Construction and modular implementation of the THETA cycle for synthetic CO2 fixation
Published in Chemistry and Cell & Molecular Biology
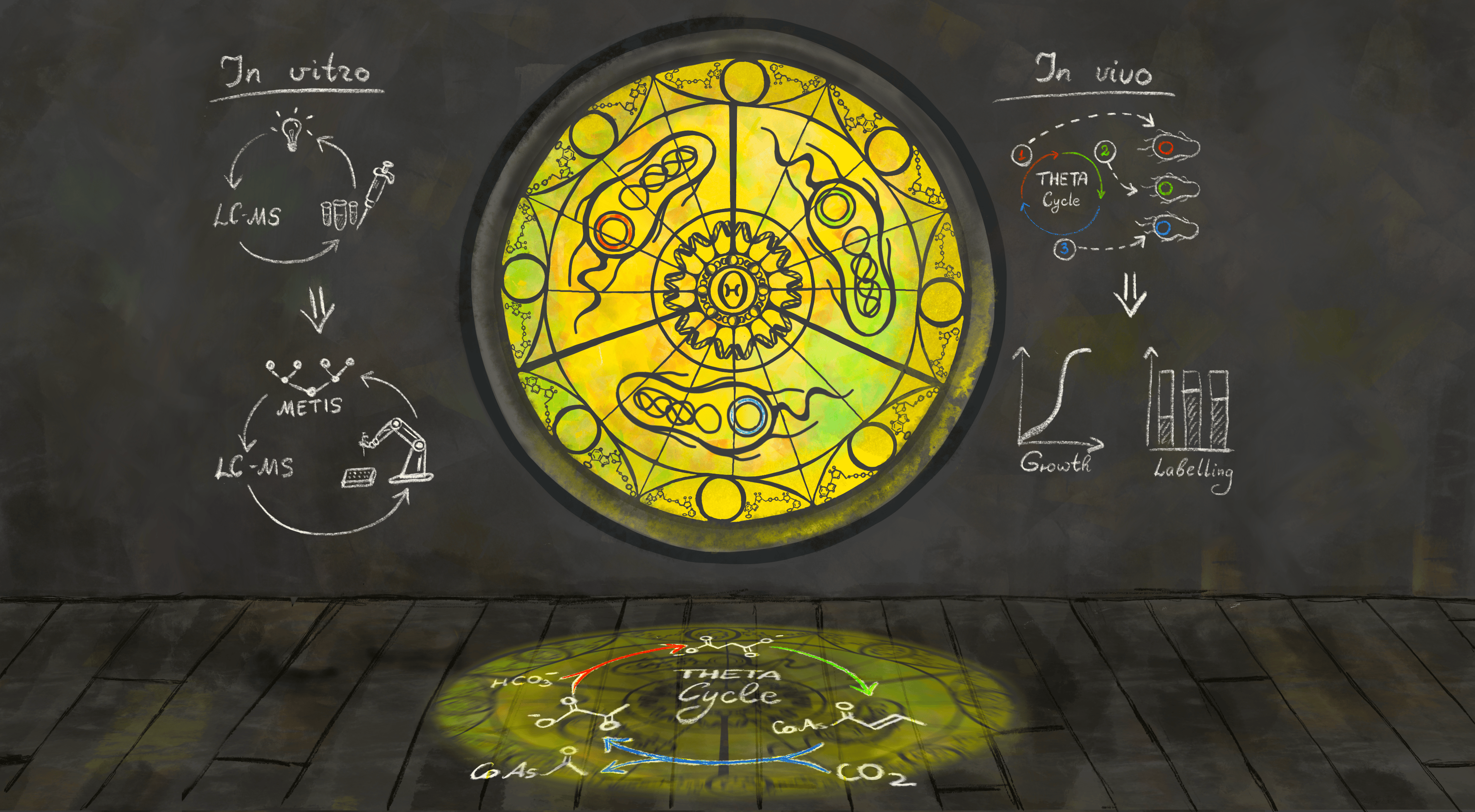
To tackle the climate crisis, developing technologies for carbon fixation, storage and utilization has become an urgent mission for our era. For this, synthetic biologists have designed novel CO2-fixation pathways that outperform natural ones. However, bringing these new-to-nature designs into reality, especially constructing them in the contexts of living cells, is still challenging. Synthetic autotrophy – the conversion of a heterotrophic organism like the model microbe E. coli into one that can grow with CO2 as sole feedstock – has so far only been achieved via the natural Calvin-Benson-Bassham (CBB) cycle1. While the latter suffers from the lacking specificity and low turnover of the core enzyme ribulose-1,5-bisphosphate carboxylase/oxygenase (RuBisCO), synthetic CO2-fixation cycles overcome these issues by employing the world’s fastest, most specific carboxylases2,3. In case of the THETA cycle, these carboxylases are crotonyl-CoA carboxylase/reductase (Ccr) and phosphoenolpyruvate carboxylase (Ppc) (Figure 1). With a cost of four ATP per pyruvate (instead of seven in the CBB cycle), the THETA cycle is amongst the most efficient designs available to date2. So far, more than 30 synthetic CO2-fixation pathways have been proposed, of which three oxygen-insensitive ones (CETCH4, rGPS-MCG5, HOPAC6) have been demonstrated in vitro. However, demonstrating the in vivo feasibility of such synthetic CO2-fixation pathways is still a challenge to be overcome in synthetic biology. While the CETCH cycle consumes only two ATP and eight NAD(P)H per pyruvate, it contains mainly CoA esters as intermediates and has glyoxylate as its output molecule. This limited overlap with E. coli's central metabolism poses challenges for its integration and evolution within the cell. The THETA cycle, like the CETCH cycle, is a complex synthetic CO2-fixation pathway. However, it also contains several central metabolites as intermediates (pyruvate, phosphoenolpyruvate, oxaloacetate, succinyl-CoA) and produces acetyl-CoA, making it easier to be plugged into the central metabolism of E. coli.
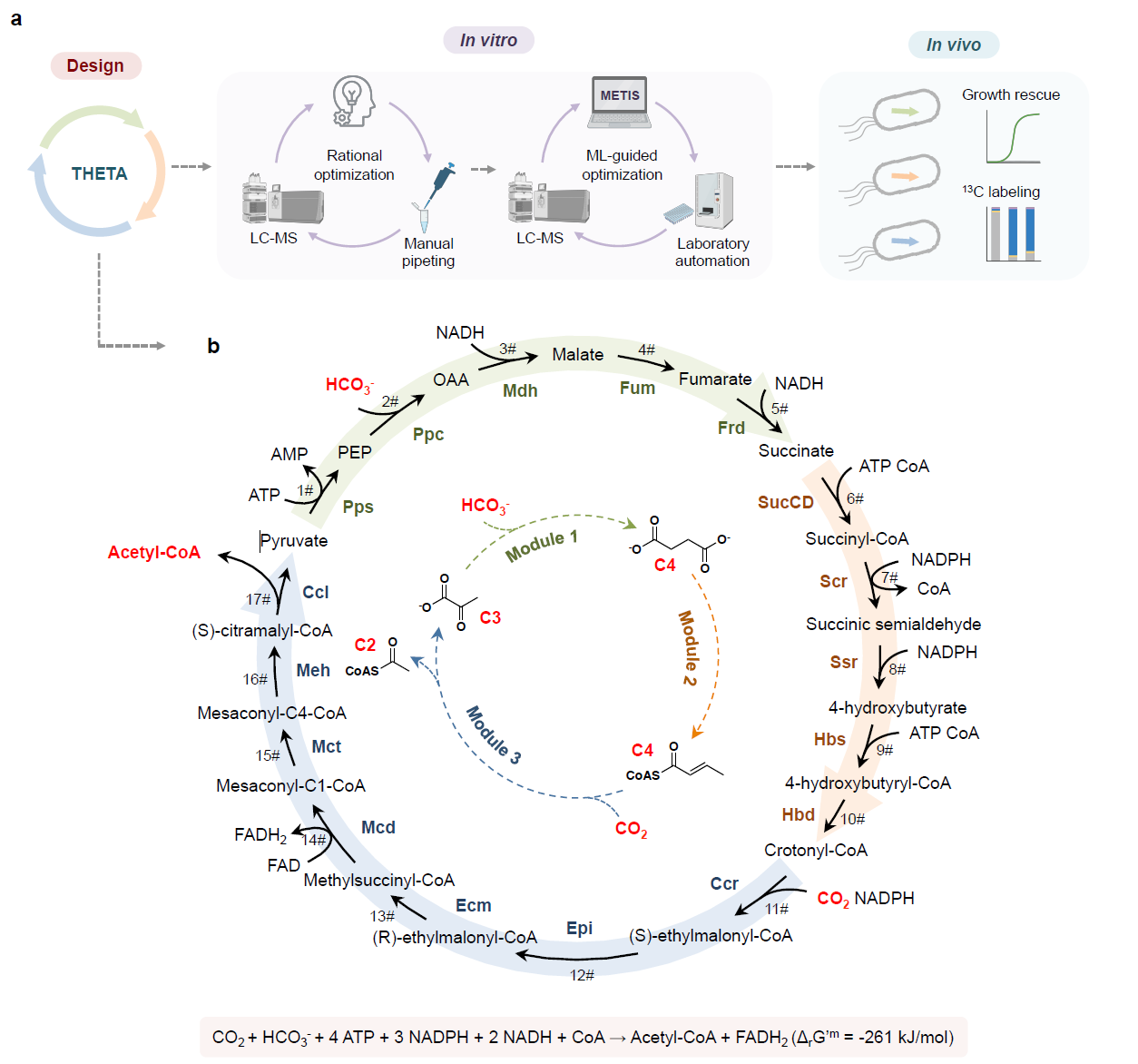
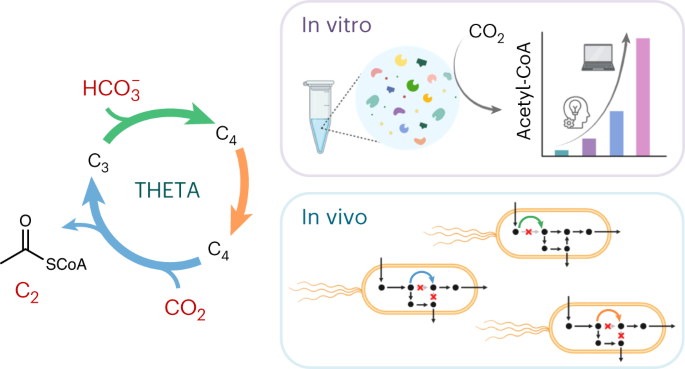
Fig. 1. The THETA cycle and its realization. (a) Overall strategy used in this work to establish the THETA cycle. The THETA cycle was designed in three modules, constructed in vitro, optimized through rational approaches and machine learning, before its three modules were implemented in E. coli. (b) Scheme of the THETA cycle. Inner circle: simplified scheme of the THETA cycle, consisting of 3 pathway modules. Outer circle: detailed scheme of the THETA cycle.
To realize the THETA cycle (Figure 1), we first demonstrated its functionality in vitro: by combining all enzyme activities and LC-MS-based intermediate quantification, we could confirm the desired cycle activity. However, further testing revealed several bottlenecks, which we could not anticipate during the theoretical cycle design: first, we noticed that a dead-end product (malyl-CoA) accumulated due to the promiscuous activity of one cycle enzyme. Here, we solved the problem by adding a proofreading enzyme to funnel the flux back into our cycle. To find further bottlenecks, we started the cycle from different intermediates, which quickly revealed that the fumarate reductase (Frd) was rate-limiting. Upon designing and incorporating a two-enzyme Frd bypass, we further increased the performance. When we could not find clear indications for bottlenecks with our rational approach, we proceeded with a machine learning-guided approach. By employing the recently developed active learning-guided workflow METIS7 and lab automation, we could iteratively vary the concentrations of enzymes, cofactors, and buffer conditions, which ultimately led to another 5-fold boost in the cycle performance. To further expand the in vitro applications of the THETA cycle, we added different pathway modules to extend the optimized THETA variant, and successfully demonstrated glycolate and malonyl-CoA production from CO2.
After confirming sufficient THETA cycle performance in vitro, we next faced the biggest challenge of current synthetic CO2 fixation: engineering the cycle in vivo, within the metabolic context of the model organism E. coli. To couple the growth of the microbe to the use of the synthetic metabolic routes, we created selection strains with auxotrophies for THETA intermediates and divided the cycle into three distinct modules to test individually: module one concerned the conversion of pyruvate into succinate via the carboxylase Ppc; module two was the reductive synthesis of crotonyl-CoA from succinate; and module three was designed around the carboxylase Ccr to yield pyruvate and acetyl-CoA from crotonyl-CoA. Of these, testing module one in a succinyl-CoA auxotrophic strain immediately confirmed the activity in vivo. For module two, we expressed additional enzymes to rescue an acetyl-CoA auxotroph from the module product crotonyl-CoA, which eventually worked after short-term evolution. The bottleneck for our in vivo implementation turned out to be module three, which could not provide enough flux to rescue our acetyl-CoA sensor strain. While metabolomics analysis revealed native thioesterases could be problematic for diverting the flux from methylsuccinyl-CoA to methylsuccinate, we also found that the native succinate dehydrogenase could convert methylsuccinate to mesaconate. These findings made us redraft an “acid-bypass” module 3, for which we confirmed in vivo activity by 13C-labeling. In sum, we demonstrated the functionality of each THETA module in vivo. Here, the number of non-native reactions and metabolites per module was correlated with our engineering difficulties. Promiscuous activities of native enzymes with our module intermediates significantly complicated the implementation, but could – once identified – in some cases be employed in favor of the engineering effort. Notably, predicting such routes from the get-go is difficult, since the intermediates of the synthetic pathway are not native to the host metabolism. Therefore, we relied on evolution and thorough system characterization for their identification. In retrospect, using growth as readout for module performance required significant module flux, so that measuring enzyme activities in cell lysates and LC-MS-based metabolomics analysis became extremely relevant for troubleshooting less functional modules.
While the current work proves the feasibility of the THETA cycle for both in vitro and in vivo applications, we aim to optimize both in the future. In vitro, we on the one hand seek to extend the output spectrum for sustainable productions. Acetyl-CoA, the output of the THETA cycle, is a central building metabolite for various valuable products, including polyhydroxyalkanoates (PHAs), ethanol, butanol, polyketides, fatty acids, as well as fatty-acid-derived fuels and chemicals8-11. This expansion could result in a versatile in vitro platform for converting CO2 into valuable chemicals. Furthermore, we recently developed a system interfacing electrical and biological energy currencies12, which we want to combine with the THETA cycle in the near future. In vivo, we are currently working on the improvement of module three to allow cellular growth. Once this is achieved, we expect that further optimization will be required to improve the joint performance of multiple modules. As was reported for the CBB cycle13, achieving synthetic autotrophy via the THETA cycle will likely require extensive laboratory evolution and a careful transition from hemiautotrophic growth to purely CO2-dependent growth.
In sum, we brought the THETA cycle from theoretical design to life and demonstrated its great potential for sustainable biosynthesis from CO2. The complexity of this venture required working in a highly interdisciplinary fashion, which was made possible by the close collaboration between all authors, each bringing expertise from their fields, including pathway design, enzyme and strain engineering, computational modeling, high-throughput workflows, and metabolomics. In that, this work hopefully serves as a blueprint for future synthetic biology endeavors to build new-to-nature solutions for a sustainable future.
Conceptualization and editing: Shanshan Luo
Writing: Helena Schulz-Mirbach
Illustration credit: Elizaveta Bobkova
References
- Gleizer, S. et al. Conversion of Escherichia coli to Generate All Biomass Carbon from CO2. Cell 179, 1255-1263.e12 (2019).
- Löwe, H. & Kremling, A. In-Depth Computational Analysis of Natural and Artificial Carbon Fixation Pathways. BioDesign Res. 2021, 1–23 (2021).
- Bar-Even, A., Noor, E., Lewis, N. E. & Milo, R. Design and analysis of synthetic carbon fixation pathways. Proc. Natl. Acad. Sci. 107, 8889–8894 (2010).
- Schwander, T., Schada von Borzyskowski, L., Burgener, S., Cortina, N. S. & Erb, T. J. A synthetic pathway for the fixation of carbon dioxide in vitro. Science 354, 900–904 (2016).
- Luo, S. et al. A cell-free self-replenishing CO2-fixing system. Nat. Catal. 5, 154–162 (2022).
- McLean, R. et al. Exploring alternative pathways for the in vitro establishment of the HOPAC cycle for synthetic CO2 fixation. Sci. Adv. 9, eadh4299 (2023).
- Pandi, A. et al. A versatile active learning workflow for optimization of genetic and metabolic networks. Nat. Commun. 13, 3876 (2022).
- Diehl, C., Gerlinger, P. D., Paczia, N. & Erb, T. J. Synthetic anaplerotic modules for the direct synthesis of complex molecules from CO2. Nat. Chem. Biol. 19, 168–175 (2023).
- Sundaram, S. et al. A Modular In Vitro Platform for the Production of Terpenes and Polyketides from CO2. Angew. Chem. Int. Ed Engl. 60, 16420–16425 (2021).
- Cruz-Morales, P. et al. Biosynthesis of polycyclopropanated high energy biofuels. Joule 6, 1590–1605 (2022).
- Lee, S. Y. et al. A comprehensive metabolic map for production of bio-based chemicals. Nat. Catal. 2, 18–33 (2019).
- Luo, S. et al. ATP production from electricity with a new-to-nature electrobiological module. Joule 7, 1745–1758 (2023).
- Ben-Nissan, R. et al. Autotrophic growth of E. coli is achieved by a small number of genetic changes. http://biorxiv.org/lookup/doi/10.1101/2023.06.03.543535 (2023) doi:10.1101/2023.06.03.543535.
Follow the Topic
-
Nature Catalysis
This journal brings together researchers from across all chemistry and related fields, publishing work on homogeneous catalysis, heterogeneous catalysis, and biocatalysts, incorporating both fundamental and applied studies.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in