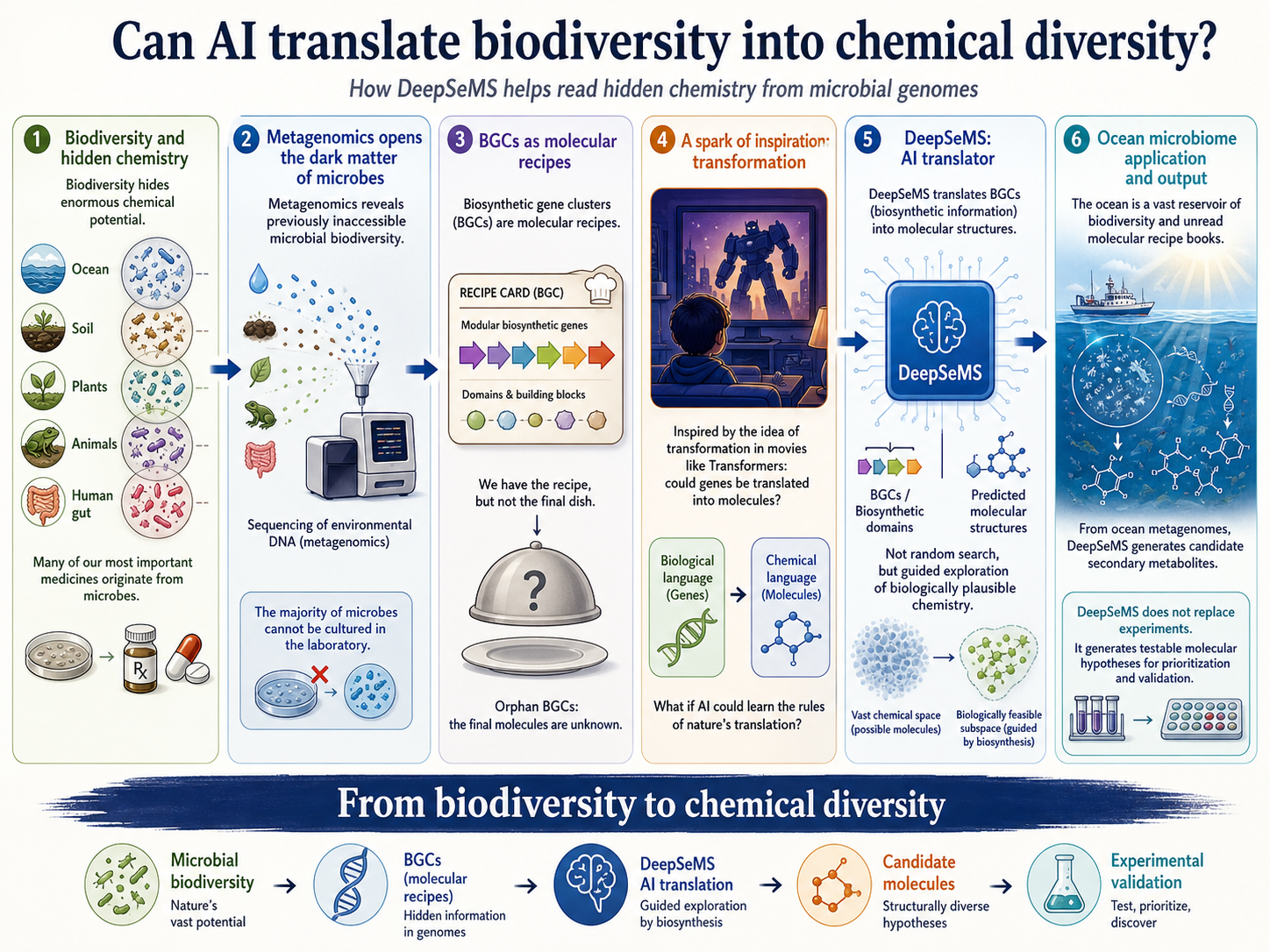
Biodiversity is not only a catalogue of organisms. Hidden within it is an enormous chemical potential. Across oceans, soils, plants, animals and our own bodies, microbes encode countless molecular recipes that shape ecosystems, mediate interactions and sometimes inspire new medicines. Many antibiotics, anticancer agents and other therapeutics have their origins in molecules made by microbes.
Yet for a long time, we could access only a small fraction of this potential. Most microbes cannot be easily grown in the laboratory, and many of their chemical products remain unknown. The rise of metagenomic sequencing changed the first part of this problem. Instead of culturing microbes one by one, researchers can now read genetic material directly from environmental samples such as seawater, soil or the human gut. This has revealed an immense landscape of microbial biodiversity, including many organisms that were previously invisible to science.
But sequencing also created a new challenge. Once we have this vast genetic information, how do we understand the chemistry it encodes? In other words, can we move from biodiversity to chemical diversity? This question led us to biosynthetic gene clusters (BGCs). A BGC is a group of genes that work together to produce a specialised molecule. One simple way to think about a BGC is as a molecular recipe. Just as a recipe contains instructions for turning ingredients into a dish, a BGC contains biological instructions for turning molecular building blocks into a chemical structure. These molecules can help microbes compete, communicate, defend themselves or adapt to their environments. Some may also have useful biological activities for humans.
The difficulty is that many BGCs are “orphan” clusters. We can identify them in genomes, but we do not know what molecules they produce. This is especially true in large metagenomic datasets, where many genes come from microbes that have never been cultured. We may have the recipe, but not the final dish. For natural product discovery, this creates both a problem and an opportunity: microbial genomes may contain a vast reservoir of hidden chemistry, but we need better ways to read it.
At the same time, generative artificial intelligence was changing how people thought about language, images and molecules. Transformer-based models showed that machines could learn complex relationships between one form of information and another. For us, the analogy was hard to ignore. Some of us grew up watching Transformers movies, where machines reconfigure themselves into entirely different forms. The scientific analogy was not perfect, of course, but it captured something important: biosynthesis also performs a kind of transformation. Information encoded in genes is converted into molecular structures.
That idea stayed with us. Could biosynthesis be treated as a translation problem?
In human language, translation converts a sentence in one language into a sentence in another while preserving meaning. In biosynthesis, genes and protein domains provide biological instructions, while molecules are the chemical outputs. A BGC is not merely a stretch of DNA. It is a structured program for molecular assembly. If an AI model could learn this program, perhaps it could translate biosynthetic information into candidate molecular structures.
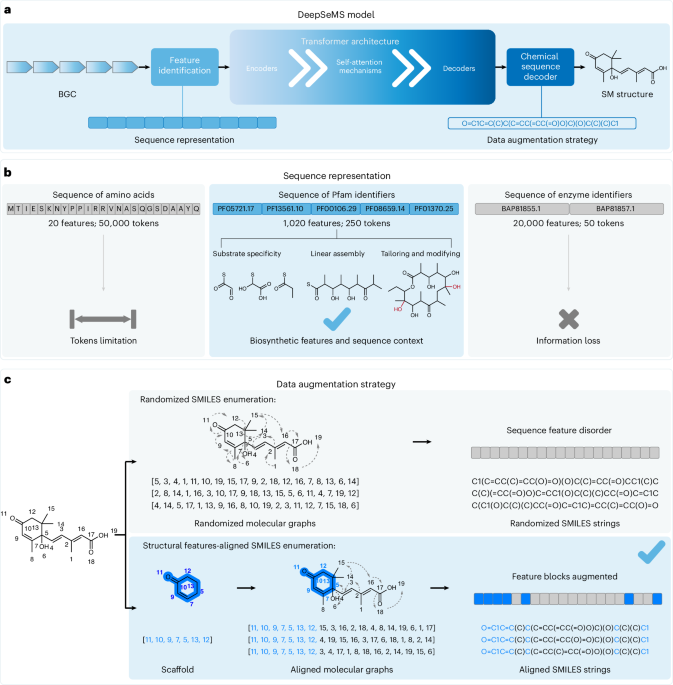
This idea became DeepSeMS. The goal of DeepSeMS is to predict plausible secondary metabolite structures from BGCs. Rather than searching randomly through the enormous universe of possible molecules, DeepSeMS uses biosynthetic information as a guide. This distinction is important. Chemical space is unimaginably large, and known molecules occupy only tiny regions within it. Random search would be inefficient. BGCs help narrow the problem because they tell us which kinds of molecular architectures are biologically plausible — that is, which molecules could reasonably be made by real microbial machinery.
Developing DeepSeMS was not straightforward. Early versions of the model were unstable, and some predicted molecules looked chemically possible but did not follow the biological logic of the gene clusters. We gradually realised that the key was not only to build a powerful model, but also to represent the biology carefully. The model needed to learn how biosynthetic domains, enzymatic rules and molecular structures correspond to one another. By improving this alignment, DeepSeMS became better at generating molecules that reflected the underlying biosynthetic logic.
One of the most exciting moments came when the model’s predictions began to make biological sense. The generated structures were not just arbitrary chemical strings. They reflected features of biosynthetic assembly, such as substrate preferences, chain extension and tailoring reactions. At that point, the project felt less like building a black-box molecular generator and more like building a translator — a translator between the language of microbial genes and the language of molecules.
We then applied DeepSeMS to the global ocean microbiome. The ocean is one of Earth’s largest reservoirs of biodiversity, and much of its microbial life remains uncultured and poorly understood. If BGCs are molecular recipes, then ocean metagenomes contain countless unread recipe books. Applying DeepSeMS allowed us to begin reading some of those pages and to generate candidate molecules from genomic information that would otherwise remain difficult to interpret.
Importantly, DeepSeMS does not replace experiments. Natural product discovery still requires chemical isolation, synthesis, activity testing and biological validation. What DeepSeMS provides is a way to make hidden chemistry more visible, searchable and testable. It turns orphan BGCs into molecular hypotheses. Researchers can then prioritise which predictions look most interesting and which gene clusters may deserve experimental follow-up.
Looking back, the most rewarding part of this work was the conceptual shift. We started with a practical problem: how can we predict the products of orphan BGCs? But the project became a broader way to think about discovery. Metagenomics reveals biodiversity, but biodiversity alone is not enough. To understand its chemical potential, we need tools that connect genes to molecules. DeepSeMS is one attempt to build that connection.
The microbial world has been encoding complex chemistry for billions of years, but much of this chemistry remains hidden in genomic data. With DeepSeMS, we hope to provide a more systematic way to explore it. Our long-term vision is to move from simply cataloguing microbial genes to understanding the molecules they may encode — and ultimately to help translate biodiversity into new chemical possibilities.
Follow the Topic
-
Nature Computational Science
A multidisciplinary journal that focuses on the development and use of computational techniques and mathematical models, as well as their application to address complex problems across a range of scientific disciplines.
Related Collections
With Collections, you can get published faster and increase your visibility.
Physics-Informed Machine Learning
Publishing Model: Hybrid
Deadline: May 31, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in