A micrometer-scale self-assembled platform around a human centriole: How does it form, and what to make of it after all?
Published in Cancer
As an electron-dense, membrane-less organelle (approximately 100–300 aL in volume before undergoing mitotic expansion), the centrosome plays a critical role in promoting various cellular events, including cell division, cell motility, and intracellular signaling1,2. Multiple super-resolution imaging studies have revealed that pericentriolar material (PCM) proteins are highly organized, exhibiting layers of concentrically localized patterns around a centriole3-6. Yet how PCM scaffold proteins organize the PCM space remains largely unknown. Given that a micrometer-scale PCM is densely packed with a few hundred copies of several hundred different proteins7, properly organizing the PCM space is essential to achieve its functional proficiency8,9. Aberrant centrosome function predisposes humans to various diseases, including cancer, microcephaly, and neurological disorders10-13. Thus, unraveling the mechanism underlying PCM organization may reveal the fundamental principles governing centrosome functionality and could help determine the etiology of centrosome-associated human disorders.
A self-assembled cylindrical Cep63•Cep152 platform formed around a human centriole.
My group stumbled into this direction of research when we unexpectedly observed that the coexpression of a polo-like kinase 4 (Plk4)-binding protein, Cep15214,15, and its binding target, Cep6316, form a sphere-like assembly in the cytosol17. Plk4 plays a key role in centriole biogenesis by triggering downstream events, including the cartwheel-like Sas6 self-assembly found in the lumen of a centriole18,19. Notably, the expression of each Cep63 or Cep152 resulted in only cytosolic aggregates, suggesting that these two proteins work together to form a higher-order assembly. Subsequent studies with Cep63 and Cep152 revealed that these two long coiled-coil-encoding proteins interact with each other only through a short region in their C-terminal and middle region (i.e., Cep63-C and Cep152-M), respectively, by forming a heterotetrameric four-helical bundle arranged in an anti-parallel manner20. Further studies with purified recombinant proteins showed that these proteins can cooperatively generate a micrometer-scale “cylindrical” self-assembly in a Cep63 N-terminus-in and Cep152 N-terminus-out fashion on a slide glass. The architectural feature of the self-assembly resembles the cylindrical localization patterns of Cep63 and Cep152 around a centriole5,6. (Note several studies call this localization pattern “toroidal.”) Thus, unlike intermediate filament-forming long coiled-coil proteins, such as keratin, lamin, and vimentin, which form a long cable-like structure in an open cytosolic environment, Cep63 and Cep152 cooperatively generate a PCM architecture by self-assembling them around a centriole.
The assembly occurs through a stepwise and reversible process.
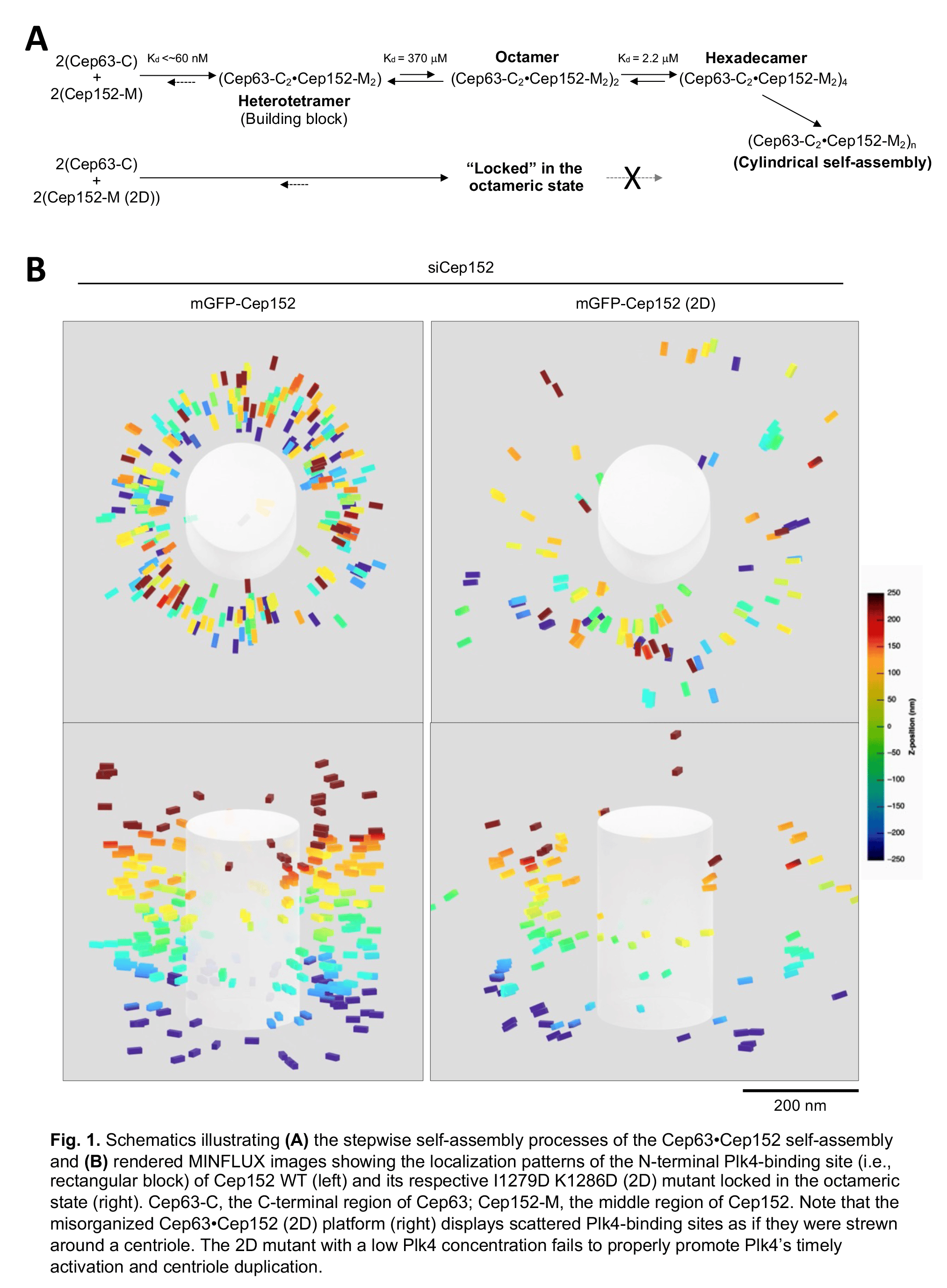
How do Cep63 and Cep152 self-assemble into a higher-order cylindrical architecture? In the Ahn, JI, et al. paper in Communications Biology (DOI: 10.1038/s42003-023-05067-8), our group combined MINFLUX nanoscopy with biophysical and structural analyses and theoretical modeling to demonstrate that the Cep63•Cep152 complex undergoes a concentration-dependent and reversible tetramer–octamer–hexadecamer transition to generate a cylindrical self-assembly21 (Fig. 1A). The formation of the hexadecameric Cep63•Cep152 complex was cooperative. With our experimental systems, we failed to detect a Cep63•Cep152 complex greater than the hexadecamer. Notably, although the assembly process appears highly dynamic, the self-assembled architecture remained stable for up to five days under physiological buffer conditions20. These two contrasting properties—intrinsic dynamics and steady-state stability—are the distinguishing characteristics of a self-organizing system22.
The octameric Cep152 (2D) mutant misorganizes the Cep63•Cep152 platform.
The significance of the stepwise self-assembly process becomes apparent from the experiment carried out with the I1279D and L1286D (2D) mutant, which forms a stable octameric form (i.e., having no reversible tetramer–octamer–hexadecamer transition) without any apparent propensity for self-association. Further characterization of the 2D mutant revealed that it forms severely defective cylindrical self-assemblies in vitro and misorganizes the pericentriolar Cep63•Cep152 platform in vivo, exhibiting a significantly reduced number of Cep152 molecules scattered in a wider radial space21 (Fig. 1B). These findings suggest that the ability of Cep152 to undergo stepwise formation into higher-molecular-weight complexes is essential for the dynamic recruitment and assembly of the Cep63•Cep152 building block around a centriole.
Effect of misorganizing the Cep63•Cep152 platform on Plk4 function and beyond.
Misorganizing the Cep63•Cep152 platform had far-reaching consequences, directly altering Plk4-dependent centriole biogenesis. This is because the functionality of Plk4, which binds to the N-terminus of Cep15214,15,23, is determined by the amount of Plk4 undergoing trans-autophosphorylation-induced liquid–liquid phase separation (LLPS) and ensuing ring-to-dot relocalization to the procentriole assembly site24-26. Moreover, since the LLPS is concentration-dependent, the dimension of the PCM space where Plk4 is distributed and the organizational pattern of Plk4 on the Cep63•Cep152 platform directly impact the efficiency of Plk4’s LLPS. Therefore, we propose that the organizational properties of the Cep63•Cep152 platform determine the timing of Plk4 activation and Plk4-dependent centriole biogenesis.
Intriguingly, multiple cancer-associated Cep63 and Cep152 mutations are found in human cancer tissues (Catalogue of Somatic Mutations in Cancer; https://cancer.sanger.ac.uk/cosmic) but not in the gnomAD (https://gnomad.broadinstitute.org), which generally represents wildtype variants. Several of these mutations are present within the regions forming the heterotetrameric Cep63•Cep152 complex. Thus, investigating the mutations’ significance could offer a deeper understanding about the architecture–function relationship of the Cep63•Cep152 complex. It may also help uncover new principles of building the Cep63•Cep152 self-assembly and provide valuable insights into the causes of PCM-associated human disorders.
References:
1 Conduit, P. T., Wainman, A. & Raff, J. W. Centrosome function and assembly in animal cells. Nature reviews. Molecular cell biology 16, 611-624 (2015). https://doi.org:10.1038/nrm4062
2 Vertii, A., Hehnly, H. & Doxsey, S. The centrosome, a multitalented renaissance organelle. Cold Spring Harb. Perspect. Biol. 8, a025049 (2016). https://doi.org:10.1101/cshperspect.a025049
3 Lawo, S., Hasegan, M., Gupta, G. D. & Pelletier, L. Subdiffraction imaging of centrosomes reveals higher-order organizational features of pericentriolar material. Nat. Cell Biol. 14, 1148-1158 (2012).
4 Mennella, V. K. et al. Subdiffraction-resolution fluorescence microscopy reveals a domain of the centrosome critical for pericentriolar material organization. Nat. Cell Biol. 14, 1159-1168 (2012).
5 Olivier, N., Keller, D., Gonczy, P. & Manley, S. Resolution doubling in 3D-STORM imaging through improved buffers. PLoS One 8, e69004 (2013). https://doi.org:10.1371/journal.pone.0069004
6 Sieben, C., Banterle, N., Douglass, K. M., Gonczy, P. & Manley, S. Multicolor single-particle reconstruction of protein complexes. Nat. Methods 15, 777-780 (2018). https://doi.org:10.1038/s41592-018-0140-x
7 Bauer, M., Cubizolles, F., Schmidt, A. & Nigg, E. A. Quantitative analysis of human centrosome architecture by targeted proteomics and fluorescence imaging. EMBO J. 35, 2152-2166 (2016). https://doi.org:10.15252/embj.201694462
8 Lee, K. S., Park, J. E., Il Ahn, J., Wei, Z. & Zhang, L. A self-assembled cylindrical platform for Plk4-induced centriole biogenesis. Open Biol. 10, 200102 (2020). https://doi.org:10.1098/rsob.200102
9 Lee, K. S., Park, J.-E., Ahn, J. I. & Zeng, Y. Constructing PCM with architecturally distinct higher-order assemblies. Curr. Opin. Str. Biol. 66, 66-73 (2021).
10 Fu, J., Hagan, I. M. & Glover, D. M. The centrosome and its duplication cycle. Cold Spring Harb Perspect Biol 7, a015800 (2015). https://doi.org:10.1101/cshperspect.a015800
11 Nigg, E. A. & Holland, A. J. Once and only once: mechanisms of centriole duplication and their deregulation in disease. Nature reviews. Molecular cell biology 19, 297-312 (2018). https://doi.org:10.1038/nrm.2017.127
12 Gonczy, P. Centrosomes and cancer: revisiting a long-standing relationship. Nat Rev Cancer 15, 639-652 (2015). https://doi.org:10.1038/nrc3995
13 Schatten, H. & Sun, Q. Y. Functions and dysfunctions of the mammalian centrosome in health, disorders, disease, and aging. Histochem Cell Biol 150, 303-325 (2018). https://doi.org:10.1007/s00418-018-1698-1
14 Hatch, E. M., Kulukian, A., Holland, A. J., Cleveland, D. W. & Stearns, T. Cep152 interacts with Plk4 and is required for centriole duplication. J. Cell Biol. 191, 721-729 (2010).
15 Cizmecioglu, O. et al. Cep152 acts as a scaffold for recruitment of Plk4 and CPAP to the centrosome. J. Cell Biol. 191, 731-739 (2010).
16 Brown, N. J., Marjanovic, M., Luders, J., Stracker, T. H. & Costanzo, V. Cep63 and cep152 cooperate to ensure centriole duplication. PLoS One 8, e69986 (2013). https://doi.org:10.1371/journal.pone.0069986
17 Ahn, J. I. et al. Phase separation of the Cep63•Cep152 complex underlies the formation of dynamic supramolecular self-assemblies at human centrosomes. Cell Cycle 19, 3437-3457 (2020).
18 van Breugel, M. et al. Structures of SAS-6 suggest its organization in centrioles. Science. 331, 1196-1199 (2011 ).
19 Kitagawa, D. et al. Structural basis of the 9-fold symmetry of centrioles. Cell. 144, 364-375. (2011).
20 Kim, T. S. et al. Molecular architecture of a cylindrical self-assembly at human centrosomes. Nat. Commun. 10, 1151 (2019). https://doi.org:10.1038/s41467-019-08838-2
21 Ahn, J. I. et al. Architectural basis for cylindrical self-assembly governing Plk4-mediated centriole duplication in human cells. Commun Biol in press (2023).
22 Misteli, T. The Self-Organizing Genome: Principles of Genome Architecture and Function. Cell 183, 28-45 (2020). https://doi.org:10.1016/j.cell.2020.09.014
23 Park, S. Y. et al. Molecular basis for unidirectional scaffold switching of human Plk4 in centriole biogenesis. Nature structural & molecular biology 21, 696-703 (2014). https://doi.org:10.1038/nsmb.2846
24 Yamamoto, S. & Kitagawa, D. Self-organization of Plk4 regulates symmetry breaking in centriole duplication. Nat. Commun. 10, 1810 (2019). https://doi.org:10.1038/s41467-019-09847-x
25 Park, J. E. et al. Phase separation of Polo-like kinase 4 by autoactivation and clustering drives centriole biogenesis. Nat. Commun. 10, 4959 (2019). https://doi.org:10.1038/s41467-019-12619-2
26 Park, J.-E. et al. Autophosphorylation-induced self-assembly and STIL-dependent reinforcement underlie Plk4’s ring-to-dot localization conversion around a human centriole. Cell Cycle, 3419-3436 (2020).
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
Mechanistic insights into human host and microbiome interactions
Publishing Model: Open Access
Deadline: May 31, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in