Mapping T cell activation one cell at a time
Published in Bioengineering & Biotechnology, Genetics & Genomics, and Biomedical Research

Explore the Research
Just a moment...
onlinelibrary.wiley.com
Classical three-signal model of T cell activation
The classical three-signal model of naïve T cell activation, T cell receptor (TCR) engagement, co-stimulation, and cytokine support, has long provided a clear, linear pathway for understanding adaptive immunity. Yet, as immunological understanding has grown and, in the last decade, technology has exponentially advanced, it has become increasingly evident that this model captures only part of a far more complex reality. T cell activation is simply not a singular, uniform event but a dynamic spatio-temporal process shaped by exquisite intra- and inter-cellular cooperation. For the journal Immunology and Cell Biology (ICB) we recently highlighted four influential studies published in 2025 illustrating how emerging technologies, particularly single-cell genomics, are redefining the foundational T cell activation paradigm.
A defining feature of the immune system is its heterogeneity, the ability to generate diverse and highly specialised cell states in response to varied challenges. Historically, this diversity has been difficult to resolve. Techniques such as flow cytometry and bulk RNA sequencing have enabled the classification of major T cell subsets, but they rely on predefined markers and signals averaged across populations. As a result, rare, transient, or intermediate cellular states, required for understanding how immune responses develop, remained elusive.
Single cell genomics is redefining cellular immunology
Single-cell genomic approaches such as single-cell RNA sequencing (scRNA-seq), when combined with TCR sequencing, trajectory inference, and multimodal profiling, now allow for the unbiased analysis of individual cells. These methods can intricately reconstruct differentiation pathways, extensively map clonal relationships, and have the potential to reveal the full spectrum of cellular states within an immune response. Importantly, they also enable the integration of temporal and spatial dimensions, offering a far more complete picture of how T cells are activated, maintained, and adapted across the body.
Spatio-temporal dynamics and cellular communication networks
One of the most significant emerging insights is that CD8+ T cell activation unfolds as a temporally staged process. Rather than a single priming event, activation involves multiple waves of signalling. A key example is the identification of a secondary priming phase in lymph nodes, occurring days after the initial antigen encounter. During this phase, CD4+ T helper cells provide critical signals, most notably IL-2, that selectively reinforce high-affinity CD8+ T cell clones. This acts as a developmental checkpoint, refining clonal selection and shaping the trajectory of effector differentiation. Such fine-grained temporal dynamics have only become apparent through single-cell approaches capable of resolving changes at the level of individual cells.
Equally important and reliant on advanced single-cell genomics technology, is the recognition that lymph nodes function as more than sites of initial activation. Instead, they act as sustained developmental hubs that maintain stem-like progenitor populations. Single-cell trajectory analyses have identified these progenitor cells (TPEX cells) as highly plastic, with the capacity to generate multiple downstream fates, including effector and exhausted states. Their presence ensures a sustained immune responses during chronic infection and cancer, highlighting the importance of maintaining a reservoir of adaptable cells capable of responding to ongoing antigen exposure.
Beyond lymphoid tissues, single-cell genomics has revealed a complex network linking lymph nodes and peripheral sites. Indeed recent clonal tracking studies demonstrate that tissue-infiltrating CD8+ T cells are frequently derived from progenitor populations maintained in lymph nodes, indicating continuous seeding across anatomical compartments. At the same time, local tissue environments exert powerful influences on T-cell fate. Signals such as TGFβ can promote tissue residency while limiting progenitor expansion, effectively partitioning immune responses across different niches. Notably, hybrid cellular states have been identified that combine features of stem-like and tissue-resident populations, underscoring the remarkable plasticity of T-cell differentiation.
Peripheral tissues themselves are increasingly recognised as active participants in immune regulation. Rather than serving solely as sites of effector function, they can provide instructive signals that reshape T-cell responses. Noncanonical helper circuits, involving interactions between CD4+ T cells and tissue-resident cells such as macrophages, can restore CD8+ T-cell function in otherwise suppressive environments. These findings highlight a distributed system of immune regulation, in which multiple anatomical sites contribute distinct yet coordinated signals.
Cellular network maps to navigate therapeutic advances
Despite major advances, current single-cell approaches remain limited by their inability to fully preserve spatial context, though emerging technologies such as spatial transcriptomics, multiplexed imaging, and lineage tracing are beginning to bridge this gap by integrating molecular, spatial, and temporal dimensions of immune responses. As these tools converge, they are shifting the field from static snapshots toward dynamic, in situ maps of T-cell activation, migration, and differentiation. In the coming years, such integrated single-cell frameworks may not only reveal how immune activation unfolds across tissues and time, but also enable its precise modulation, opening the door to therapies that can steer immune responses with unprecedented accuracy and durability.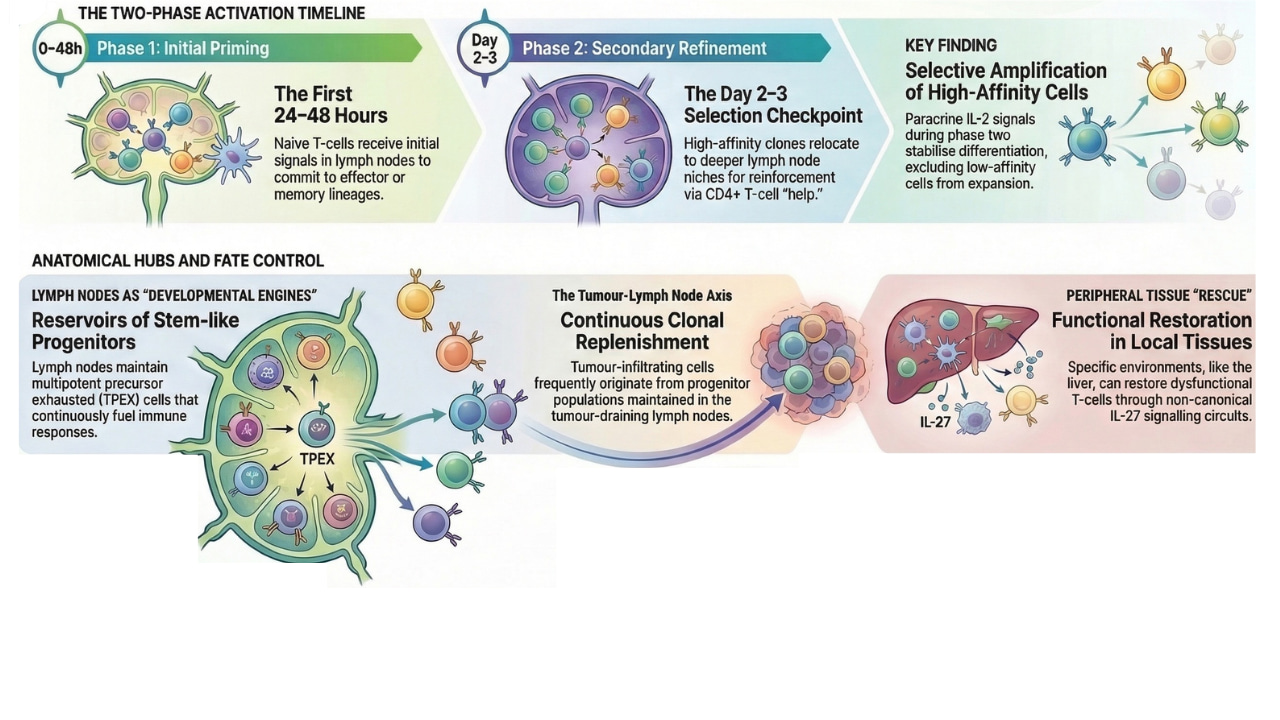
Follow the Topic
Ask the Editor - Immunology, Pathogenesis, Inflammation and Innate Immunity
Got a question for the editor about the complement system in health and disease? Ask it here!
Continue reading announcement

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in