Looking for cancer in the bloodstream – one patient at a time
Published in Research Data and Biomedical Research
From a clear goal to a complicated reality
The project began in 2018 with what seemed like a straightforward idea: to map cancer-related changes in blood plasma cell-free RNA (cfRNA) across different cancer types. At the time, circulating DNA was already well established as a source of cancer biomarkers, and we wanted to explore whether cfRNA could provide complementary information given that it can reflect gene activity across the body.
But the data had other ideas.
As we began analyzing the first datasets, one pattern became immediately clear: variability. For blood cancers, the signal was relatively strong and consistent. But for solid tumors the differences between cancer patients and healthy individuals were far less uniform. Each patient seemed to show their own unique pattern of changes.
At first, this felt like a problem. Much of biomarker research relies on finding signals that behave consistently across large groups of patients. Our results didn’t fit that model. The more samples we analyzed, the clearer it became that a “one-size-fits-all” biomarker might simply not exist for cfRNA in solid tumors.
What also became clear over time was that the project itself was evolving. What started as a relatively small cohort grew into a dataset of more than 600 plasma samples across multiple cancer types. With each expansion came new questions and additional layers of analysis, making it increasingly challenging to bring everything together into a coherent story.
Turning variability into a signal
The turning point came when we stopped trying to reduce variability and instead asked whether we could use it.
Rather than searching for genes that are consistently different on average between patients and controls, we shifted to a patient-centered perspective: what stands out in an individual sample? For each gene, we compared its abundance in a patient’s plasma to the distribution observed in healthy individuals. Genes that fell far outside the typical range - either much higher or much lower - were labeled as “tail genes.”
This idea is conceptually simple, but it changed how we looked at the data.
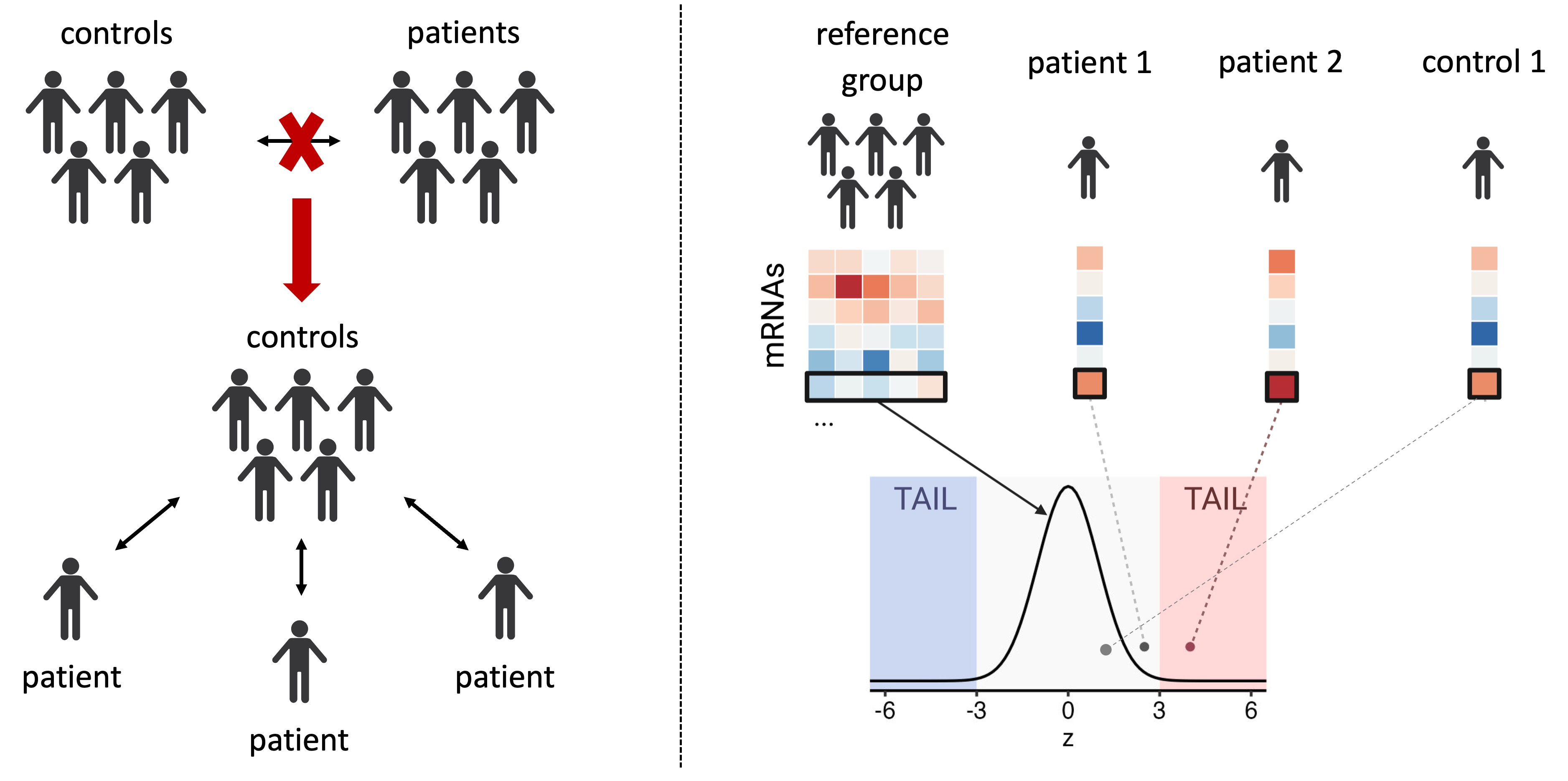
For each gene, we compare its abundance in a patient’s sample to the range observed in healthy individuals. If a gene’s transcript levels are much higher or lower than expected (far outside the typical range) it is classified as a “tail gene” for that patient. Cancer patient samples generally have more of these strongly deviating genes than those from healthy individuals, and this difference can be used to distinguish them. Figure created with BioRender.com.
When we applied this approach, a clear pattern emerged: cancer patients consistently had more of these strongly deviating genes than healthy individuals. Remarkably, simply counting the number of “tail genes” in a sample was enough to distinguish cancer patients from controls with good accuracy. This held true across independent datasets and even extended beyond blood plasma to urine cfRNA profiles.
What surprised us most was not just that this worked, but how robust such a simple measure turned out to be. While the exact genes differed from patient to patient, there was still partial overlap, suggesting a mix of shared biological signals and individual variation.
A long journey - inside and outside the lab
Research rarely follows a straight line, and neither does life. This project evolved over the course of my PhD, shaped both by the work itself and by what was happening outside the lab. There were periods when I had to step back, and others when the project demanded full attention - with multiple rounds of revision and a dataset that kept growing faster than I could keep up with. Those shifts in pace became part of the process: sometimes frustrating, sometimes motivating, but always there in the background. Looking back, I’m proud not just of the final result, but of having seen it through.
Looking beyond the average
This study changed how we think about variability in biological data. What initially appeared as noise turned out to be a signal in its own right - one that becomes visible when we stop averaging across individuals and start looking more closely at each patient.
Sometimes, the most informative patterns are not the ones that everyone shares, but the ones that make each patient unique.
As the field moves toward more personalized approaches to diagnosis and treatment, embracing this kind of individual variation may become increasingly important. While more work is needed before clinical application, our results suggest that cfRNA, and patient-specific deviations within it, could play a valuable role in future cancer diagnostics.
This work was carried out as part of the OncoRNALab, and would not have been possible without the many collaborators who contributed samples, expertise, and critical feedback along the way. We are also very grateful for the funding that supported this project.
Follow the Topic
-
Communications Medicine
A selective open access journal from Nature Portfolio publishing high-quality research, reviews and commentary across all clinical, translational, and public health research fields.
Related Collections
With Collections, you can get published faster and increase your visibility.
Public health and health governance in China
Publishing Model: Open Access
Deadline: Jul 31, 2026
Life Course Epidemiology
Publishing Model: Open Access
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in