United by the goal of improving precision medicine, Hematologics Inc and the Lamba Lab collaborated to publish ‘Interstitial Loss of Heterozygosity at Chr19q13 in CD33-Negative AML‘.
Published in Biomedical Research
Explore the Research
Just a moment...
ascopubs.org
An Enormous Clinical Data Repository
Hematologics Inc, located in Seattle, WA, is a CLIA certified clinical laboratory that offers an expansive test menu. Few facilities offer flow cytometry, cytogenetics, and molecular testing with speciality in hematopoiesis. These testing capabilities, at production scale, characterize incredibly rare and unique leukemias. One such discovery was Acute Myeloid Leukemia (AML) that is CD33 negative by flow cytometry in eight different patient samples. This was a significant observation because Leukemias that completely lack surface expression of CD33 cannot be targeted by CD33-directed antibody-drug conjugates. But how, on a genetic level, is the expression of CD33 lost? As a clinical production lab, Hematologics relies on the research tools and expertise of esteemed collaborators to get at these basic research questions.
A Tried-and-True Collaboration
Enter the Lamba Laboratory, led by Dr. Jatinder Lamba at the University of Florida in Gainesville, Florida. Dr. Lamba’s team applies a multi-omics toolkit to study AML at the genomic level. By dissecting a patient’s unique leukemia down to its nucleotides, this research makes tailored treatment possible. Previously, Hematologics and the Lamba Lab joined forces to publish another story connecting different CD33 expression levels and treatment efficacy. Specific single nucleotide polymorphisms (SNPs) were shown to modulate CD33 expression in AML. Depending on which SNP the patient possessed, low, medium, or high responsiveness to treatment was observed. What a difference a nucleotide can make! Once again, Hematologics collaborated with the Lamba Lab, this time to further investigate the genetic basis for the complete loss of CD33 in AML.
CCGG Deletion: The Tip of the Iceberg
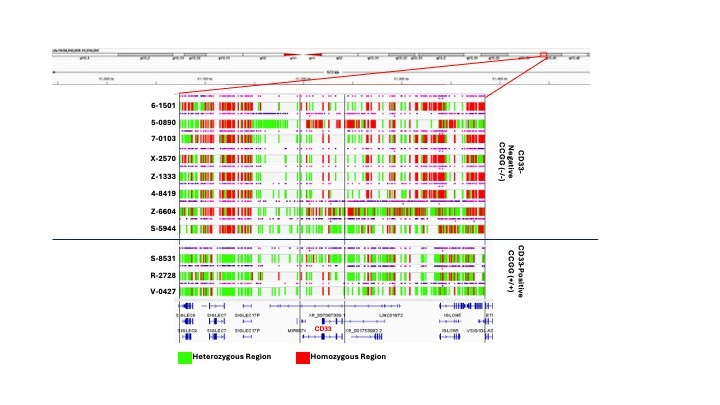
After comparing the CD33 positive and CD33 negative groups, the Lamba lab quickly identified a ‘CCGG’ deletion in all eight CD33 negative patients that was not observed in the CD33 positive patients. This 4bp deletion, present in both alleles, created a premature termination codon and clearly explained the complete loss of CD33 protein. However, the molecular profile of CD33 negative AML proved to be far more complex, and this homozygous mutation was the tip of the iceberg. There was a larger pattern at play. Large stretches of the CD33 locus were homozygous in the CD33 negative patient group harbouring the homozygous CCGG deletion. These same stretches were almost entirely heterozygous in CD33 positive patients. This evidence of loss of heterozygosity in the CD33 locus was unprecedented.
Iterative Improvement in Patient Care
What started as CD33 negative myeloid cells identified by a clinical lab became the research discovery of loss of heterozygosity at the CD33 locus. This is only half of the story. The characterization of rare phenotypes in AML will then inform new genetic test development. As more clinical data is collected, the molecular landscape of AML will continue disentangle ultimately treatment design will improve. When researchers and the clinical lab collaborate, like Hematologics and the Lamba Lab, both basic biology and the clinical test menu win, which means that patients win.
Follow the Topic
-
Leukemia
This journal publishes high quality, peer reviewed research that covers all aspects of the research and treatment of leukemia and allied diseases. Topics of interest include oncogenes, growth factors, stem cells, leukemia genomics, cell cycle, signal transduction and molecular targets for therapy.


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in