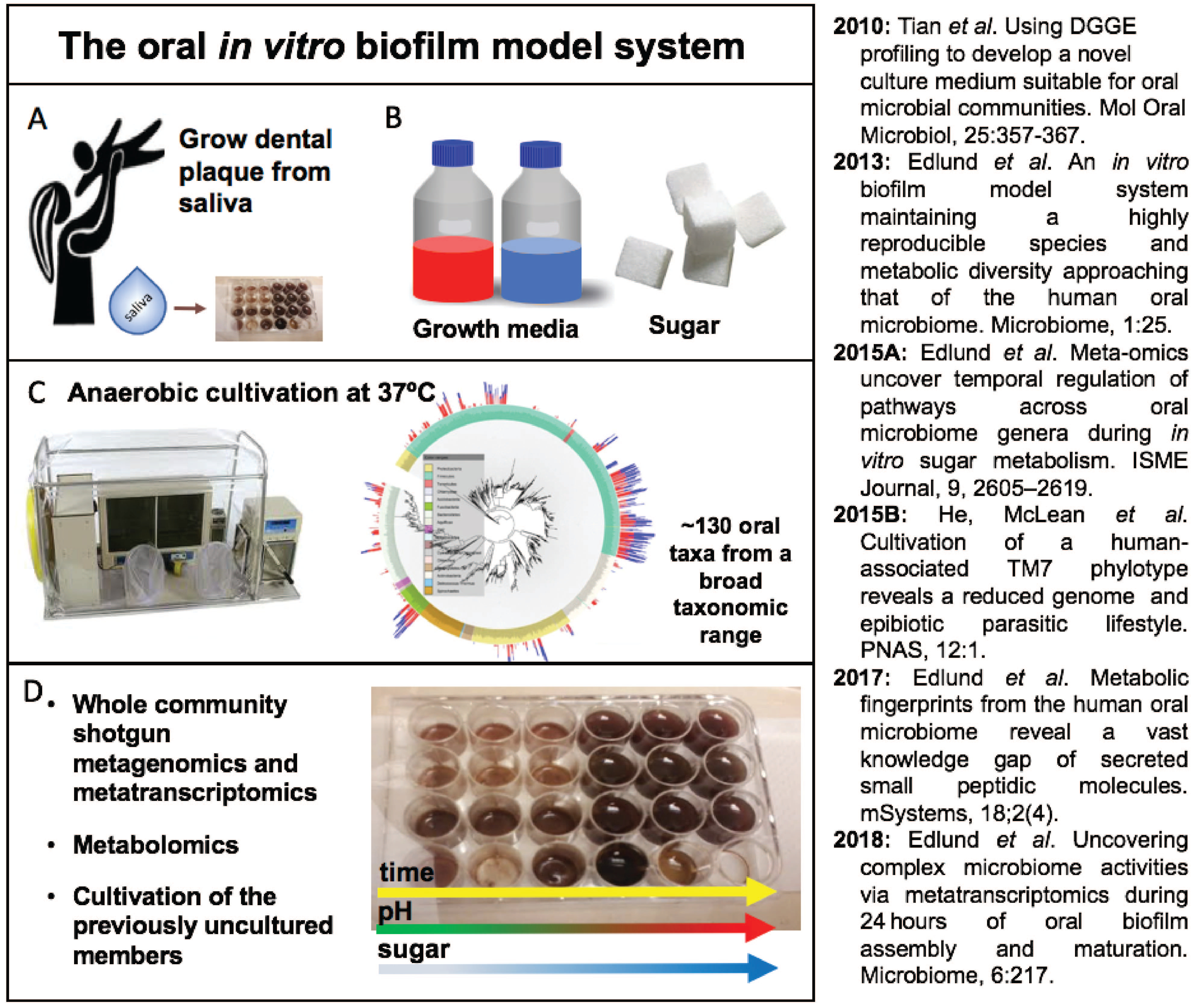

We made a major discovery when we found that many of the biofilm community members were previously uncultivated and belonged to the enigmatic Candidate Phyla Radiation, such as TM7 and SR1. By using the complex community as a selection platform our team continued the isolation and domestication of the first TM7 member, TM7x, and its bacterial host, an Actinomyces odontolythicus strain, XH001.
Next, we set out to answer questions about how health-associated communities can recover from the recurring pH drop after eating a sugary snack, and which metabolites and signaling molecules are produced during sugar catabolism that could possibly be used for bacterial interspecies, as well as bacterial-host communication.
The results from these studies lead to the opportunity to follow the growth expansion and bloom of the cariogenic pathogen Lactobacillus fermentum, as highlighted our study in the Microbiome journal. This represents the first study that addresses bacterial behaviors and ecosystem functions as a complex oral biofilm community initially assembles and matures. We captured species-unique gene expression patterns related to biofilm community invasion resistance, cell-to-cell signaling, cell attachment mechanisms, iron sequestration, low pH stress responses, and unique secondary metabolite biosynthetic pathways throughout 24 hours of growth.
With this new knowledge, we can prioritize studies of clinically and ecologically important bacterial taxa and molecular mechanisms, which hopefully will lead to future developments of novel therapies to prevent and treat caries disease.
Follow the Topic
-
Microbiome
This journal hopes to integrate researchers with common scientific objectives across a broad cross-section of sub-disciplines within microbial ecology. It covers studies of microbiomes colonizing humans, animals, plants or the environment, both built and natural or manipulated, as in agriculture.
Related Collections
With Collections, you can get published faster and increase your visibility.
Animal Gut Nutrition and Greenhouse Gas Mitigation
Animal Microbiome, Journal of Animal Science and Biotechnology and Microbiome call for submissions to the collection on Animal Gut Nutrition and Greenhouse Gas Mitigation.
Efforts to reduce greenhouse gas emissions from livestock systems increasingly hinge on innovations in animal gut nutrition. The dynamic relationship between the gut microbiome and nutrient utilization plays a pivotal role in shaping methane output, feed efficiency, and overall sustainability. Advances in microbial ecology—particularly in understanding the role of gut microbiome in nutrient metabolism—are opening new pathways for mitigating emissions while enhancing productivity. These developments support the implementation of climate-smart agricultural strategies to address climate change and its impacts.
Looking ahead, continued research in this field has the potential to yield innovative solutions such as targeted probiotic supplementation, which could further optimize gut function and enhance nutrient absorption. These advancements may lead to reduced greenhouse gas emissions while improving animal health and productivity. By deepening our understanding of the animal gut microbiome, we can contribute significantly to sustainable agricultural practices that benefit both the environment and food security.
We invite researchers to contribute to this special Collection on Animal Gut Nutrition and Greenhouse Gas Mitigation. Topics of interest include but are not limited to:
- Animal Gut Microbiome and Feed Efficiency
- Greenhouse Gas Mitigation Strategies
- Rumen Fermentation Dynamics
- Nutrient Utilization in Livestock
- Probiotic Supplementation Effects
- Sustainable Livestock Production Practices
- Climate-Smart Agriculture Innovations
This Collection supports and amplifies research related to SDG 13, Climate action.
All submissions in this collection undergo the relevant journal’s standard peer review process. Similarly, all manuscripts authored by a Guest Editor(s) will be handled by the Editor-in-Chief of the relevant journal. As an open access publication, participating journals levy an article processing fee (Animal Microbiome fees, Journal of Animal Science and Biotechnology fees, Microbiome fees). We recognize that many key stakeholders may not have access to such resources and are committed to supporting participation in this issue wherever resources are a barrier. For more information about what support may be available, please visit OA funding and support, or email OAfundingpolicy@springernature.com or the Editor-in-Chief of the journal where the article is being submitted.
Publishing Model: Open Access
Deadline: Sep 04, 2026
Harnessing plant microbiomes to improve performance and mechanistic understanding
This is a Cross-Journal Collection with Microbiome, Environmental Microbiome, npj Science of Plants, and npj Biofilms and Microbiomes. Please click here to see the collection page for npj Science of Plants and npj Biofilms and Microbiomes.
Modern agriculture needs to sustainably increase crop productivity while preserving ecosystem health. As soil degradation, climate variability, and diminishing input efficiency continue to threaten agricultural outputs, there is a pressing need to enhance plant performance through ecologically-sound strategies. In this context, plant-associated microbiomes represent a powerful, yet underexploited, resource to improve plant vigor, nutrient acquisition, stress resilience, and overall productivity.
The plant microbiome—comprising bacteria, fungi, and other microorganisms inhabiting the rhizosphere, endosphere, and phyllosphere—plays a fundamental role in shaping plant physiology and development. Increasing evidence demonstrates that beneficial microbes mediate key processes such as nutrient solubilization and uptake, hormonal regulation, photosynthetic efficiency, and systemic resistance to (a)biotic stresses. However, to fully harness these capabilities, a mechanistic understanding of the molecular dialogues and functional traits underpinning plant-microbe interactions is essential.
Recent advances in multi-omics technologies, synthetic biology, and high-throughput functional screening have accelerated our ability to dissect these interactions at molecular, cellular, and system levels. Yet, significant challenges remain in translating these mechanistic insights into robust microbiome-based applications for agriculture. Core knowledge gaps include identifying microbial functions that are conserved across environments and hosts, understanding the signaling networks and metabolic exchanges between partners, and predicting microbiome assembly and stability under field conditions.
This Research Topic welcomes Original Research, Reviews, Perspectives, and Meta-analyses that delve into the functional and mechanistic basis of plant-microbiome interactions. We are particularly interested in contributions that integrate molecular microbiology, systems biology, plant physiology, and computational modeling to unravel the mechanisms by which microbial communities enhance plant performance and/or mechanisms employed by plant hosts to assemble beneficial microbiomes. Studies ranging from controlled experimental systems to applied field trials are encouraged, especially those aiming to bridge the gap between fundamental understanding and translational outcomes such as microbial consortia, engineered strains, or microbiome-informed management practices.
Ultimately, this collection aims to advance our ability to rationally design and apply microbiome-based strategies by deepening our mechanistic insight into how plants select beneficial microbiomes and in turn how microbes shape plant health and productivity.
This collection is open for submissions from all authors on the condition that the manuscript falls within both the scope of the collection and the journal it is submitted to.
All submissions in this collection undergo the relevant journal’s standard peer review process. Similarly, all manuscripts authored by a Guest Editor(s) will be handled by the Editor-in-Chief of the relevant journal. As an open access publication, participating journals levy an article processing fee (Microbiome, Environmental Microbiome). We recognize that many key stakeholders may not have access to such resources and are committed to supporting participation in this issue wherever resources are a barrier. For more information about what support may be available, please visit OA funding and support, or email OAfundingpolicy@springernature.com or the Editor-in-Chief of the journal where the article is being submitted.
Collection policies for Microbiome and Environmental Microbiome:
Please refer to this page. Please only submit to one journal, but note authors have the option to transfer to another participating journal following the editors’ recommendation.
Collection policies for npj Science of Plants and npj Biofilms and Microbiomes:
Please refer to npj's Collection policies page for full details.
Publishing Model: Open Access
Deadline: Jun 01, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in