Growth phase estimation using human stool metagenomic time series data
Published in Bioengineering & Biotechnology
Longitudinal sampling of stool has provided important insights into the ecological dynamics of the human gut 1,2. There has been increasing interest in long-term monitoring of the gut microbiome for clinical trials, enabled by technologies like smart toilets 3,4. However, the interpretation of these dynamics has been fraught due to an under-appreciated disconnect between population dynamics in the gut (that occur on minute-to-hour timescales) and sampling timescales of human stool (around once per day).
Intuitively, if abundance fluctuations in gut microbial time series data were proportional to the growth or death of microbial populations, we would expect that these deltas (tn+1 – tn) at each sampling interval would be positively correlated to the baseline abundance (tn) in the case of net growth and negatively correlated in cases of net death. However, we observed these relationships to be uniformly negative across all taxa, across all gut microbiome time series sampled from healthy adults, which is most consistent with regression-to-the-mean (i.e., randomly sampling from a stationary distribution). Overall, we contend that population growth/death processes cannot be accurately inferred from day-to-day fluctuations in steady-state abundances in human stool.
We next sought to find alternative ways to infer bacterial population dynamics from metagenomic time series. To tackle this problem, we calculated instantaneous microbial replication rates (i.e., peak-to-trough ratios, or PTRs) for abundant taxa in gut microbiome time series 5, which we paired with abundance time series. We used log2(PTR)s as proxies for effective bacterial population growth rates. For the same gut bacterial taxon, we observed a range of positive, negative, and null associations between log2(PTR) and centered log ratio (CLR) abundance across donors, which was not consistent with regression-to-the-mean. These patterns were complex, and required a more sophisticated interpretation.
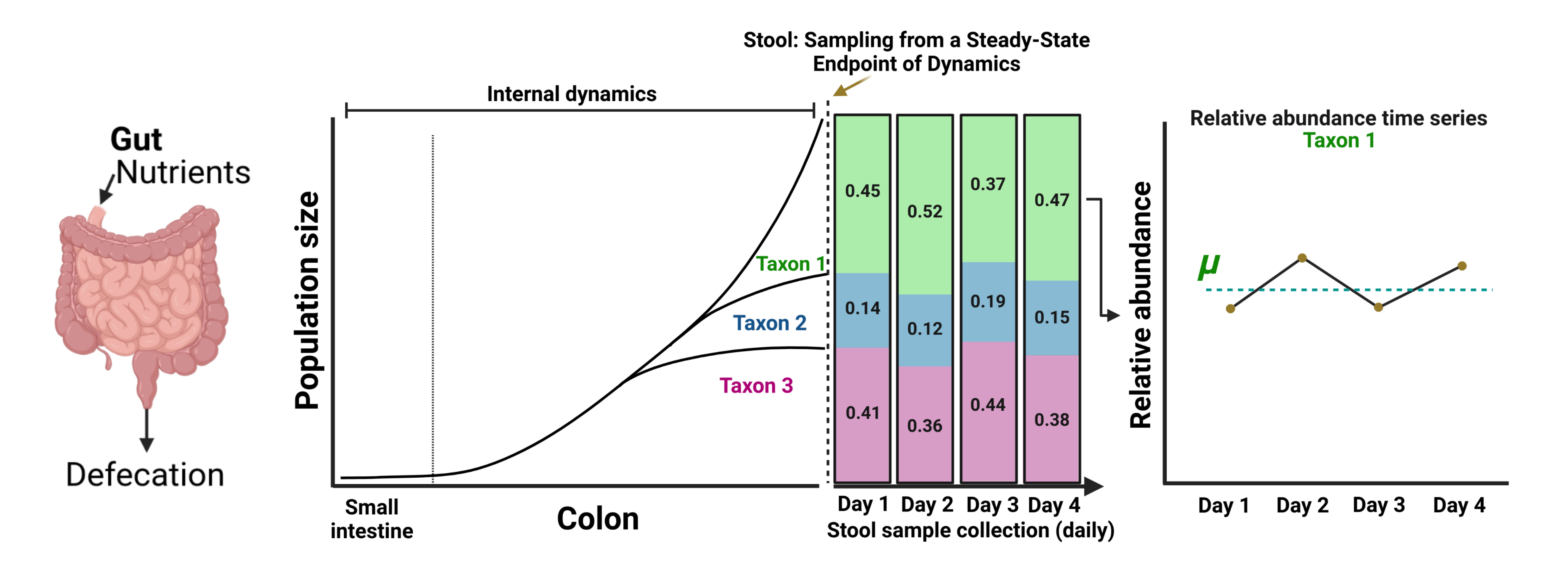
We took a modeling approach to better understand and interpret the observed relationships between log2(PTR) and abundance. The logistic growth equation (LGE) is a classic model of self-limiting growth, resulting in a sigmoidal curve. The first derivative of this curve with respect to time yields the change in abundance (i.e., the effective growth rate). We introduced a stochastic term to the LGE to account for the technical and biological noise we expect to see in empirical data (sLGE). We defined 4 growth phases (i.e., acceleration, mid-log, deceleration, and stationary) depicting major regions in the growth curve. Upon simulation, we observed that the correlation between the first derivative (effective growth rate) and population abundance curves were positive in acceleration phase and negative in deceleration phase, while growth rates and abundances were not correlated in the mid-log and stationary phases. We leveraged these observations to identify putative growth phases for gut bacterial populations, sampled at a regular frequency over many time points.
We assessed our model predictions using replicate E. coli populations sampled at high temporal resolution across their growth curves. We performed metagenomic shotgun sequencing, obtaining log2(PTR) and abundance values for E. coli. We found that the relationships between log2(PTR)s and abundances were significantly positively and negatively correlated in acceleration and deceleration phases, respectively, matching our sLGE findings. Furthermore, we found no statistical significance between log2(PTR) and abundance in the E. coli samples in mid-log or stationary phases, as expected, but we observed that samples in mid-log phase had a mean log2(PTR) ~3.5-fold greater than those in stationary phase. This difference clearly distinguished between mid-log and stationary phases across multiple different gut bacterial species grown in vitro. Overall, this result allowed us to propose a lower PTR threshold that could be leveraged to identify stationary phase (i.e., dormant, non-growing) taxa 5, which further refined our ability to identify in situ growth phases.
Finally, we used our novel growth phase inference approach to assign putative in situ growth phases to abundant commensals present across four metagenomic time series from healthy human donors. For taxa with average log2(PTRs) above the empirical stationary phase threshold, significantly positive associations between log2(PTRs) and CLR abundances likely indicate acceleration phase and significantly negative associations likely indicate deceleration phase. Furthermore, we were able to distinguish putative mid-log and stationary phases using our PTR threshold. Taxa that showed no log2(PTRs)/CLR correlation and had PTR values above the stationarity threshold were strong candidates for mid-log phase, but these taxa might also be in acceleration or deceleration phase (i.e., we were underpowered to detect the correlation). Therefore, these taxa were labeled as ‘non-stationary’.
Our findings provide a new path towards improving statistical analyses and mechanistic modeling of human fecal microbiome time series data. In addition, this work could inform more accurate genome-scale metabolic modeling of gut microbiome communities, which assume that all organisms are growing exponentially at steady-state (e.g., community models could exclude taxa that are likely in stationary phase, as they are probably not contributing that much to colonic metabolism). We hope our in situ growth phase estimation approach will be useful for advancing microbiome research across a range of domains, from human health to agriculture and industrial bioreactor design.
References
- Hou, K. et al. Microbiota in health and diseases. Signal Transduct Target Ther 7, 135 (2022).
- Fan, Y. & Pedersen, O. Gut microbiota in human metabolic health and disease. Nat. Rev. Microbiol. 19, 55–71 (2021).
- Park, S.-M. et al. A mountable toilet system for personalized health monitoring via the analysis of excreta. Nat Biomed Eng 4, 624–635 (2020).
- Ge, T. J. et al. Passive monitoring by smart toilets for precision health. Sci. Transl. Med. 15, eabk3489 (2023).
- Korem, T. et al. Growth dynamics of gut microbiota in health and disease inferred from single metagenomic samples. Science 349, 1101–1106 (2015).
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in