What’s in my mRNA vaccine? mRNA vaccine quality analysis using direct RNA sequencing.
Published in Genetics & Genomics
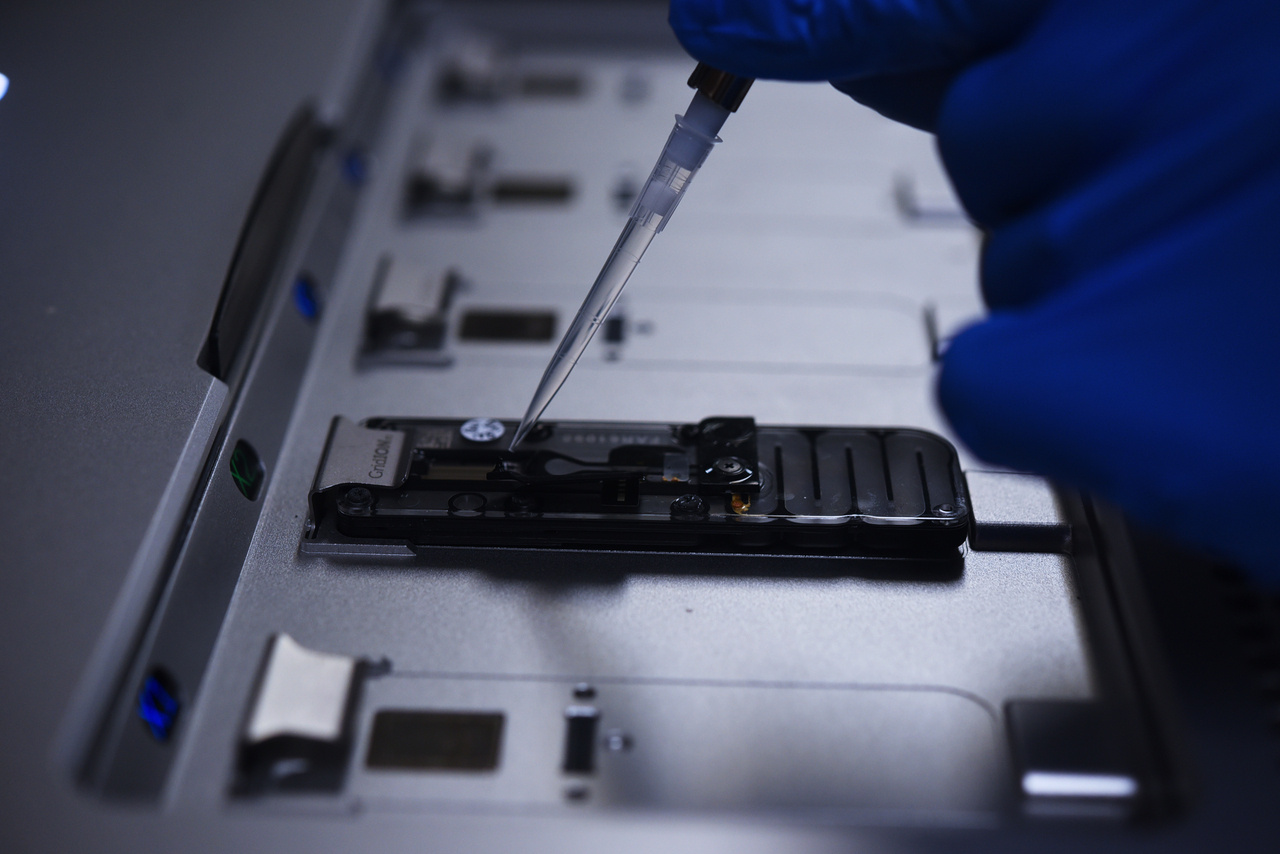

At present, therapeutic mRNAs are in vitro transcribed from a linearised DNA template using an RNA polymerase. Each mRNA has a 5’ cap, a poly(A) tail, 5’ and 3’ UTRs, and a coding region, which can encode any protein. The 5’ cap, UTRs, and poly(A) tail are optimised to ensure mRNA stability and translation by the host cell. mRNA quality evaluations test mRNA for a range of critical quality control attributes, including identity, integrity, and purity (e.g. the presence of double-stranded RNAs, off-target RNAs transcribed from the DNA template, and RNAs from contaminating microorganisms). As these contaminants can cause a wide range of detrimental effects to the patient, their identification is important for the safety and efficacy of mRNA therapies.
Currently, the mRNA manufacturing industry employs a range of protocols to test mRNA quality. These include Sanger sequencing, capillary electrophoresis, Mass Spectrometry, IP-RP HPLC, and immunoblotting. However, supporting such a range of tests can be time-consuming and expensive.
For more than a decade, RNA sequencing has been used for gene expression and splice variant analyses, however to date, it has not been used extensively in mRNA vaccine quality testing. Our research aimed to test whether RNA sequencing could be used to analyse mRNA vaccine quality. DNA sequencing was also used to investigate the quality of the plasmid DNA template. Our team developed a comprehensive method for mRNA vaccine quality testing using Oxford Nanopore sequencing and a custom bioinformatics pipeline.
For the development and benchmarking of our test, we designed an Enhanced Green Fluorescent Protein (eGFP) mRNA vaccine control construct. The plasmid template also includes 5’ and 3’ UTRs and a poly(A) tail, in a pUC-57 plasmid backbone, and can be in vitro transcribed to generate eGFP mRNA. Both short read (Illumina) and long read (Oxford Nanopore) DNA sequencing were used to test the sequence identity and purity of the plasmid template. Both methods confirmed the consensus accuracy of the plasmid sequence. Additionally, we detected trace amounts of E. coli sequences that were not fully removed during plasmid purification, which prompted the optimisation of our plasmid extraction protocols.
Next, Illumina and Oxford Nanopore cDNA sequencing were used to analyse mRNA vaccine quality. Both library preparation methods involved the Reverse Transcription of mRNA into cDNA. Both Illumina and Oxford Nanopore sequencing confirmed the consensus accuracy of the cDNA sequence. Illumina sequencing showed a lower per-nucleotide error rate than Oxford Nanopore sequencing.
However, Oxford Nanopore long-read sequencing permitted additional analyses on mRNA vaccine identity, integrity, and purity, that were not possible using short-read sequencing (Figure 1). As Oxford Nanopore cDNA sequencing libraries do not include a fragmentation step, library fragment length is an indicator of mRNA integrity. Indeed, our cDNA library fragment sizes reflected our capillary electrophoresis testing of mRNA integrity. Additionally, we detected RNA polymerase mis-binding and mRNA transcription of regions outside the T7 promoter, which included the plasmid backbone. The analysis of polymerase mis-binding in long reads can assist in optimising in vitro transcription reaction conditions and plasmid backbone sequence.
Moreover, we could accurately measure poly(A) tail length from Oxford Nanopore cDNA sequencing data. Long homopolymer tracts such as poly(A) tails are inherently error-prone, as the boundaries between adjacent bases can be difficult to distinguish. We used tailfindr to calculate Poly(A) tail length, as it calibrates homopolymer length measurements for individual mRNA molecules, based on the speed at which they traverse the nanopore. The tailfindr calculations accurately reflected our expected poly(A) tail lengths. This method is therefore suitable for identifying poly(A) deletions, which can accumulate in the plasmid template during the plasmid propagation step.
Uniquely, mRNA can also be sequenced without a Reverse Transcription step using Oxford Nanopore chemistry (direct RNA sequencing). This method avoids the errors introduced by Reverse Transcriptases, and polymerases, which are both used to prepare cDNA sequencing libraries. It also maintains the poly(A) tail length information, providing accurate tail measurements, again, using software such as tailfindr. Moreover, direct RNA sequencing preserves the modified bases used in mRNA vaccines, such as N1-methylpseudouridine. Modified bases can be used to boost the efficacy and safety of mRNA drugs by evading the host inflammatory response. At the time of writing, the commercially available Oxford Nanopore base callers do not recognise N1-methylpseudouridine. However, as we observed a characteristic error profile for N1-methylpseudouridine, we predict that subsequent base callers can be trained to recognise it.
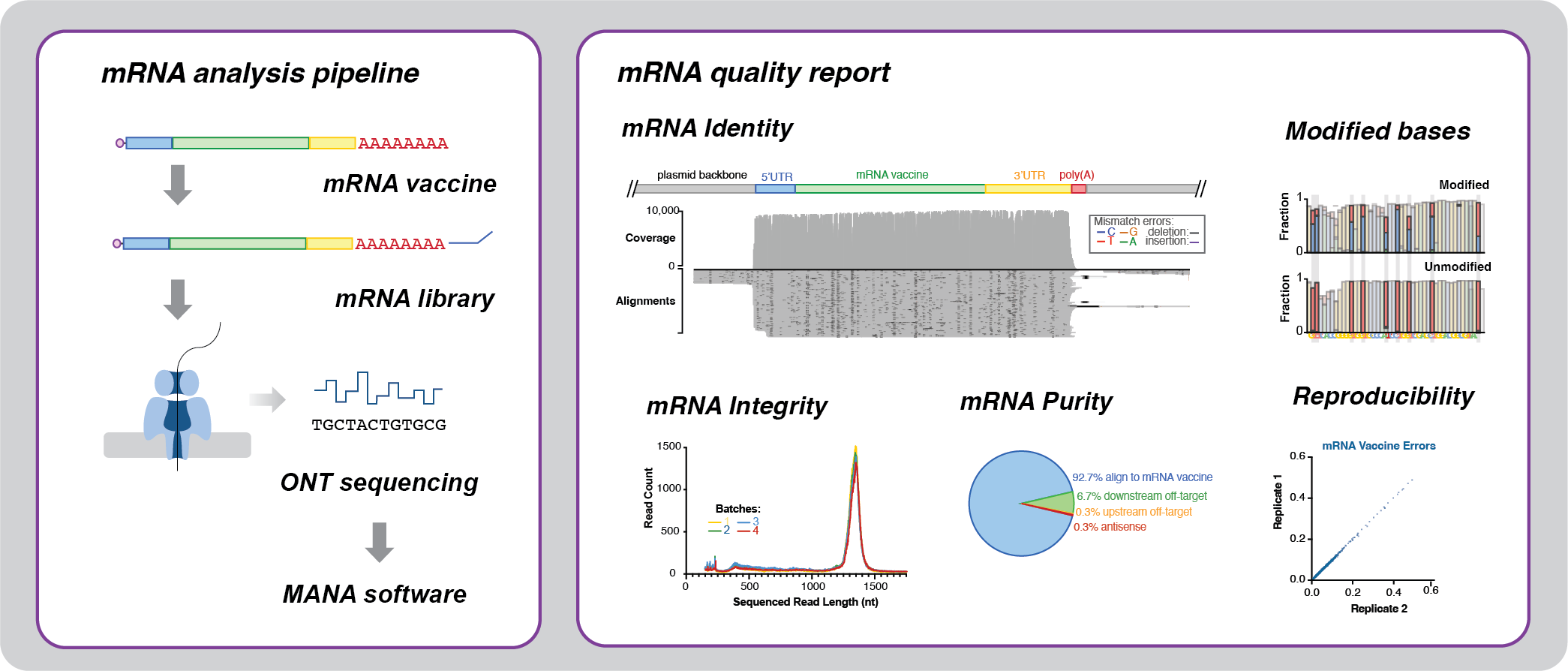
Our manuscript shows that Oxford Nanopore sequencing can be used to evaluate mRNA vaccine quality metrics such as mRNA identity, integrity, and purity. Our work provides proof of concept that Oxford Nanopore sequencing can comprehensively measure mRNA vaccine quality. However, our results are based on a single, control mRNA construct.
Our next challenge is to optimise direct RNA sequencing for use by the mRNA manufacturing industry. To achieve this, we are partnering with Oxford Nanopore and the BASE mRNA facility at the University of Queensland to use the most recent RNA sequencing chemistries to evaluate a range of critical quality attributes for different mRNA constructs.
Through measuring mRNA vaccine critical quality attributes in a single test, our research shows that mRNA vaccine quality tests using Oxford Nanopore sequencing have the potential to improve the efficiency and speed of mRNA manufacture. We are excited to see how quality tests such as ours may be used to manufacture the mRNA therapeutics of the future.
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in