Hematopoietic stem cells use chromatin-remodeling factors for specific processes: the story of Smarca5.
Published in Cell & Molecular Biology and Immunology

The production of blood cells relies on coordinated regulation of gene expression, timing of the cell cycle, and suppression of inappropriate processes by apoptosis. Like any stem cell, the hematopoietic stem cell deals with the balance between stemness and differentiation, but also with the quantitative requirements for the production of different elements at specific stages of development and during everyday situations that might include bleeding or infections. The different decisions in the cell nucleus of stem cells are related to the structure of the DNA, which is wrapped in a bun of histone proteins. Different critical sequences have to be made available, and this is done by an enzymatic motor that releases DNA from the histone octamer, called the nucleosome, with the expenditure of energy, creating a region of freely accessible DNA, while other unnecessary DNA sequences are twisted like a thread around a spool. The unwinding of the DNA strand from the nucleosome is called the process of chromatin remodeling. It is generally accepted that stem cells have very little gene expression and replication, and thus we were interested in how dependent the various blood subtypes are on the function of the chromatin remodeling enzymatic engine in stem cells during their early differentiation. Specifically, we tested how the ISWI ATPase SMARCA5 (SNF2H), which represents an absolutely essential and unique enzyme, effects the production of lymphoid, erythroid and myeloid cells.
This type of project involves mouse transgenic models that we have gradually developed over the past 20 years. My lab group took up the project around 2005 in Prague, Czech Republic, but the roots go back to 1998 when we cloned the mouse mRNA for Smarca5 and found that Smarca5 is very important for blood production. In 2003, we published with Art Skoultchi (Albert Einstein College of Medicine, Bronx, NY) a mouse knockout for Smarca5 that showed its indispensability for early embryonic development. Further research led to the development of a mouse model of Smarca5 conditional inactivation that demonstrated the importance of Smarca5 for hematopoietic stem cell, lymphoid and erythroid development.
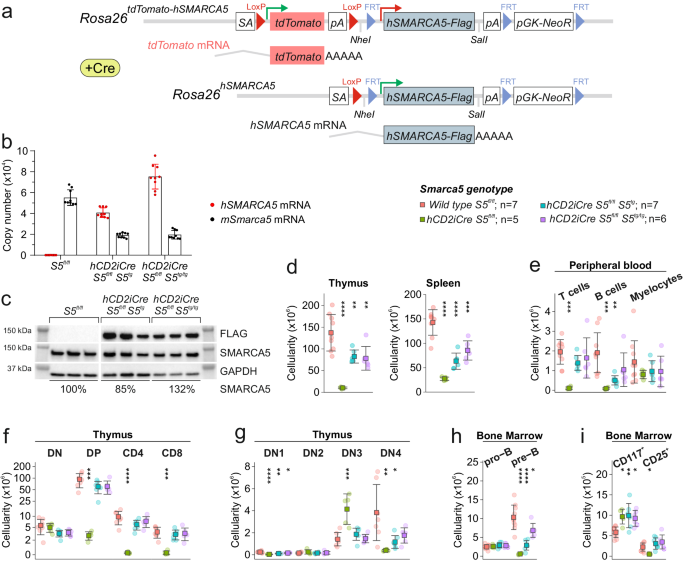
When we used up the possibilities of the knockout mouse and decided to create a conditional transgenic model for Smarca5. Although we created models for human as well as murine Smarca5, a publication now coming out in Communications Biology covers the first results with the transgene for human SMARCA5 entitled: Differential requirements for Smarca5 expression during hematopoietic stem cell commitment. We observed that the transgenic allele only provides about 10% of the expression of the normal allele. Therefore, we crossed the transgenic model with the previous model with a knockout allele to generate a set of supermodels in which the different Smarca5 expression levels exist. This allowed us to show that the hypomorphic (producing lower protein levels) SMARCA5 allele (S5tg) is fully functional and can rescue, in a dose-dependent manner, the gene inactivation phenotypes from previous k.o. projects, i.e. defects in lymphoid and erythroid development. In addition, transgenic Smarca5 binds all canonical complexes that have been incrementally discovered. Once we express low level of Smarca5 in stem cells, we see that progenitors entering lymphoid T and B development are the most sensitive to Smarca5 reduction, while other progenitors are less or not affected (see figure & legend).
To support our hypothesis that SMARCA5 expression level has a direct effect on development at stem cell level, we employed competitive transplantation experiment of transgenic and normal stem cells into normal but also transgenic recipients. Both of these experiments confirmed our hypothesis that it is primarily lymphocyte development that is strongly dependent on Smarca5 levels and that the defect originates in stem cells. Low SMARCA5 levels lead to significant permissiveness of the bone marrow niche for wild type stem cells. Transgenic hematopoietic stem cells were not able to produce lymphocytes, indicating that these lymphoid early progenitors are most sensitive to SMARCA5 levels. You think that's the end of the story? But no. Our current results show that Smarca5 is indeed a very important factor that affects the life of cancer stem cells. Therefore, we are also focusing on the question of how to effectively get rid of these cancer stem cells and one of the mechanisms is to focus on the biology of Smarca5 in this regard.

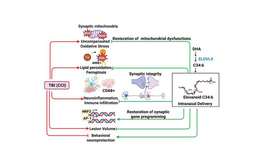
Figure legend: Hematopoietic stem cells in bone marrow regulate the expression of the epigenetic regulator Smarca5, which influences the development of blood elements through ATP-dependent effects on nucleosome architecture. Our data show that the development of lymphoid progenitors of both the B and T lineages is most dependent on Smarca5 levels, erythropoiesis is less dependent, while a very low level of the studied ISWI ATPase is sufficient for myeloid cell development. Created with BioRender.com.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
From RNA Detection to Molecular Mechanisms
Publishing Model: Open Access
Deadline: May 05, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in