Understanding the humoral response to hMPV F protein: Enlighting the therapeutic antibody and vaccine development for a common respiratory infectious pathogen
Published in Microbiology
Humoral response, a key component of adaptive immunity, is mediated by antibody molecules to target specific antigens, such as foreign viral antigens during natural infection. Isolation of antibodies with potent neutralization activity from convalescent patients has been demonstrated as an effective approach to identify antiviral agents against infectious pathogens, such as SARS-COV-2 and RSV, providing protections from further infections. Furthermore, characterization of the humoral immunity repertoire, which consists of a large variety of antibodies targeting different epitopes of the antigen, helps us better understand how our immune system works to recognize different regions of viral antigens and generate protective antibodies. Such information may provide guidance to design an engineered vaccine, which could shape our immune response towards generating the most potent neutralizing antibodies for protections, better than the natural infection.
Human metapneumovirus (hMPV) is a common pathogen that causes acute lower respiratory infection. The hMPV infections in high-risk populations, including infants, young children, elderly people, and immunocompromised patients, are more likely to develop severe symptoms such as bronchiolitis and/or pneumonia that results in considerable hospitalization, mortality, and morbidity. Unfortunately, an effective prophylactic modality, such as neutralizing antibody, antiviral drugs, or vaccine, is not yet available.
The trimeric fusion (F) protein of hMPV is a major antigenic target for neutralizing antibodies and vaccine development. hMPV F plays a critical role to aid the virus entry to the host cell, by structural rearrangement from prefusion (PreF) to postfusion (PostF) status. In the current manuscript, we isolated over one hundred antibodies targeting the hMPV F protein from memory B cells of multiple donors. Some of these antibodies have shown extremely high neutralization potency in vitro, suggesting the potential of developing them as effective antiviral therapies. We mapped most isolated neutralizing antibodies to the structure of hMPV F antigen by various antibody characterization approaches, revealing multiple distinct antibody recognition sites. We identified the correlations between the neutralization potency of antibodies and their recognition sites – for example, the antibodies recognize the side regions of hMPV F tend to have stronger neutralization potencies, whereas the antibodies targeting the apex regions is very rare. This is different from the understanding of respiratory syncytial virus (RSV), a closely related virus to hMPV from the same Pneumoviridae family, where most dominant neutralizing antibodies target the apex region of prefusion F protein. Furthermore, the maturation of trimeric hMPV PreF is known to require protease-mediated cleavage process. We found that different processed forms of hMPV F antigen are recognized by the antibodies differently – a subset of antibodies that bind processed PreF show greater neutralization potency than the antibodies bind preferentially to unprocessed PreF, and antibodies specifically binding to postF has poor neutralization activity, suggesting that the processed PreF might be the better framework for vaccine development. These findings provide us a roadmap to design an hMPV vaccine with desired properties to be recognized by the most potent neutralizing antibodies.
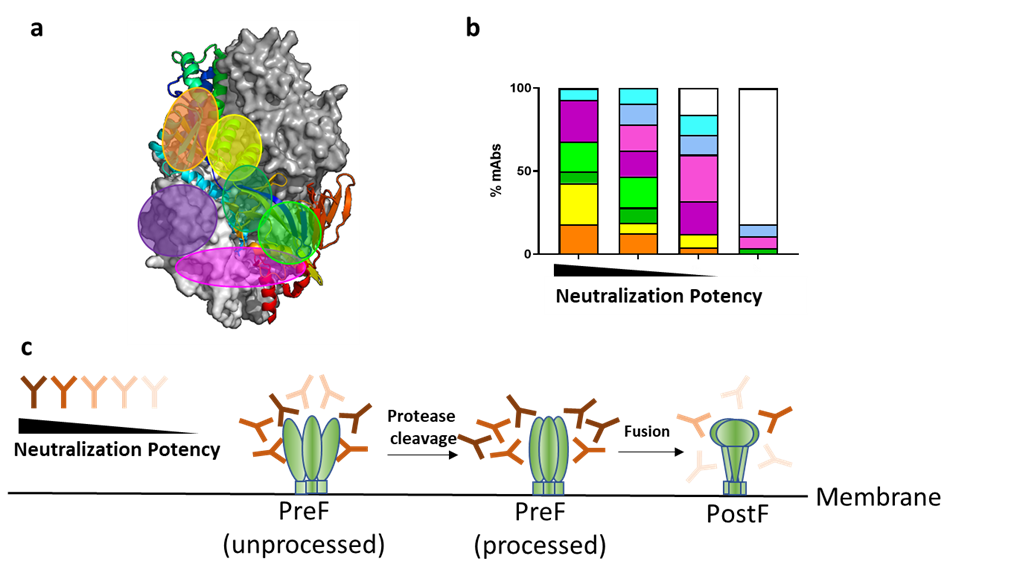
Figure. (A) Different antibody recognition sites mapped on hMPV F trimer structure, represented in different colors. (B) Percentage of antibodies targeting each recognition site, grouped by neutralization potency. (C) Different formats of hMPV F antigens are recognized by the antibodies with different neutralization potency.
Read the full article here: https://www.nature.com/articles/s41467-022-30205-x
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in