Mosaic variant barcode analysis to turn back the clock and understand human development
Published in Cell & Molecular Biology
Tracing cellular lineages has been a fundamental question in understanding embryonic development. Over the past decades, if not centuries, numerous methodologies, including direct observation, cell intrinsic pigmentation, injected paints or dyes, viral tracing, isotope labeling, genetic recombination, fluorescent labeling, and the recent CRISPR-based tracing techniques, have been established to facilitate the study of embryonic development at resolution from whole organism all the way to every single cell. Although these techniques work really well for model organisms, it has been exceptionally hard to use traditional interventions to perform these tracings in humans, for many complicated reasons.
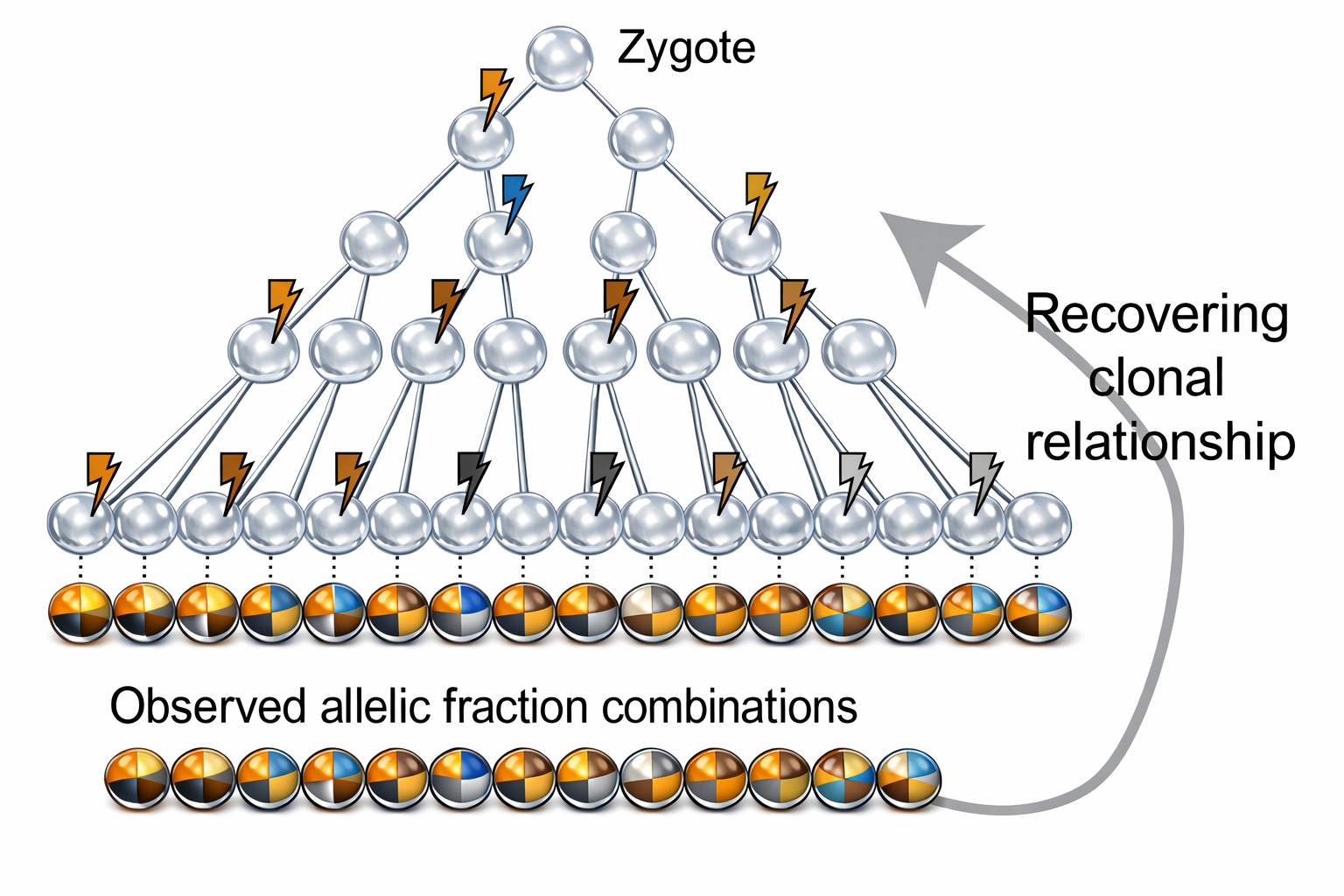
The idea of lineage tracing in humans with mutations first came about in cancer studies for clonal evolution, where acquired genetic instability permits stepwise selection of variant sublines and underlies tumor progression. Different from tumor driver variants that are selected with high allelic fractions, naturally occurring noncancer mosaic variants come in significantly lower fractions and are hard to distinguish from noise. With the recent dramatic price drop of whole-genome sequencing and AI-powered ultra-specific variant calling algorithms, we can accurately detect and quantify fractions of mosaic variants happening across the genome, which are likely neutral barcodes, inherited by daughter cells, at unprecedented resolution.
Based on the observation that different organ, tissue, and even cell from the same individual all possess their own mutations, and these postzygotic mutations get inherited by daughter cells and result in patchy genomic compositions called mosaicism. The idea of using these noncancer mosaicism to trace clonal relationships arises. With the cell over-proliferation, programmed cell death, and different biological processes, the mutant/variant allelic fraction (fraction of mutant/variant alleles over reference + alternative alleles, or 2* fraction of mutant cells for a diploid genome), we previously built a mathematical model and proved the feasibility of using these fractions, to reflect the genetic relationship between multiple samples obtained at similar development levels.
Practically, to perform the mosaic variant analysis for clonal reconstructions in human tissues, an extensive sampling and innovative experimental designs are required. In some cases, the experiment design never existed. For example, in a previous study where we discovered the left-right neocortical hemispheric genetic differences before the rostral-caudal lineage distribution barriers, we extensively sampled both hemispheres from multiple human donors that we collected ourselves, because in most brain banks, after harvesting the tissue, one hemisphere was stored frozen while the other was formalin-fixed, a standard procedure that has existed for decades.
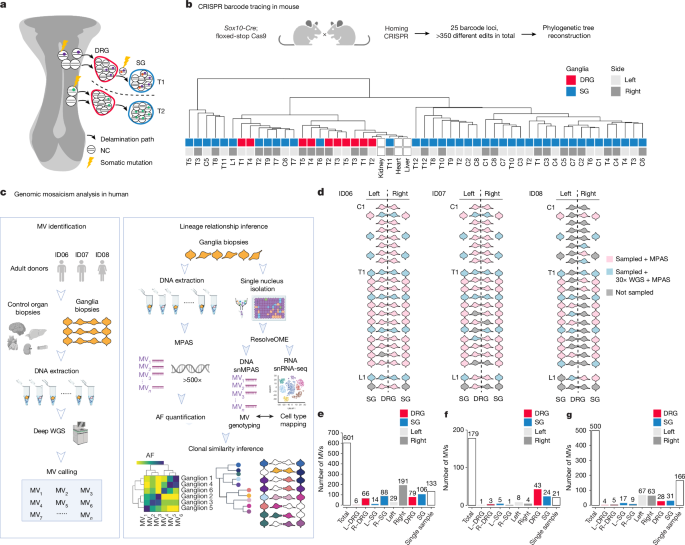
In other cases, to overcome the low sample size because of this experimental setup, we have to be creative in selecting intrinsic controls: as we already know the left-right lineage difference in the human brain, when we look at the inhibitory and excitatory neruonal clonal relationships, we can use similar set of mosaic variants, with significantly different allelic fractions between the left and right hemisphere, as independent biological replicates. Similarly, in the current study to uncover fate determination in neural crest-derived ganglia, a similar distribution pattern was observed in independent individuals, multiple mosaic CRISPR lines, as well as by traditional live imaging in quail models, demonstrating the power of mosaic variant-based barcode analyses.
In this current study, interestingly, the dorsal root ganglia and sympathetic ganglia, although bilaterally placed along the body, showed astonishing left-right genetic similarity distributed rostral-caudally in contrast to the neocortex. This likely reflected the early fate determinations as pointed out in this publication. Quantitative modeling, as well as evidence in multiple model organisms, also supports this observation. This further shows that mosaic variant barcode analyses not only pick up human-specific early developmental events, but also those conserved events between vertebrates, even with indications of how the vertebrate nerve system evolves. The observations here might also shift our views towards human disease diagnosis and treatment. We further hope that the mosaic variant barcode analysis we established through these studies can shed new light on embryonic developmental studies, especially in humans.
Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in