When cancer hides its origins: unique insights from Barrett's esophagus and esophageal adenocarcinoma
Published in Cancer and General & Internal Medicine
Explore the Research

Page not found | Nature
Page not found
There is a question that has followed esophageal cancer clinicians and researchers for a long time, and it goes something like this: if Barrett's esophagus is the precursor to esophageal adenocarcinoma, the cancer of the food pipe, then why do about half of all esophageal cancer patients have no detectable Barrett's at the time of diagnosis?
We set out to answer it. What we found challenged our own assumptions time and again.
Why this question kept us up at night
Esophageal adenocarcinoma (cancer of the food pipe) is one of the worst-outcome solid tumours, and its incidence has risen sharply in Western countries over the past four decades. The only clinically-established early detection route is surveillance of Barrett's esophagus, an endoscopically visible patch of abnormal lining that is considered the pre-cancerous state. The uncomfortable reality is that almost all patients (>90%)1 with this cancer have never been found to have Barrett's esophagus with the current cancer control approaches. Either we are missing an unknown, independent cancer progression pathway that bypasses Barrett's entirely, or Barrett's is always present, but we simply cannot see it. These two possibilities carry very different consequences for how we screen, clinically monitor, and how we framework cancer early detection.
The centrality of this question meant we needed a large, carefully annotated, and deeply phenotyped cohort of patients with research-grade matching normal, Barrett’s, and tumour biospecimens collected cross-sectionally and longitudinally for next generation sequencing. After two decades of work building the OCCAMS cohort, we thought the data and research conditions were finally satisfactory enough to begin studying this question.
What we did
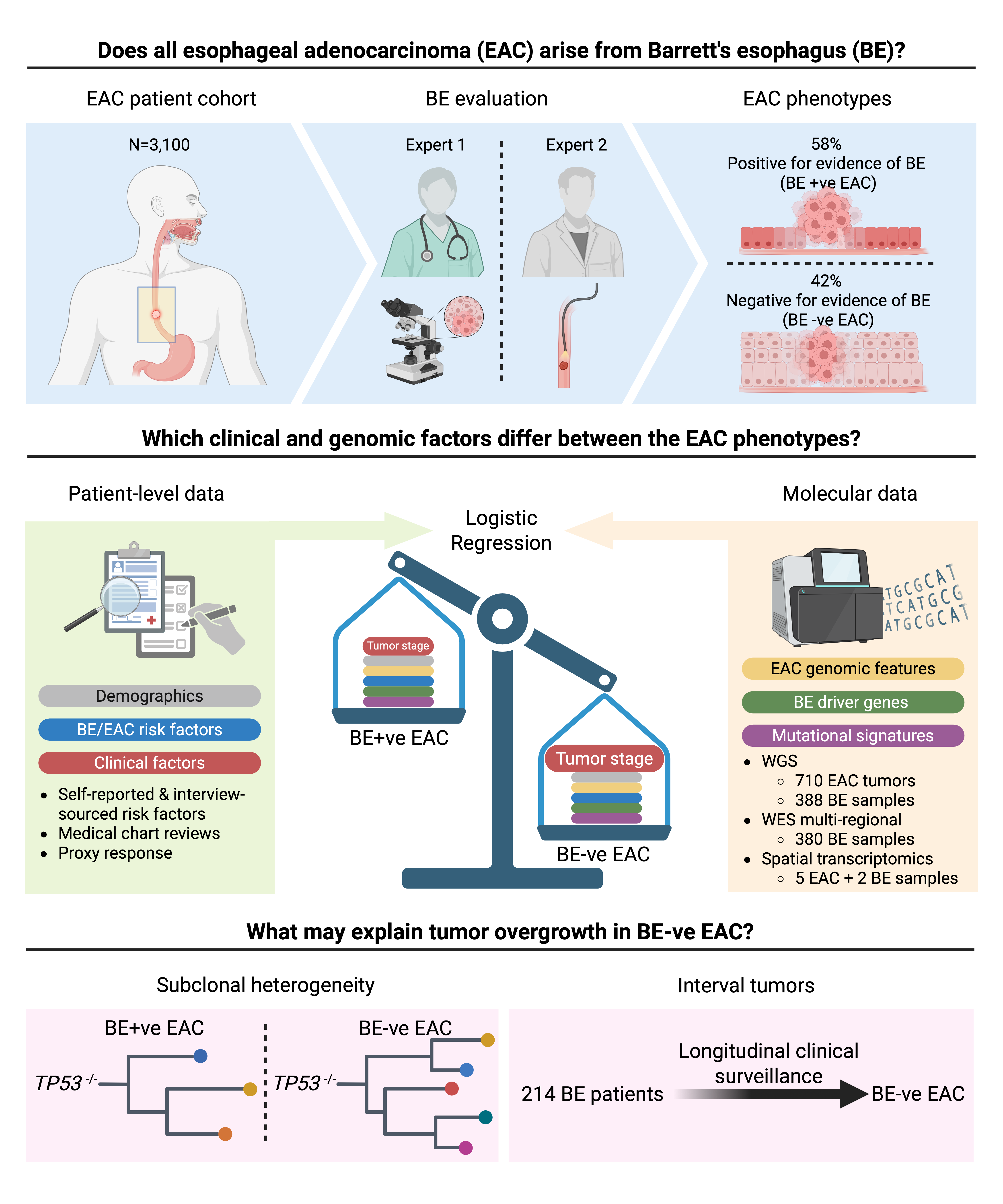
Through the OCCAMS consortium,2 a network of 25 UK hospitals, we have prospectively enrolled >4,000 patients with esophageal adenocarcinoma between 2002 and 2022 and included 3,100 in our study. For every single tumor, at least two expert pathologists reviewed an array of clinical and pathological reports, surgical specimens, and microscopy slides to determine whether Barrett's esophagus was present, absent, or unascertainable at the time of cancer diagnosis.
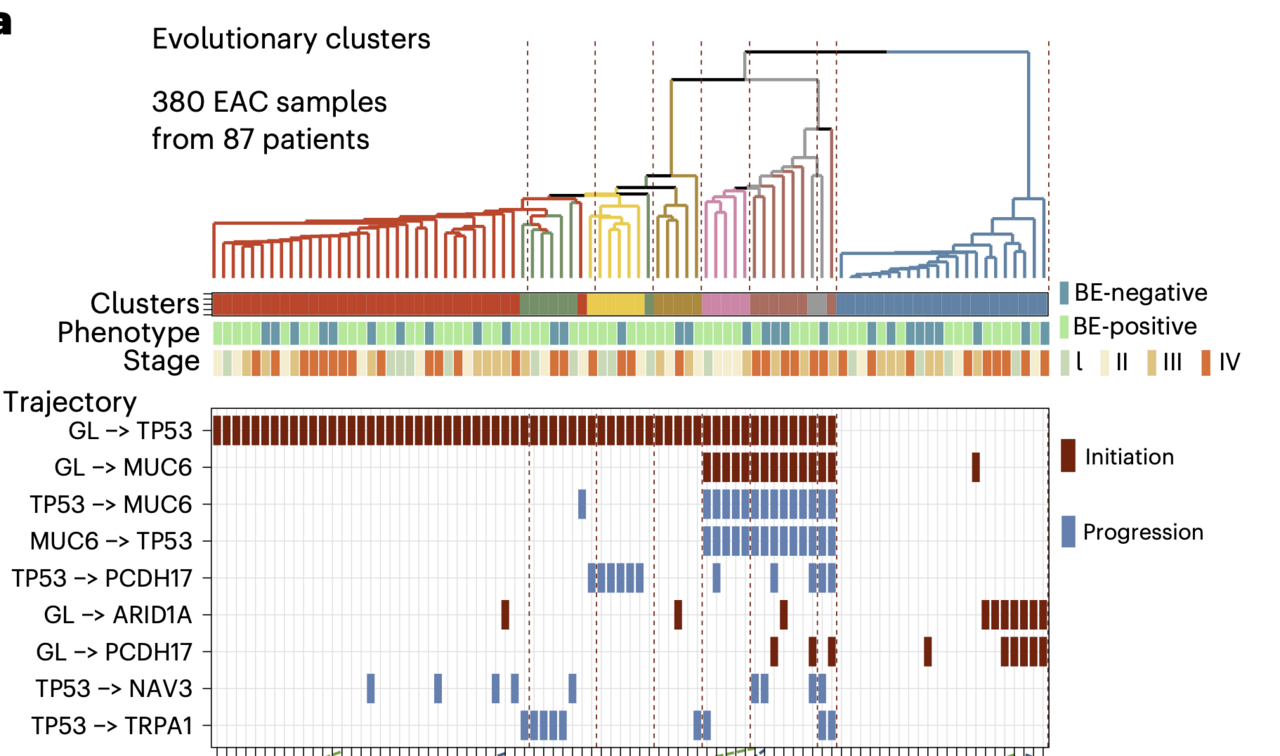
From this cohort, 710 patients had whole-genome sequencing of their tumours basically a complete readout of every DNA change the cancer cells had accumulated. A further 87 patients contributed whole-exome sequencing data from multiple regions within the same tumour, allowing us to reconstruct each cancer's evolutionary history, much like building a family tree. We also assembled a new cohort of 388 cancer-free Barrett's esophagus samples to give us a genomic picture of what the pre-cancerous state actually looks like.
And then, we added one more data layer: spatial transcriptomics that maps gene activity at near-cellular resolution across a tissue section, to ask whether the molecular identity of Barrett's esophagus was retained within tumours, even in patients without any visible Barrett's adjacent to their cancer.
What we found and what surprised us
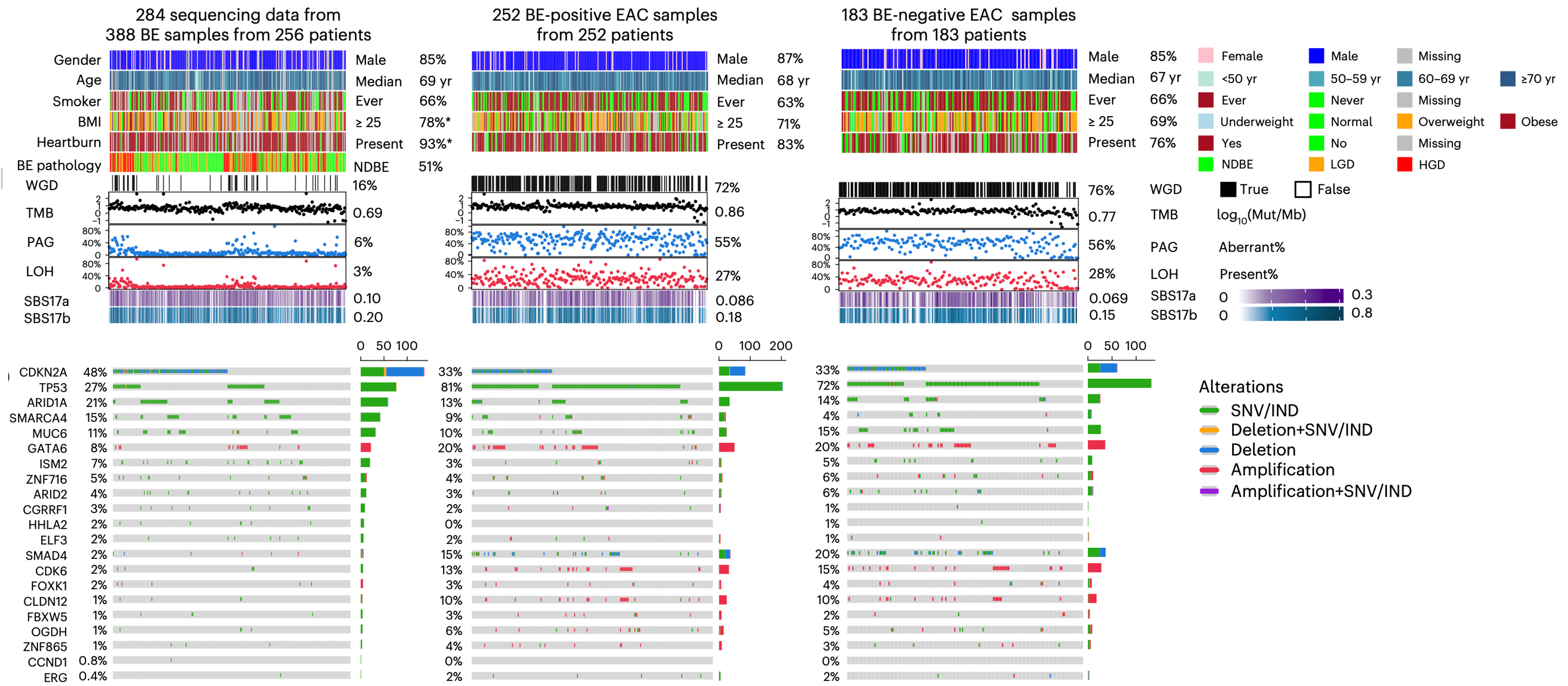
When the genomic data came back, the similarity between patients with and without Barrett's esophagus was striking. This was in addition to the remarkable similarity we had already observed given the patient-level epidemiological data analysis. The same genes were mutated at basically the same rates. The so-called mutational signatures, chemical "fingerprints" left in DNA by repeated acid and bile exposure, characteristic of Barrett's esophagus, were equally present in both groups. Tumour evolution also followed the same trajectories. Regardless of how many angles we looked from76 known esophageal cancer-driving genes, copy number changes across every chromosome arm, rare catastrophic genomic events, we could not find a feature that distinguished a "Barrett's-independent" cancer from a Barrett's-associated one.
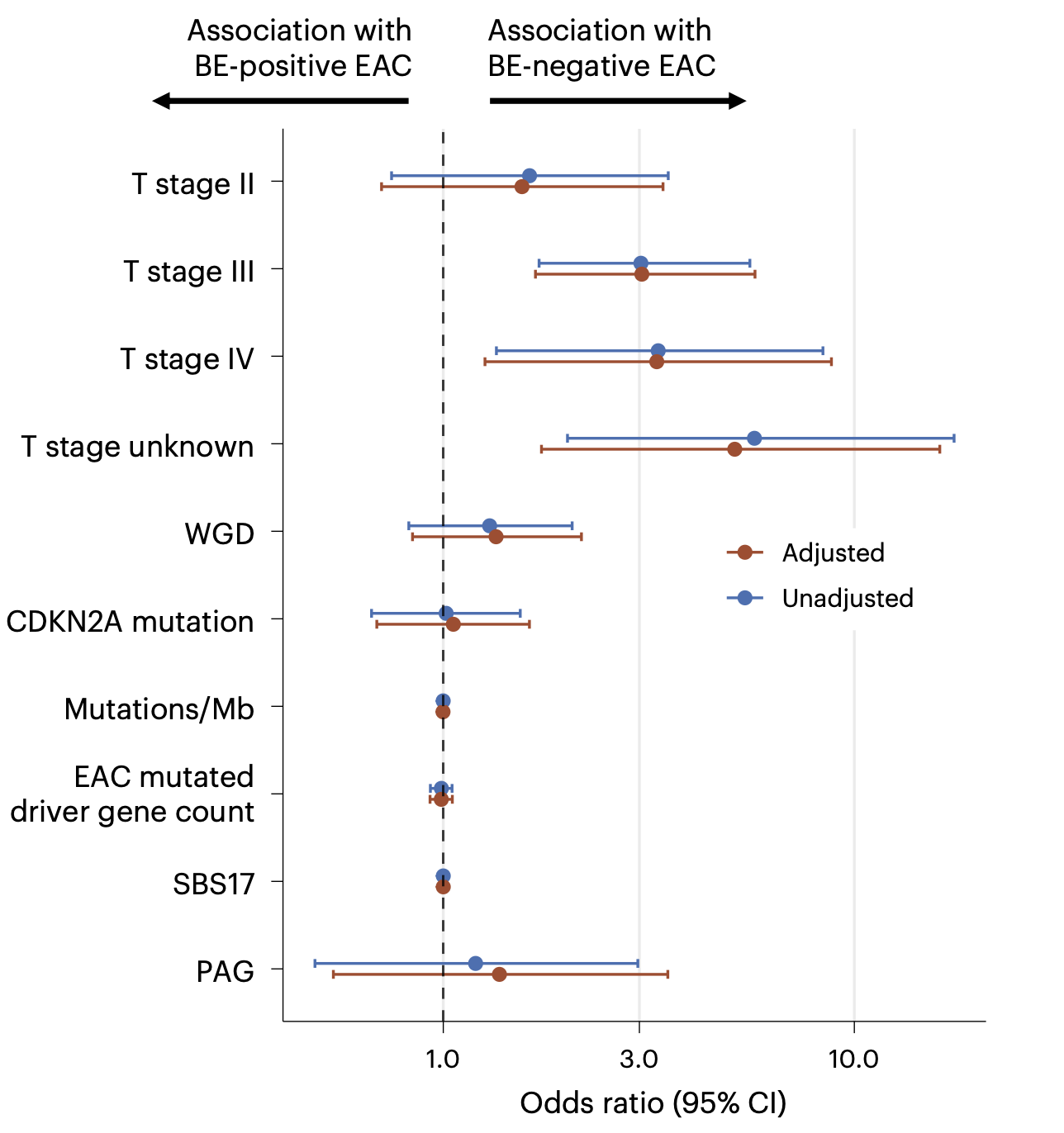
We next integrated the clinical and molecular data in an adjusted logistic regression model and found the only variable that reliably separated the two groups was tumour stage: patients without Barrett's presented with more advanced cancer.
A specific subgroup of patients held the key to the whole puzzle. Among patients who had been under Barrett's surveillance before their cancer diagnosis, 12.7% were Barrett's-negative when the cancer was found. Their Barrett's had not failed to exist, it had been consumed and overwritten by the growing tumour.
The spatial transcriptomics data sealed the argument. Even in tumours with no adjacent Barrett's tissue visible on pathology, the molecular programme of intestinal metaplasia, the cellular change that defines Barrett's, was still running inside the cancer cells. On average, 56% of tumour tissue, and up to 90% in some samples, expressed canonical Barrett's lineage genes. The precancer molecular signals were reverberating in the tumor, even after Barrett's had disappeared.
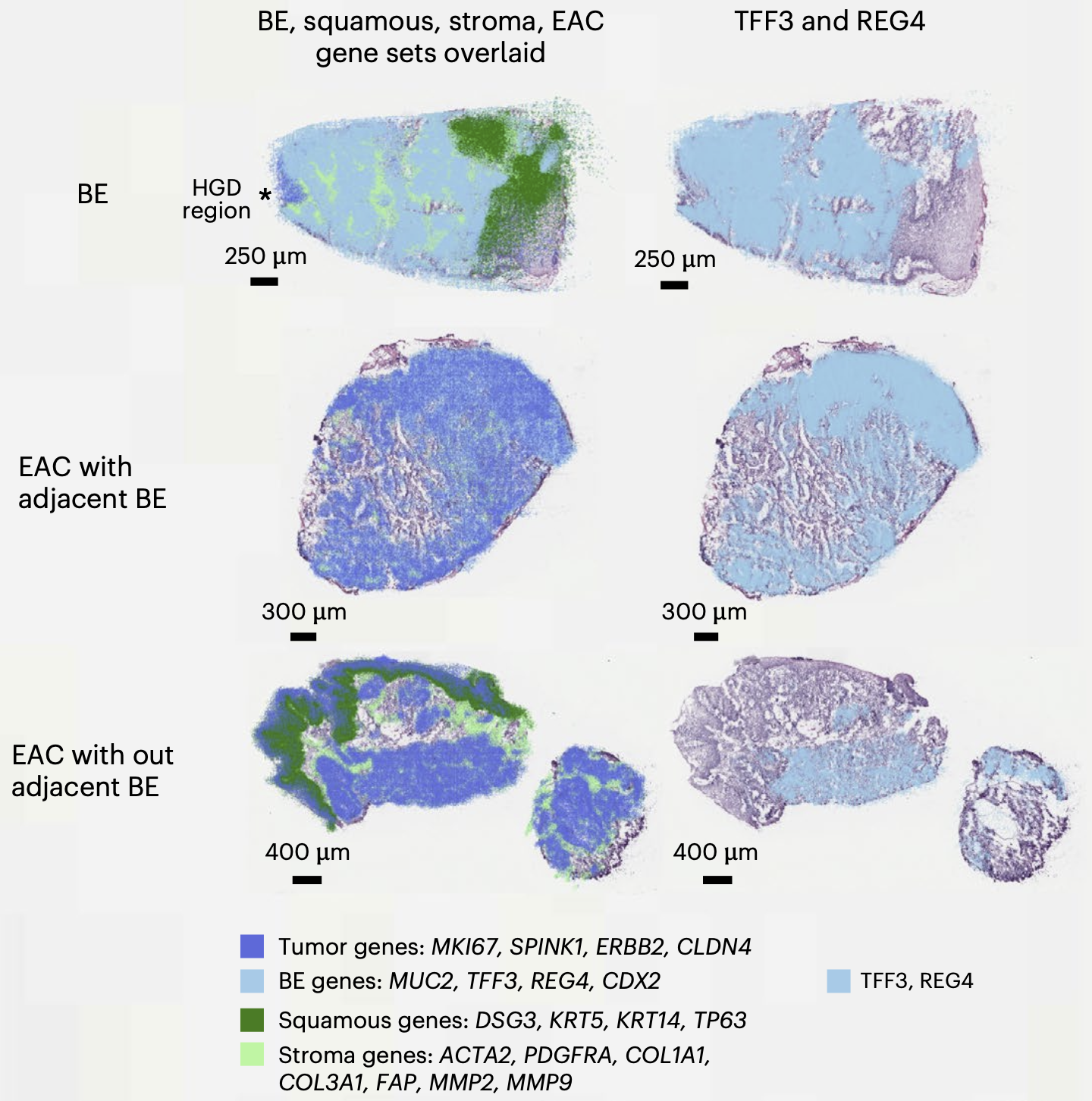
What this means for patients and the future of the field
Collectively, and with orthogonal findings from other independent researchers,3 our findings make a compelling case that there is one cancer pathway to oesophageal adenocarcinoma. Barrett's esophagus, specifically the intestinal metaplasia that defines it at a cellular level, is the universal precursor to this cancer. When it cannot be found at the time of cancer diagnosis, we think that is likely explained by a limitation of visual recognition, not evidence of a fundamentally different tumor biology at work.
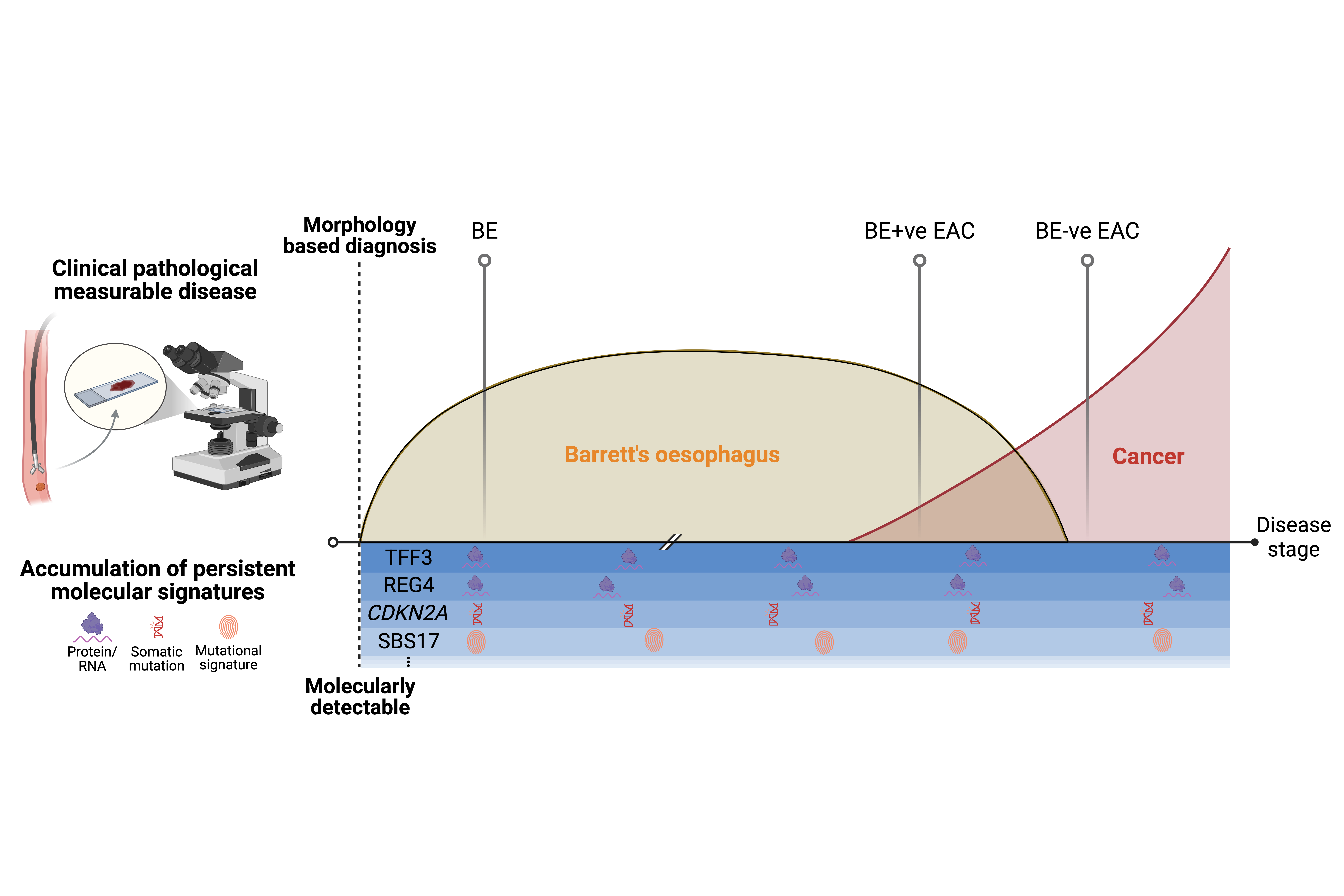
This matters significantly for how we approach cancer early detection. Current screening guidelines focus on patients with a visible, measurable Barrett's segment. But the true precursor may be smaller, shorter, or confined to microscopic changes that escape standard endoscopy. Protein markers like REG4 (OMIM: 609846) and the Trefoil Factors (collection of secretory proteins from Barrett's-type cells used in non-endoscopic screening tests) could offer a minimally invasive way to identify people at risk before cancer develops.
There are limitations we are honest about. We could not always ascertain Barrett's status with certainty, and our cohort is comprised of UK participants, therefore, limiting the generalizability of our findings. While we did adjust our analyses for potential confounders, we cannot completely rule out residual confounding or selection bias.
Future work should focus on improving clinical tools to more robustly localize intestinal metaplasia within the esophagus, and to risk-stratify patients when the Barrett’s metaplasia is on a path to cancer. The answer to the old question of "does esophageal cancer always begin with Barrett's?" now appears to be yes. The challenge now is to make sure we find it in time.
###
References
- Vaughan, T., Fitzgerald, R. Precision prevention of oesophageal adenocarcinoma. Nat Rev Gastroenterol Hepatol 12, 243–248 (2015). https://doi.org/10.1038/nrgastro.2015.24
- Peters, C.J., Ang, Y., Ciccarelli, F.D. et al. A decade of the Oesophageal Cancer Clinical and Molecular Stratification Consortium. Nat Med 30, 14–16 (2024). https://doi.org/10.1038/s41591-023-02676-y
-
-
Curtius K., Rubenstein J.H., Chak A., et al. Computational modelling suggests that Barrett’s oesophagus may be the precursor of all oesophageal adenocarcinomas. Gut 70, 1435-1440 (2021).
-
Follow the Topic
-
Nature Medicine

This journal encompasses original research ranging from new concepts in human biology and disease pathogenesis to new therapeutic modalities and drug development, to all phases of clinical work, as well as innovative technologies aimed at improving human health.
Related Collections
With Collections, you can get published faster and increase your visibility.
Microbiome and energy metabolism
Publishing Model: Hybrid
Deadline: Dec 06, 2026
Digital Medicine for Infectious Diseases
Publishing Model: Hybrid
Deadline: Nov 09, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in