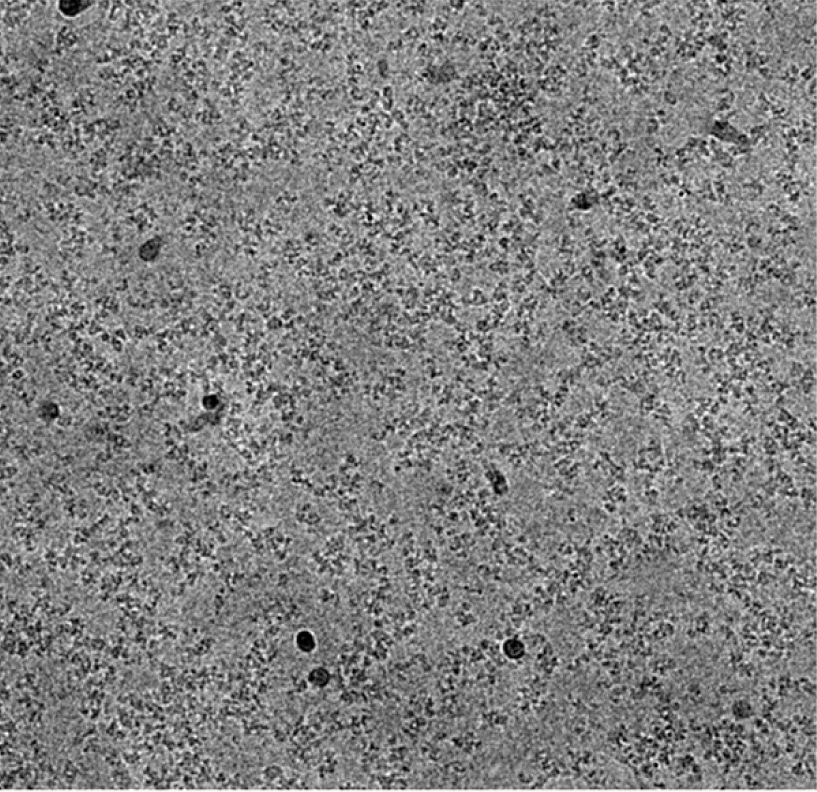

Retroviruses are ancient infectious elements that are ubiquitous in nature. Retroviruses had many evolutionary roles in different biological species by integrating their viral DNA, produced by reverse transcription of their RNA genome, into cellular chromosomes. The viral encoded integrase (IN) mediates the integration of the viral DNA.
As a microbiology student at the University of Iowa in 1970, I attended a virology class which sparked my interest in this field. This course initiated my career studying avian retroviruses at Saint Louis University, leading to the discovery of the avian myeloblastosis virus IN by my lab in 1978. By 1983, the retrovirus research field was primed to unravel the mysterious and harmful human immunodeficiency virus type 1 (HIV-1) that caused acquired immunodeficiency syndrome (AIDS).
Besides our biological approaches using Rous sarcoma virus (RSV), biochemistry played an important role in understanding IN functions. Numerous other labs studied IN in the vast retroviridae family including the genera lentivirus, betaretrovirus, and simiispumavirus among others. Near-atomic resolution structural studies of various IN/viral DNA complexes generally termed intasomes, including the presence of target DNA in these complexes, were sure to play an important role for understanding integration. We determined the structure of the RSV intasome containing viral DNA by single-particle cryo-EM (1). The intasome showed significant conformational flexibility in its overall architecture as well as the interactions of an HIV-1 IN strand transfer inhibitor (INSTI) in the active site. The availability of the state of the art cryo-EM facility at Washington University in St. Louis as well as our long collaboration with the structural biologist Dr. Hideki Aihara (University of Minnesota) made this project possible. Our work complements earlier pioneering x-ray crystallography studies of simiispumavirus prototype foamy virus (PFV) IN in complex with viral DNA, in the absence and presence of HIV-1 INSTIs (2). Structural studies of intasomes revealed different number of IN subunits ranging from 4 for the PFV and delta-retroviruses human T-cell leukemia virus type 1 (HTLV-1) (3) and simian T-lymphotropic virus type 1 (4), 8 for alpha-retrovirus RSV (5) and beta-retrovirus mouse mammary tumor virus (MMTV) (6). For lentiviral intasomes, variable number of IN subunits ranging from 4 to 12 for HIV-1 and simian immunodeficiency virus (SIV) (7-9) and up to 16 subunits for maedi visna virus (MVV) (10) were revealed. These studies provided a highly needed understanding of INSTIs interactions within the conserved intasome core (CIC) containing the active site. The CIC structure is maintained across all retrovirus intasomes.
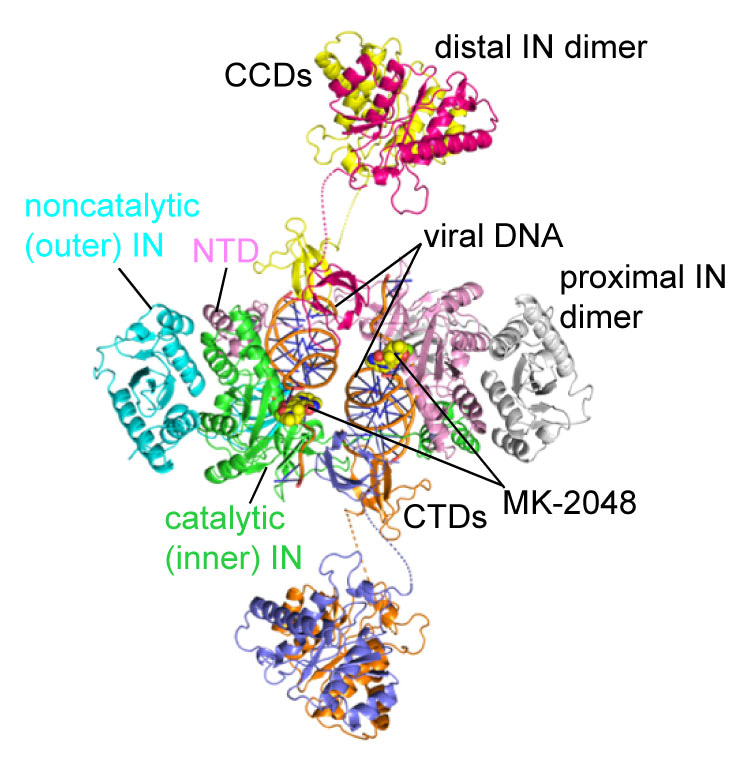
Fig. 1. Cryo-EM structure of the RSV octameric intasome trapped with MK-2048. Ribbon model representation of RSV octameric intasome viewed down its pseudo 2-fold axis. The two proximal IN dimers (green/cyan and pink/gray) plus the distal IN CTDs (magenta, slate blue) bridging between them constitute the core of the complex, referred as ‘conserved intasome core (CIC)’. Disordered CCD-CTD linkers for the distal IN dimers are shown as dashed lines.
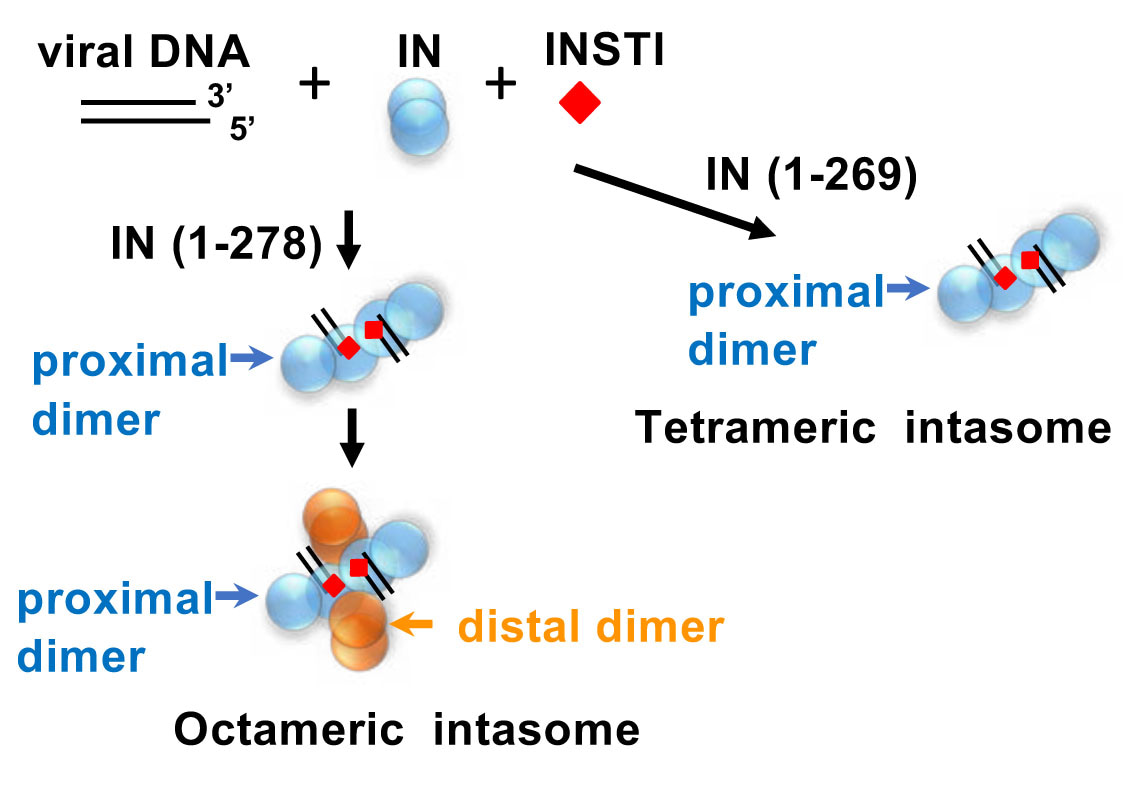
Fig. 2. Assembly pathway of RSV octameric and tetrameric intasomes. Octameric intasome can be assembled with IN 1-278 while a tetrameric intasome is formed with IN 1-269. INSTI stabilizes the intasomes.
What are the basic assembly mechanisms associated with these architecturally diverse IN intasome structures that carry out integration? We approached this question using the RSV octameric intasome containing 8 IN subunits with viral DNA as a model (1) (Fig. 1). Earlier studies demonstrated that a tetrameric RSV intasome, the precursor to the octameric form, was preferentially assembled by a C-terminal truncated RSV IN (1-269) that lacks the last 17 amino acids (11,12) (Fig. 2). We utilized HIV-1 INSTIs as “effective tools” to stabilize the tetrameric intasome, a necessary step for isolation by size-exclusion chromatography and, future structural determination by cryo-EM. RSV octameric intasome is produced and isolated in a similar fashion with IN 1-278, the optimum size truncation for its assembly in comparison to wt IN 1-286. Single-particle cryo-EM analysis produced surprising results. It revealed significant flexibility of the two non-catalytic distal IN dimers along with previously unrecognized CIC movement, suggesting ordered conformational transitions between intermediates that may be important to capture the target DNA. In collaboration with Dr. Alan Engelman (Harvard Medical School), we investigated how C-terminal IN truncations and structure-based missense mutations affected assembly in vitro and infectivity of pseudotyped RSV virions. Truncations of the C-terminal region up to Ile269 did not affect viral infectivity despite regulating the transition of the tetrameric intasome to the octameric form in vitro. Missense mutations helped identify critical residues that allowed the identification of amino acids affecting the assembly of either the tetrameric or octameric intasomes in vitro and under physiological conditions. Hopefully, these studies will provide further insights into investigating the assembly of different retrovirus intasomes including HIV-1.
References
- Pandey, K. K., Bera, S., Shi, K., Rau, M. J., Oleru, A. V., Fitzpatrick, J. A. J., Engelman, A. N., Aihara, H., and Grandgenett, D. P. Cryo-EM structure of the Rous sarcoma virus octameric cleaved synaptic complex intasome. Commun Biol 4, 330 (2021). DOI 10.1038/s42003-021-01855-2.
- Hare, S., Gupta, S. S., Valkov, E., Engelman, A., and Cherepanov, P. Retroviral intasome assembly and inhibition of DNA strand transfer. Nature 464, 232-236 (2010).
- Bhatt, V., Shi, K., Salamango, D. J., Moeller, N. H., Pandey, K. K., Bera, S., Bohl, T. E., Kurniawan, F., Orellana, K., Zhang, W., Grandgenett, D. P., Harris, R. S., Sundborger-Lunna, A. C., and Aihara, H. Structural basis of host protein hijacking in human T-cell leukemia virus integration. Nat Commun 11, 3121 (2020).
- Barski, M. S., Minnell, J. J., Hodakova, Z., Pye, V. E., Nans, A., Cherepanov, P., and Maertens, G. N. Cryo-EM structure of the deltaretroviral intasome in complex with the PP2A regulatory subunit B56gamma. Nat Commun 11, 5043 (2020).
- Yin, Z., Shi, K., Banerjee, S., Pandey, K. K., Bera, S., Grandgenett, D. P., and Aihara, H. Crystal structure of the Rous sarcoma virus intasome. Nature 530, 362-366 (2016).
- Ballandras-Colas, A., Brown, M., Cook, N. J., Dewdney, T. G., Demeler, B., Cherepanov, P., Lyumkis, D., and Engelman, A. N. Cryo-EM reveals a novel octameric integrase structure for betaretroviral intasome function. Nature 530, 358-361 (2016).
- Passos, D. O., Li, M., Yang, R., Rebensburg, S. V., Ghirlando, R., Jeon, Y., Shkriabai, N., Kvaratskhelia, M., Craigie, R., and Lyumkis, D. Cryo-EM structures and atomic model of the HIV-1 strand transfer complex intasome. Science 355, 89-92 (2017).
- Passos, D. O., Li, M., Jozwik, I. K., Zhao, X. Z., Santos-Martins, D., Yang, R., Smith, S. J., Jeon, Y., Forli, S., Hughes, S. H., Burke, T. R., Jr., Craigie, R., and Lyumkis, D. Structural basis for strand-transfer inhibitor binding to HIV intasomes. Science 367, 810-814 (2020).
- Cook, N. J., Li, W., Berta, D., Badaoui, M., Ballandras-Colas, A., Nans, A., Kotecha, A., Rosta, E., Engelman, A. N., and Cherepanov, P. Structural basis of second-generation HIV integrase inhibitor action and viral resistance. Science 367, 806-810 (2020).
- Ballandras-Colas, A., Maskell, D. P., Serrao, E., Locke, J., Swuec, P., Jonsson, S. R., Kotecha, A., Cook, N. J., Pye, V. E., Taylor, I. A., Andresdottir, V., Engelman, A. N., Costa, A., and Cherepanov, P. A supramolecular assembly mediates lentiviral DNA integration. Science 355, 93-95 (2017).
- Pandey, K. K., Bera, S., Korolev, S., Campbell, M., Yin, Z., Aihara, H., and Grandgenett, D. P. Rous sarcoma virus synaptic complex capable of concerted integration is kinetically trapped by human immunodeficiency virus integrase strand transfer inhibitors. J Biol Chem 289, 19648-19658 (2014).
- Bera, S., Pandey, K. K., Aihara, H., and Grandgenett, D. P. Differential assembly of Rous sarcoma virus tetrameric and octameric intasomes is regulated by the C-terminal domain and tail region of integrase. J Biol Chem 293, 16440-16452 (2018).
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
DNA repair and human disease
Publishing Model: Hybrid
Deadline: Oct 31, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in