Soil conditions and the plant microbiome boost the accumulation of monoterpenes in the fruit of Citrus reticulata ‘Chachi’
Published in Microbiology
The bioactive components of the medical plants are the key factor determining the therapeutic effect of traditional Chinese medicine. The medicinal material quality of Citrus reticulata ‘Chachi’ differs depending on the bioactive components influenced by the planting area. However, plant-specific metabolism is poorly understood in relation to environmental factors such as soil nutrients and root-associated microbiome.
In the current study, a multi-omics approach was used to unravel the role of environmental factors in the accumulation of monoterpenes in the peel of Citrus reticulata ‘Chachi’ procured from the core (geo-authentic product region) and non-core (non-geo-authentic product region) geographical regions, respectively. In the core region, seven monoterpenes exhibited significantly higher concentrations in citrus peel compared to non-core regions. we revealed for the first time that the soil environment (high salinity, Mg, Mn and K) enhanced the monoterpene content by promoting expression of salt stress-responsive genes and terpene backbone synthase in host plant from the core region, while rhizosphere soil and endophyte microorganisms triggered the plant immune response or provided precursors of monoterpenes. The results provide a fundamental basis for improving the fruit quality depending on bioactive components by formula fertilization and precise microbiome management in agricultural practice.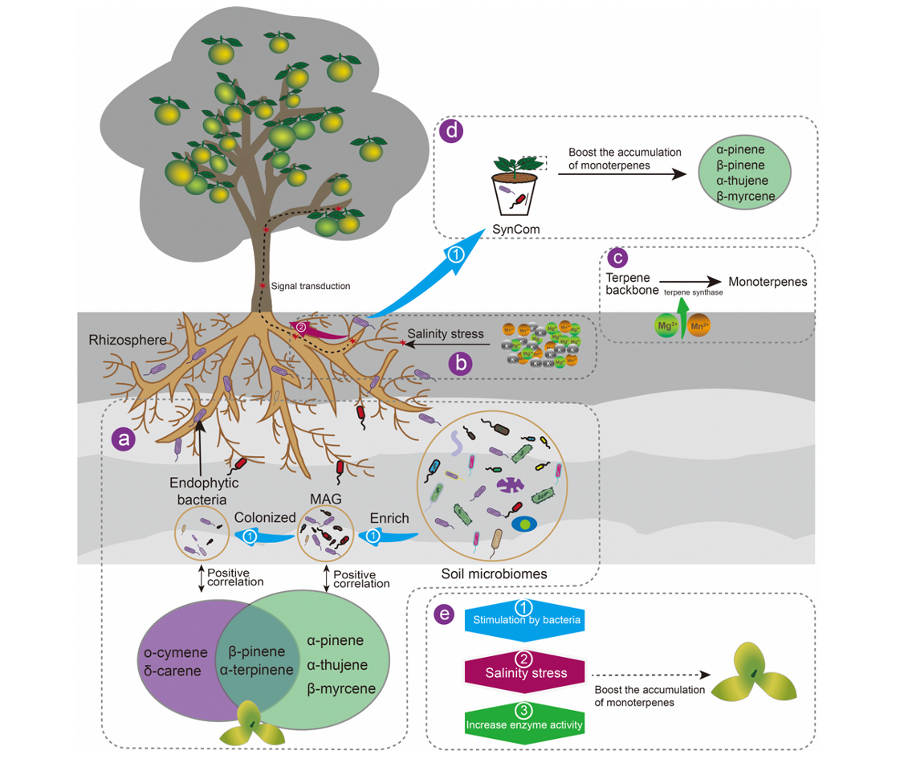
Jianmu Su1, Yayu Wang2, Huan Liu2 and Hong Wu1.
1State Key Laboratory for Conservation and Utilization of Subtropical Agro-bioresources, Guangdong Laboratory for Lingnan Modern Agriculture, College of Life Sciences, South China Agricultural University, Guangzhou 510642, China.
2State Key Laboratory of Agricultural Genomics, BGI-Shenzhen, Shenzhen 518083, China.
Follow the Topic
-
Microbiome
This journal hopes to integrate researchers with common scientific objectives across a broad cross-section of sub-disciplines within microbial ecology. It covers studies of microbiomes colonizing humans, animals, plants or the environment, both built and natural or manipulated, as in agriculture.
Related Collections
With Collections, you can get published faster and increase your visibility.
Animal Gut Nutrition and Greenhouse Gas Mitigation
Animal Microbiome, Journal of Animal Science and Biotechnology and Microbiome call for submissions to the collection on Animal Gut Nutrition and Greenhouse Gas Mitigation.
Efforts to reduce greenhouse gas emissions from livestock systems increasingly hinge on innovations in animal gut nutrition. The dynamic relationship between the gut microbiome and nutrient utilization plays a pivotal role in shaping methane output, feed efficiency, and overall sustainability. Advances in microbial ecology—particularly in understanding the role of gut microbiome in nutrient metabolism—are opening new pathways for mitigating emissions while enhancing productivity. These developments support the implementation of climate-smart agricultural strategies to address climate change and its impacts.
Looking ahead, continued research in this field has the potential to yield innovative solutions such as targeted probiotic supplementation, which could further optimize gut function and enhance nutrient absorption. These advancements may lead to reduced greenhouse gas emissions while improving animal health and productivity. By deepening our understanding of the animal gut microbiome, we can contribute significantly to sustainable agricultural practices that benefit both the environment and food security.
We invite researchers to contribute to this special Collection on Animal Gut Nutrition and Greenhouse Gas Mitigation. Topics of interest include but are not limited to:
- Animal Gut Microbiome and Feed Efficiency
- Greenhouse Gas Mitigation Strategies
- Rumen Fermentation Dynamics
- Nutrient Utilization in Livestock
- Probiotic Supplementation Effects
- Sustainable Livestock Production Practices
- Climate-Smart Agriculture Innovations
This Collection supports and amplifies research related to SDG 13, Climate action.
All submissions in this collection undergo the relevant journal’s standard peer review process. Similarly, all manuscripts authored by a Guest Editor(s) will be handled by the Editor-in-Chief of the relevant journal. As an open access publication, participating journals levy an article processing fee (Animal Microbiome fees, Journal of Animal Science and Biotechnology fees, Microbiome fees). We recognize that many key stakeholders may not have access to such resources and are committed to supporting participation in this issue wherever resources are a barrier. For more information about what support may be available, please visit OA funding and support, or email OAfundingpolicy@springernature.com or the Editor-in-Chief of the journal where the article is being submitted.
Publishing Model: Open Access
Deadline: Sep 04, 2026
The Apple Microbiome
Microbiome and Environmental Microbiome are calling for submissions to our Collection on the Apple Microbiome.
With world apple production estimated at 84 million tons, the microbiome of the apple has significant implications for agriculture, food security, and human health. Understanding the complex interactions between apple plants and their associated microbial communities can lead to improved crop management strategies, enhanced fruit quality and longevity, and sustainable agricultural practices. Recent advances have highlighted the role of specific bacteria and fungi in promoting plant health and resilience against specific pathogens. Moreover, detailed profiling of these microbial communities, revealing their diversity and functional potential facilitate exciting future developments, such as the identification of beneficial microbial consortia for biocontrol and the formulation of tailored probiotic treatments for both plants and humans. By advancing our collective understanding in this area, we can work towards a more sustainable and resilient agricultural system.
Topics of interest include but are not limited to:
-Microbial diversity and function associated with apples
-Effects of soil health and rhizosphere interactions on apple production
-Impact of climate change on the apple microbiome
-Role of the apple microbiome in fruit quality
-Microbiome-driven strategies for disease resistance
This collection is open for submissions from all authors on the condition that the manuscript falls within both the scope of the collection and the journal it is submitted to.
All submissions in this collection undergo the relevant journal’s standard peer review process. Similarly, all manuscripts authored by a Guest Editor(s) will be handled by the Editor-in-Chief of the relevant journal. As an open access publication, participating journals levy an article processing fee (Microbiome, Environmental Microbiome). We recognize that many key stakeholders may not have access to such resources and are committed to supporting participation in this issue wherever resources are a barrier. For more information about what support may be available, please visit OA funding and support, or email OAfundingpolicy@springernature.com or the Editor-in-Chief of the journal where the article is being submitted.
Publishing Model: Open Access
Deadline: Aug 05, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in