
Antimicrobial resistance (AMR) is a major problem for public health. Antimicrobial resistant bacteria produce diseases difficult to treat, raising the mortality rates and economic costs associated with infections. Since the introduction of antibiotics in the clinical practice, the emergence and progressive dissemination of AMR mechanisms among bacteria have increased. In this scenario, conjugative plasmids are important drivers of AMR dissemination by allowing resistance genes to rapidly spread between bacterial strains. However, plasmid carriage usually imposes a burden on bacterial physiology, which translates into a reduction in bacterial fitness. This is especially relevant in the absence of selection for plasmid-encoded genes, and therefore it is difficult to explain the high plasmid prevalence in bacteria, a puzzle known as “the plasmid paradox”. In the recent years, several studies have characterized the ecological and evolutionary solutions that explain the widespread of plasmids in bacterial populations. These studies have shown that in laboratory settings one of the most important mechanisms of plasmid stabilization are compensatory mutations that ameliorate the plasmid-associated fitness costs. However, the impact of these mutations in the real world is still unexplored.
When I joined the Plasmid Biology and Evolution lab as a young and enthusiastic PhD student, I was proposed to collaborate in a huge project in which our main objective was to understand the evolutionary dynamics of plasmid-bacteria associations in the gut of hospitalised patients. The gut of hospitalised patients is of particular interest as it is believed to be one of the main hotspots of AMR exchange. But are compensatory evolution events shaping AMR evolution in a complex environment such as the human gut? To test our hypothesis, we focused on a widespread conjugative AMR-plasmid from a collection of enterobacteria isolated from hospitalised patients. We had previously described how plasmid-mediated antimicrobial resistance was transmitted between and within patients in the hospital. A key finding of this study was that once a plasmid-carrying clone arrives to the gut of a patient, the plasmid is able to spread horizontally to other members of the gut community. This, creates the perfect conditions for the apparition (and selection) of plasmid compensatory mutations as theory predicts that plasmid costs could be higher in recent events of plasmid transmission. Additionally, patients included in the study were sampled periodically, generating timelines of isolates which should theoretically allow us to investigate the in vivo evolution of plasmid-bacteria associations in the human gut.
Each patient was sampled periodically, generating timelines of isolates
Our project was as exciting as it was challenging. The study involved the manipulation of different species of clinical multi-drug resistant bacteria, in which “standardized things” such as transformation or plasmid-curing are not optimized and can become troublesome. For example, most isolates were resistant to almost all antibiotics and we had to find a proper selective marker for specifically manipulate each organism. A particularly tough case, which I will someday explain to my grandchildren, was the genetic manipulation of a plasmid which did not confer any antibiotic resistance (PV-A in the manuscript).
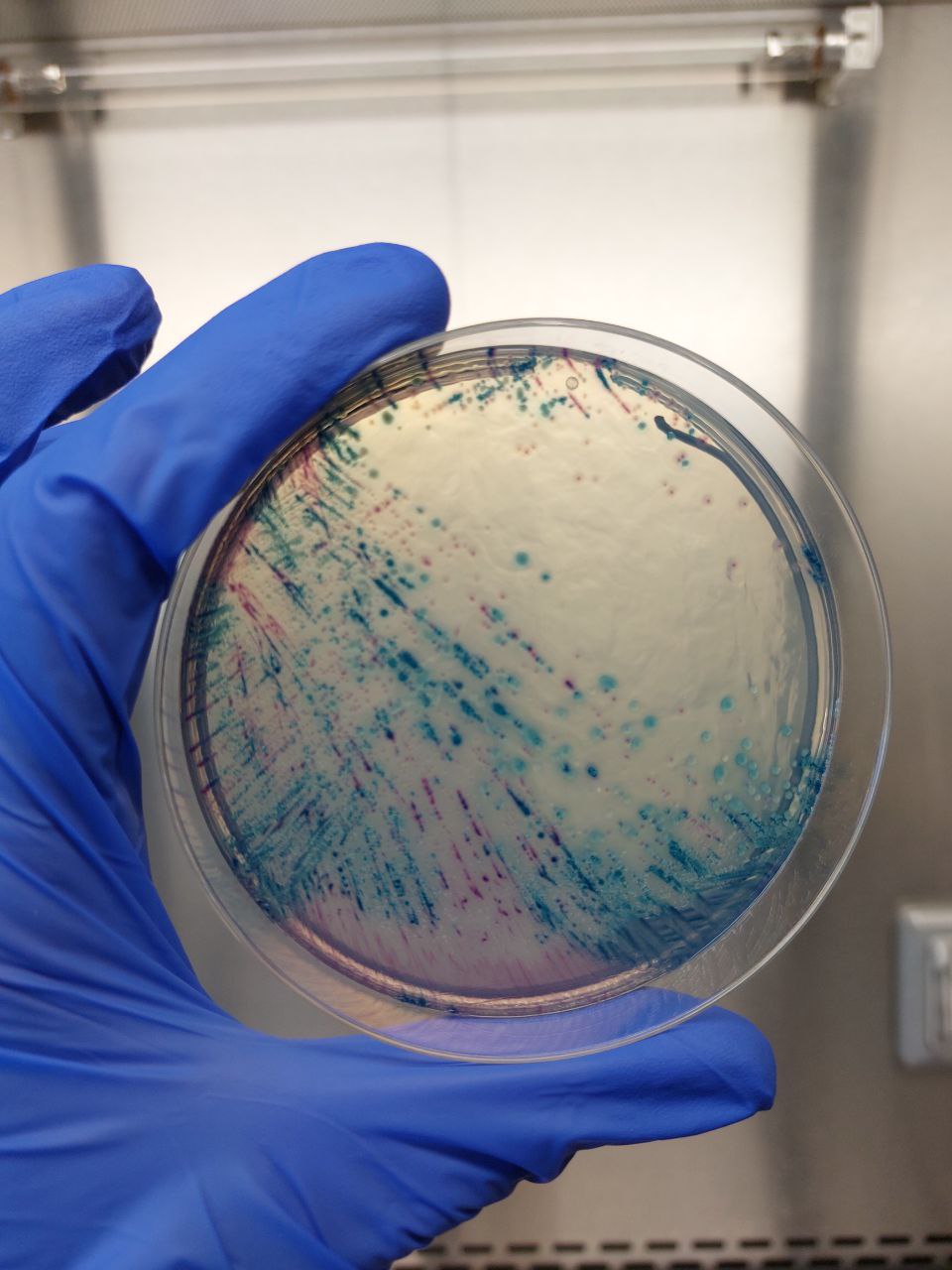
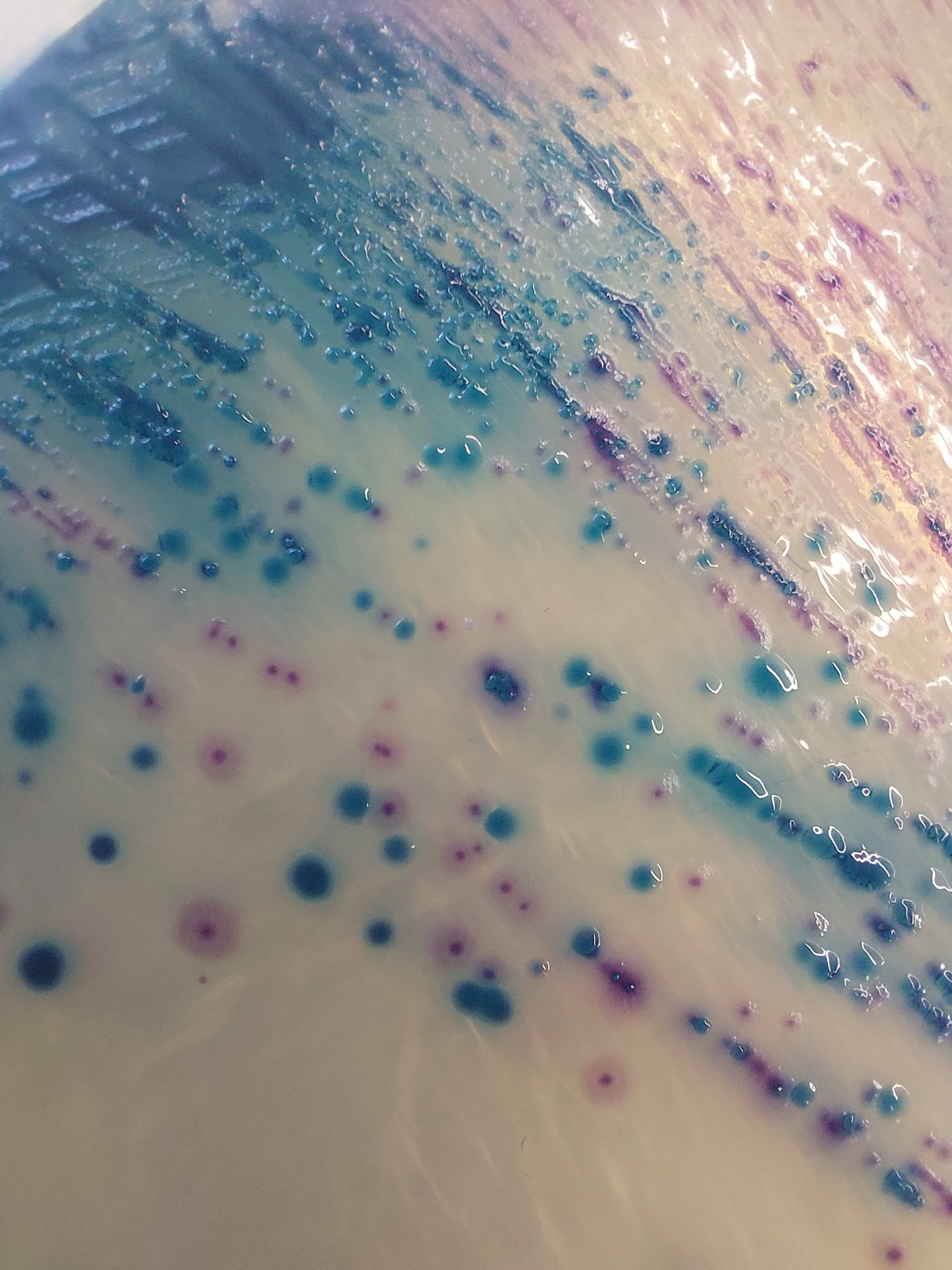
A differential medium agar plate containing multi-drug resistant Escherichia coli (red) and Klebsiella pneumoniae (blue) isolated from a single hospitalized patient.
But our efforts were worth it. First, we identified a bunch of different plasmid variants carrying a wide-range of mutations, suggesting plasmid-evolution. Then, we parallelly introduced a subset of these variants in the same genetic background and we observed that they were actually conferring different phenotypes. Importantly, when we analysed the phenotype data of these plasmids and we unveiled a fitness-resistance trade-off, that could be contributing to the evolution of pOXA-48-mediated AMR in vivo. Finally, we identified cases of within patient plasmid evolution. By combining patients’ metadata, enterobacterial sequencing data and by successfully managing to cure pOXA-48 in wild-type multidrug resistant enterobacteria, we spotted several events of compensatory evolution events in plasmids and we confirmed that bacteria evolving within the gut of the patients are constrained by a trade-off which may be modulated by the different antibiotic treatments. Importantly, data also strongly suggest that bacterial clones were coexisting in the gut at the same time. This, together with high horizontal transfer rates, may be key factors allowing these bacteria to overpass the constraints imposed by the fitness-resistance trade-off.
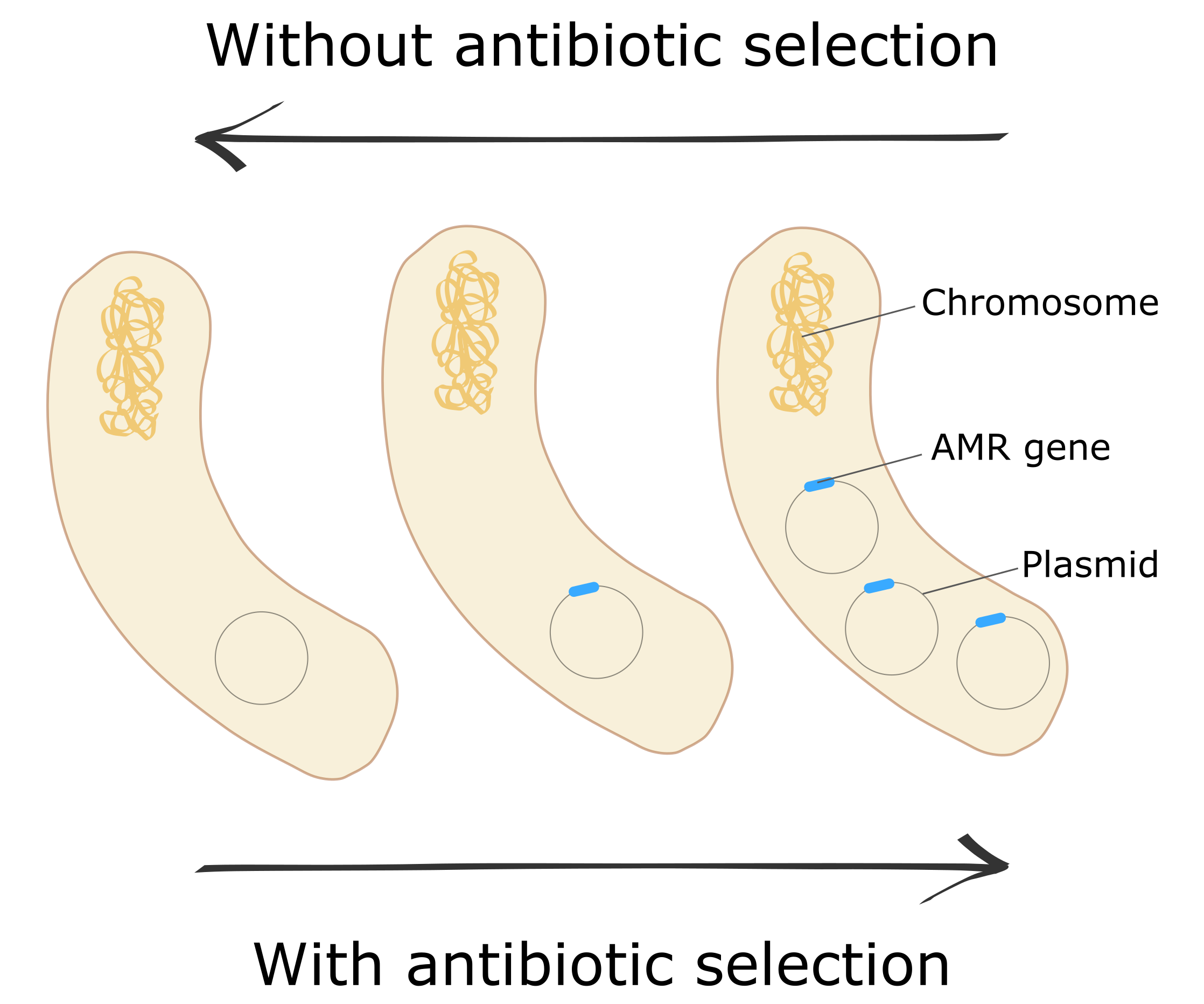
A trade-off governs the evolutionary trajectory of plasmids in the clinic.
From a broader perspective our results go in line with other studies and highlight the necessity of considering the genetic diversity present in the patients' microbiome to fully understand the ecology and evolution of AMR in bacteria.
Find also a nice News&Views post about this article here.
Follow the Topic
-
Nature Ecology & Evolution
This journal is interested in the full spectrum of ecological and evolutionary biology, encompassing approaches at the molecular, organismal, population, community and ecosystem levels, as well as relevant parts of the social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Biodiversity and ecosystem functioning of global peatlands
Publishing Model: Hybrid
Deadline: Jul 27, 2026
Understanding species redistributions under global climate change
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in