The story behind publishing “HorusEye” in Nature Computational Science
Published in Protocols & Methods and General & Internal Medicine
Our paper, HorusEye: a self-supervised foundation model for generalizable X-ray tomography restoration (https://doi.org/10.1038/s43588-026-00973-3), has finally been published in Nature Computational Science. As the first author, I want to take this opportunity to share the journey behind the paper, from writing to publication, as well as some of my own reflections along the way. I hope this post offers something beyond the paper itself: a few thoughts and lessons that may resonate in a different way.
Origins and Motivation
The starting point of this work was actually quite far from what it eventually became.
It goes back to the first project I worked on after starting my PhD, around 2021: pulmonary artery and vein segmentation (see Deep learning-driven pulmonary artery and vein segmentation reveals demography-associated vasculature anatomical differences. Nature Communications, https://doi.org/10.1038/s41467-025-56505-6). At that time, the methodology was already mostly in place when our hospital collaborators sent us a new batch of data and asked us to test the model on it.

That dataset was extremely noisy. The segmentation results were dramatically worse than what we had seen during testing. Many vessel branches appeared fragmented or broken, far beyond anything we had expected. We first tried to improve the robustness of the segmentation model itself, but none of those attempts really worked. In the end, we began to wonder whether denoising the images before segmentation might be a better direction.
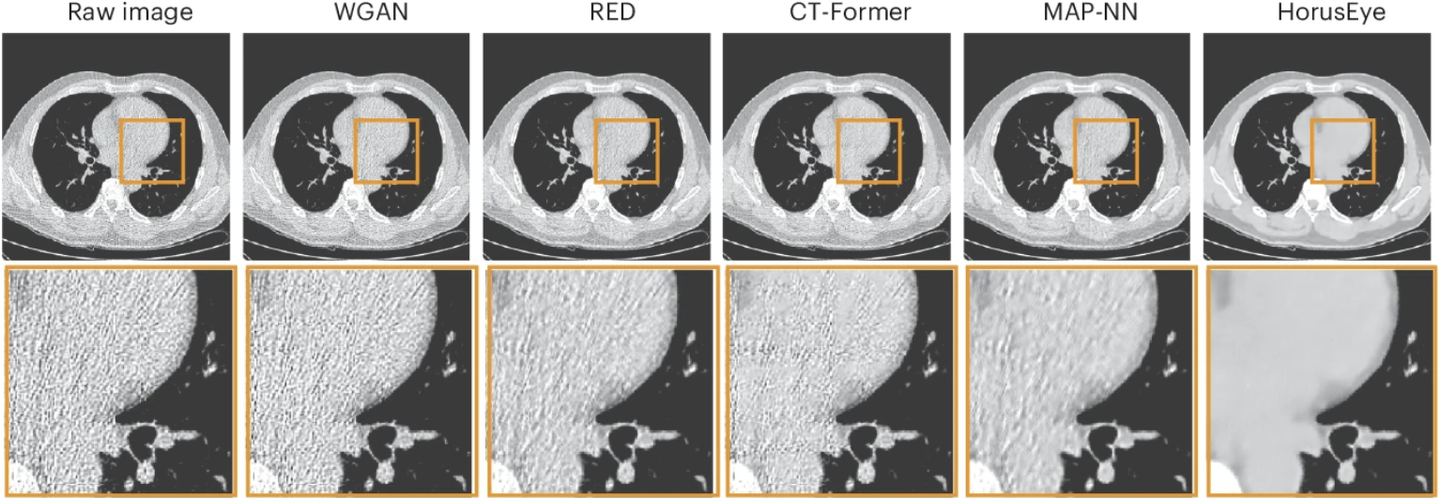
After digging into the literature, we found that CT denoising was already a fairly mature field, with many open-source methods available. So we went to GitHub, tried several representative approaches, and quickly realized that none of them worked well on this dataset either. Much of the noise remained, and the downstream segmentation problem was still unresolved. That was the moment we started asking a deeper question: why were these methods failing?
One obvious reason was that most existing methods relied on the log-Poisson noise assumption. Roughly speaking, this assumes that photon attenuation through matter follows a Poisson process and that the resulting noise can therefore be modeled accordingly. But as we looked more carefully across different datasets, we found that this assumption often did not hold. Real-world noise was usually much more complex, and the correlations between pixels were often much stronger than what log-Poisson noise would suggest. That observation pushed us toward self-supervised denoising, in the hope of finding a different way forward.
The Turning Point
At the time, Deep Image Prior (DIP) was very popular. Its key idea is that neural networks tend to recover continuous signals more easily, and this inductive bias had been successfully applied to many restoration problems.
But in CT, the problem is that the noise itself can also exhibit a surprising amount of continuity. So DIP did not work as well as we had hoped. That led us to ask a more specific question: what aspect of the noise is actually discontinuous?
A natural answer was inter-slice noise.
We started to think about transferring the DIP idea from within-slice modeling to across-slice modeling, and that was where the core idea of this paper was born.
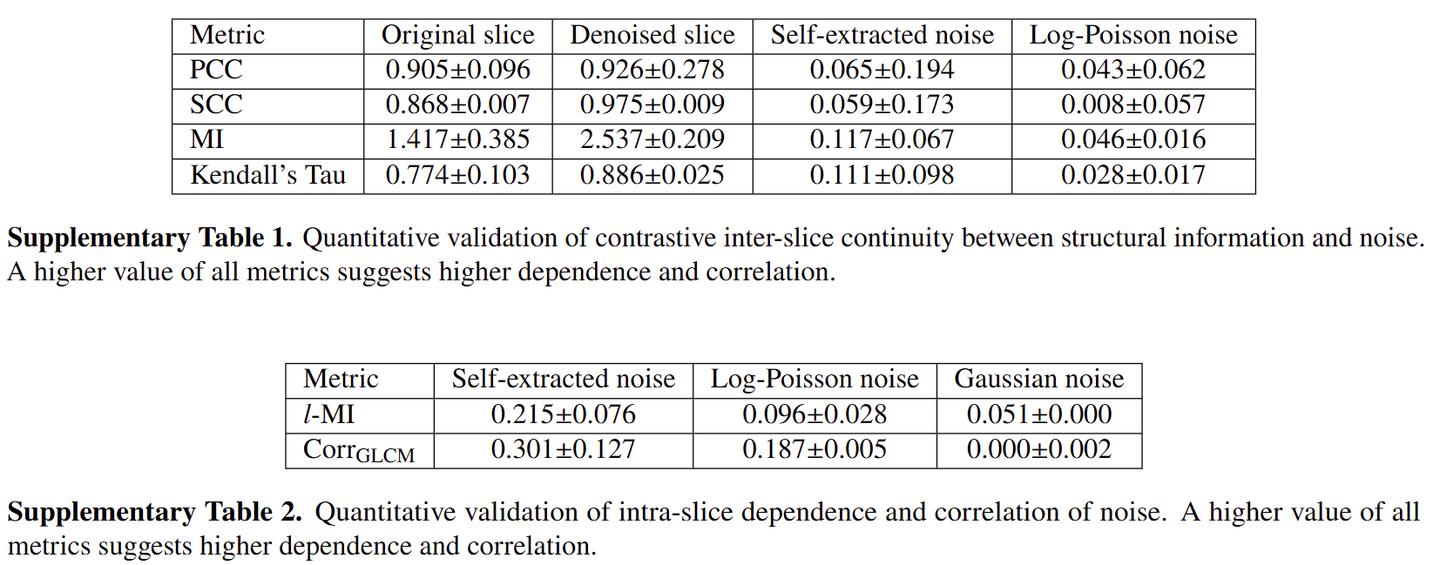
In simple terms, we wanted to use DIP to separate noise from a sequence of continuous slices. This led to the first module: we take the slices above and below as input, and the model predicts the middle slice. The residual between the prediction and the original image then largely captures the noise components that are discontinuous across adjacent slices.
Of course, that residual inevitably contains prediction errors as well. So we applied a high-frequency filter to the residual to remove the low-frequency part of the prediction error. After that, what remained could be treated, to a large extent, as noise extracted directly from real data.
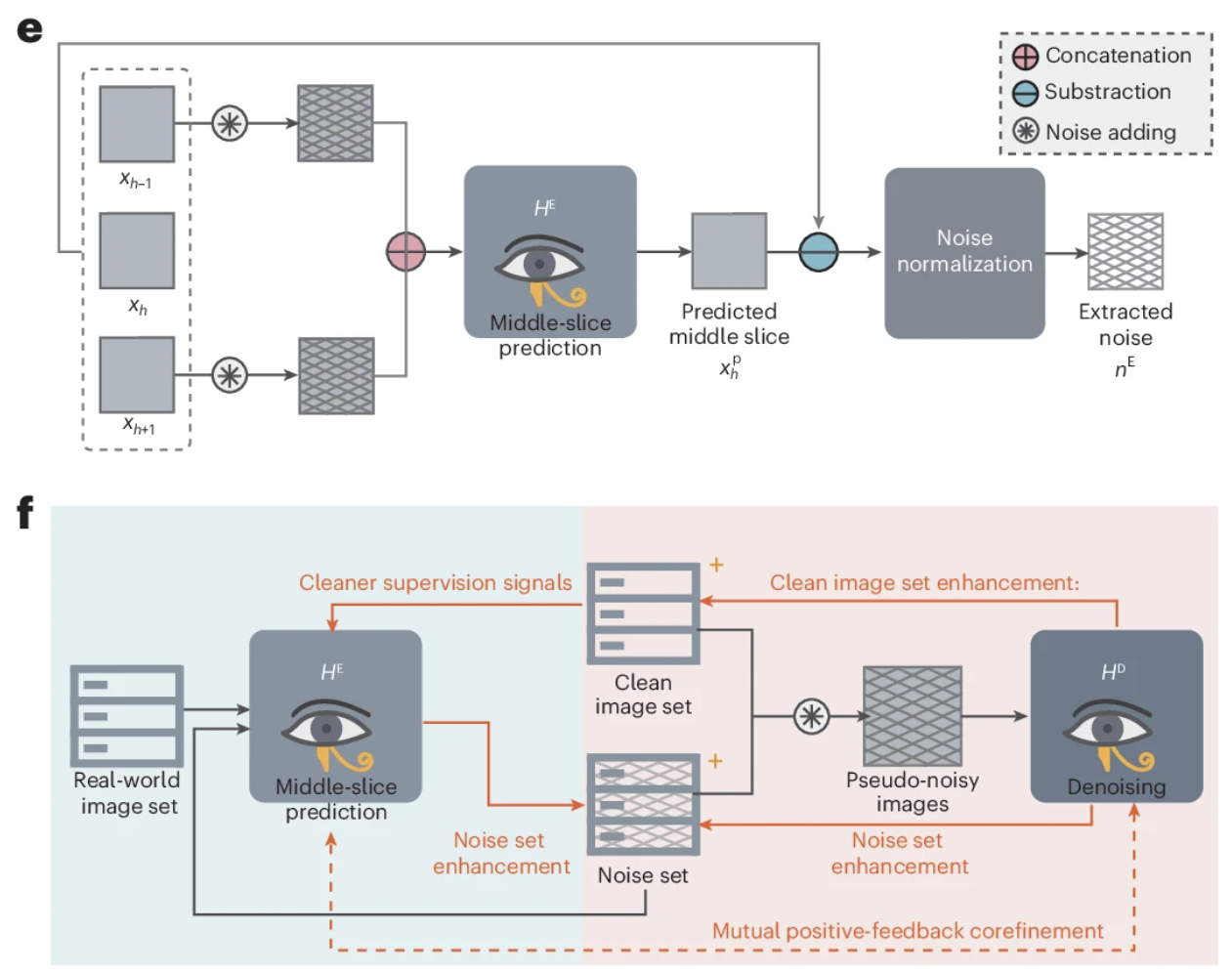
To further preserve structure, we introduced a second module: a direct denoising module. We add the extracted noise to clean images, which turns the training process into something closer to a standard denoising setup.
However, the whole idea depends on one crucial step being accurate: extracting noise by predicting the target slice from its neighbors. And that is exactly where things can go wrong, because the extracted “noise” may still contain some continuous components inherited from adjacent slices.
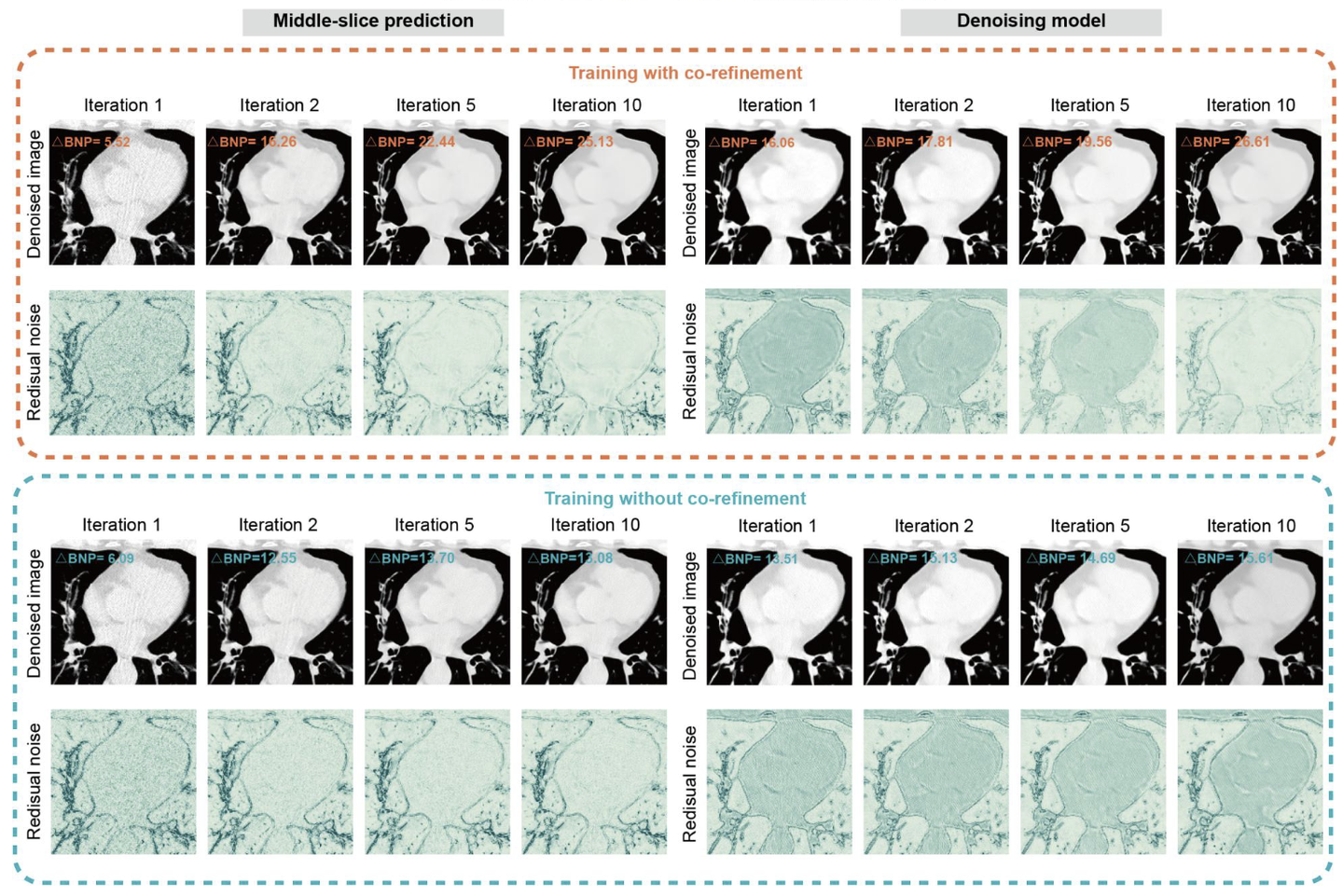
To improve this, we introduced a mutual positive-feedback loop for co-refinement. The intuition is fairly simple. By injecting the extracted noise back into the process, we can artificially amplify inter-slice discontinuity in the noise, making the model more sensitive to continuous structural information and therefore better at isolating truly discontinuous noise. At the same time, this co-refinement process also acts as a form of data augmentation, which is important because the number of clean images available for the direct denoising module was actually quite limited. Empirically, the co-refinement strategy did help. It created a positive loop that further improved the model’s capability (see Supplementary Note 6). The full training process is described in detail in the paper, so I will not repeat it all here.

The Long and Winding Road of Writing and Publishing
Once the full pipeline was in place, the results were much better than we had expected.
The denoising performance was very strong, and when applied to that noisy dataset, it also led to a clear improvement in artery-vein segmentation. At the time, our advisor felt that one of the main strengths of the work was that it challenged the traditional Poisson-based assumption, so we decided to aim for ambitious journals.
Looking back now, CT denoising is a much deeper field than we had realized at the time. Our understanding was still limited, and we also had very little experience in shaping a paper for publication. In hindsight, an important challenge was not simply the method itself, but how the work was framed. The project had originally grown out of a very concrete application problem, whereas publishing in top AI venues often requires a rather different narrative structure, with a clearer emphasis on broader methodological questions and generalizable technical contributions. At that stage, we had not yet fully figured out how to present the work in that way. As a result, the manuscript went through several rounds of rejection, and for quite a long time I felt uncertain about where the project should go next, despite the enormous amount of effort we had already invested.
What gradually changed the trajectory of the project was the realization that the work might matter not only as a solution to one denoising problem, but as part of a more general restoration framework. Around that time, we became interested in the Nature Methods paper Pretraining a foundation model for generalizable fluorescence microscopy-based image restoration. It was also centered on restoration, but it framed the contribution as a foundation model connected to multiple downstream tasks. That made us wonder whether our work could be developed in a similar direction.
In fact, from the very beginning, we had already noticed that our methodology could potentially generalize across many CT modalities, including animal CT and even some micro-CT applications such as fossils and minerals. We also reached out to veterinary hospitals for data access and collaboration. This gave us more confidence that the work could be extended naturally into a foundation-model framework. From there, we started to push the project in that direction: collecting as many modalities as possible, expanding the scope from conventional CT to X-ray tomography, and evaluating the model on a wider range of downstream tasks.
The results turned out to be encouraging, and that eventually led to the current paper.
When we first submitted the work to Nature Computational Science, we were excited to see it move into review. However, the reviewer comments were far more demanding than we had hoped. In particular, they raised serious questions about the sufficiency of the experimental evidence, the support for some of our claims, and the overall persuasiveness of several proposed innovations. In response, we prepared a rebuttal of nearly 200 pages and carried out a large number of additional experiments, many of which were eventually included in the supplementary materials. In the end, it was this round of substantial revision and clarification that made the paper much stronger and helped us address the reviewers’ concerns.
Reflections and Limitations
There is certainly no shortage of CT denoising work in the literature, and our contribution also has its own limitations. Looking back, the eventual publication of this paper was shaped not only by the work itself but also by timing, framing, and a fair amount of persistence. The broader interest in foundation models certainly provided a helpful context, but what ultimately mattered was that the work could be developed and validated at a scale that supported that framing.
From today’s perspective, I still would not consider the methodology perfect.
The first issue is the long-standing problem of fidelity in image restoration. Although we designed the framework with additional constraints to preserve structure, some information loss is still inevitable during denoising. A very typical example is trabecular bone: its texture can look remarkably similar to noise, and existing denoising methods often struggle to distinguish the two. As a result, over-smoothing can appear in spinal regions.
Another issue is that the overall training framework remains fairly complex, which means its robustness could still be improved. In practice, we also found that the model is sensitive to the choice of CT window width and window level. Some structures become much easier to detect under certain settings, whereas under a larger window width they may become over-smoothed. For this reason, in our collaborations with hospitals, we usually recommend treating the denoising model as an optional denoising layer rather than as part of a mandatory preprocessing pipeline, so that clinicians can choose between denoised and original images according to the specific diagnostic context.
These are, in many ways, common limitations of image-domain denoising and restoration methods. As for projection-domain reconstruction methods, we did not consider them at the time because of the original application scenario that motivated this work. Combining image-domain and projection-domain approaches is something worth exploring in future work, and it is also becoming an important direction in CT denoising research.
Thank you again for taking the time to read this blog. If this story has left you interested in our work, I would be very glad if you chose to read our original paper for more technical details. We also sincerely welcome any suggestions or questions, whether about the manuscript itself or the story behind it. I hope this blog has shared not only the paper, but also some of the thoughts, struggles, and reflections that came with the journey.
Follow the Topic
-
Nature Computational Science
A multidisciplinary journal that focuses on the development and use of computational techniques and mathematical models, as well as their application to address complex problems across a range of scientific disciplines.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in