A chemical catalyst that promotes histone acetylation like enzymes in live cells
Published in Chemistry

The research landscape
Life emerges from a network of biomolecules and chemical reactions catalyzed by enzymes. Since enzyme abnormalities are often associated with a variety of diseases, a chemical catalyst that could replace dysfunctional endogenous enzymes and promote physiologically important intracellular reactions would greatly aid in the elucidation and treatment of disease. Such a strategy would be superior to traditional therapeutic approaches of controlling endogenous enzyme function with inhibitors, activators, etc. in that therapeutic efficacy would not depend on the activity of endogenous enzymes and vulnerability to the acquisition of resistance could be avoided.
We previously developed chemical catalysts that can promote one of the fundamental physiological reactions, lysine residue acetylation1,2. Notably, our catalysts have already achieved histone lysine acetylation, known as an epigenetic modification in the regulation of gene transcription, in living cells3,4. However, these catalysts required the addition of an exogenous acetyl donor, despite the presence of the metabolite Ac-CoA as an endogenous acetyl source. The use of high concentrations of exogenous donors could be a barrier to future therapeutic applications due to side effects from nonspecific acetylation and the difficulty of simultaneous catalyst-donor bimolecular targeting. In addition, sustained histone acetylation is difficult due to the depletion of external donors in the reaction. Therefore, achieving catalytic histone acetylation using the metabolically replenished endogenous Ac-CoA as a sole acetyl source, which is what we have addressed in this work, is a difficult but must-be-accomplished task.
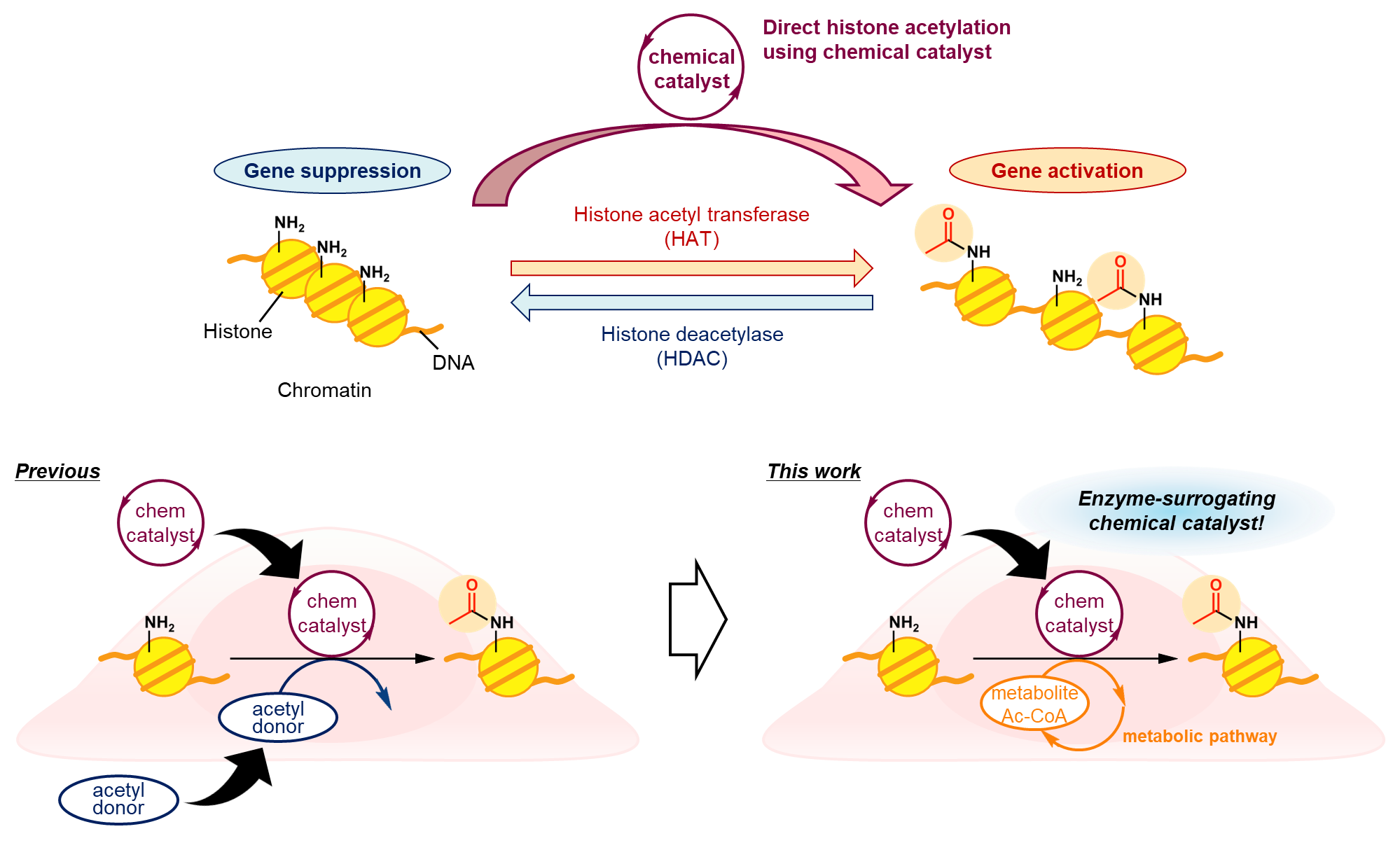
Which is better; e-deficient or e-rich?
To efficiently and potently activate the acetyl group of Ac-CoA, we devised a design that combines a thiol group as an Ac-capturing motif with a hydroxamic acid moiety (HXA) as a catalytic core for nucleophilic activation5. Contrary to our expectations, however, the histone acetylation activity of the initially synthesized catalyst, pHXA, was low in a test tube. Improving the catalyst activity was essential, but formulating a strategy for structural modifications was not a straightforward task; if HXA is more electron-deficient, the reactivity of the active "OAc" intermediate is likely to increase, and if it is more electron-rich, the rate and amount of "OAc" formation is expected to increase. It was difficult to predict whether e-deficient or e-rich would be more effective. Still, we finally found that the reactivity of the "OAc" intermediate is sufficient and that increasing its production is the key to improving activity. We reported on this catalyst improvement process in more detail in a preprint version of this paper6.
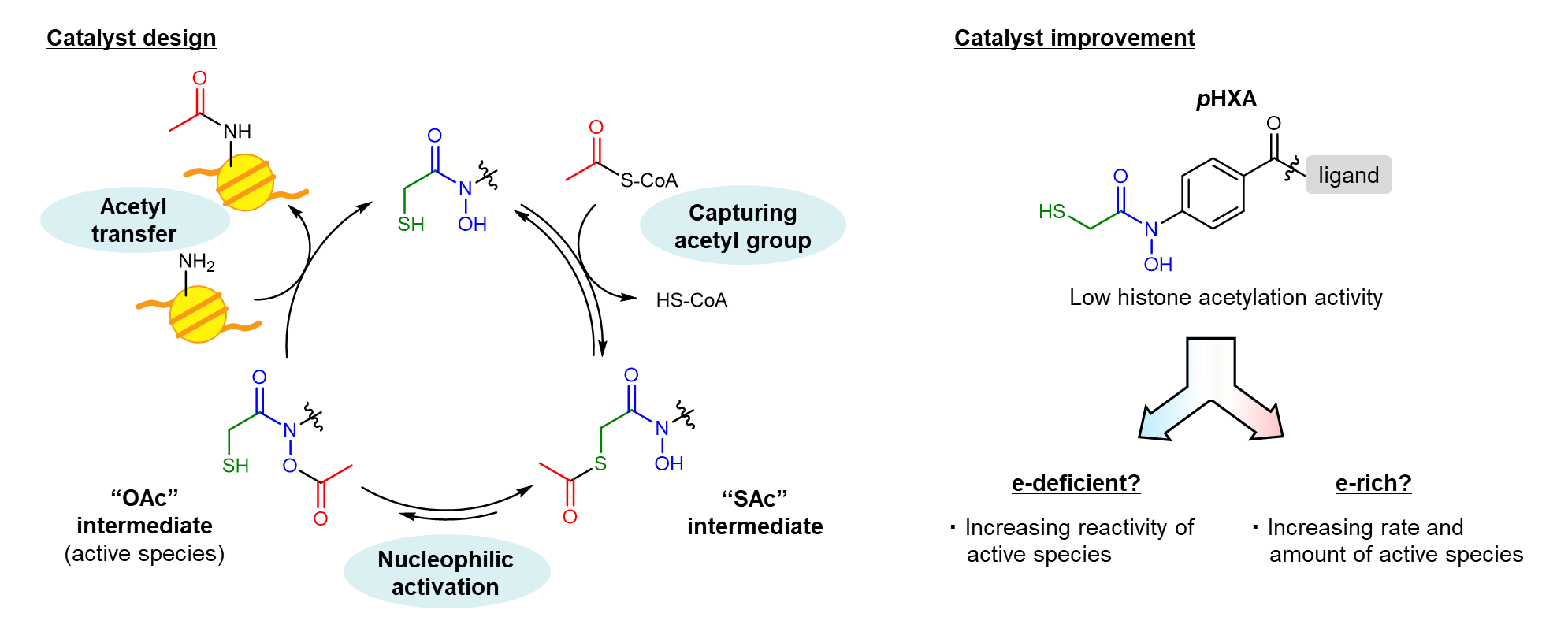
Progress from zero to one
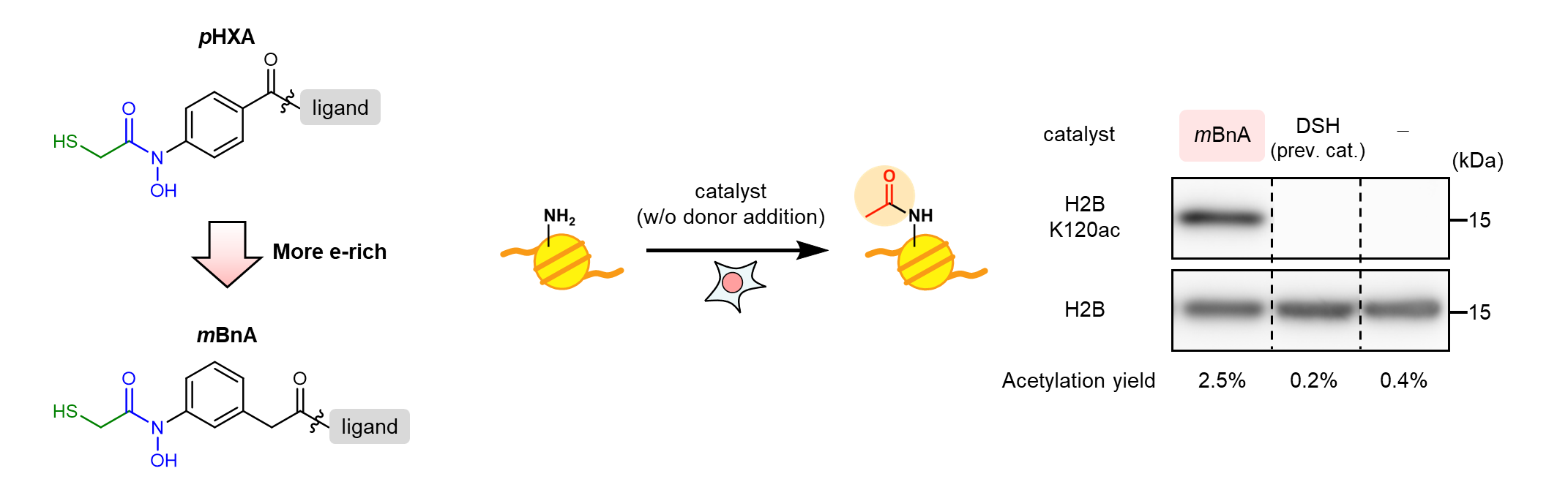
Based on our findings, we synthesized a mBnA catalyst with a more electron-rich HXA. As expected, the histone acetylation activity was significantly improved. As a next step, we attempted to catalyze histone acetylation in living cells. In initial trials, reactions with a high concentration of an exogenous acetyl donor were performed. We also tried the reactions without external donors as negative control experiments. Interestingly, the mBnA catalyst promoted histone acetylation using only endogenous Ac-CoA without adding exogenous acetyl donor, the phenomenon which had never been observed with the previously developed catalysts. This was the most exciting moment in this research. It was delightful that the incorporation of the hydroxamic-acid catalytic core has yielded such a remarkable "0 to 1" achievement.
Probing in-cell Ac-CoA level and new biological findings
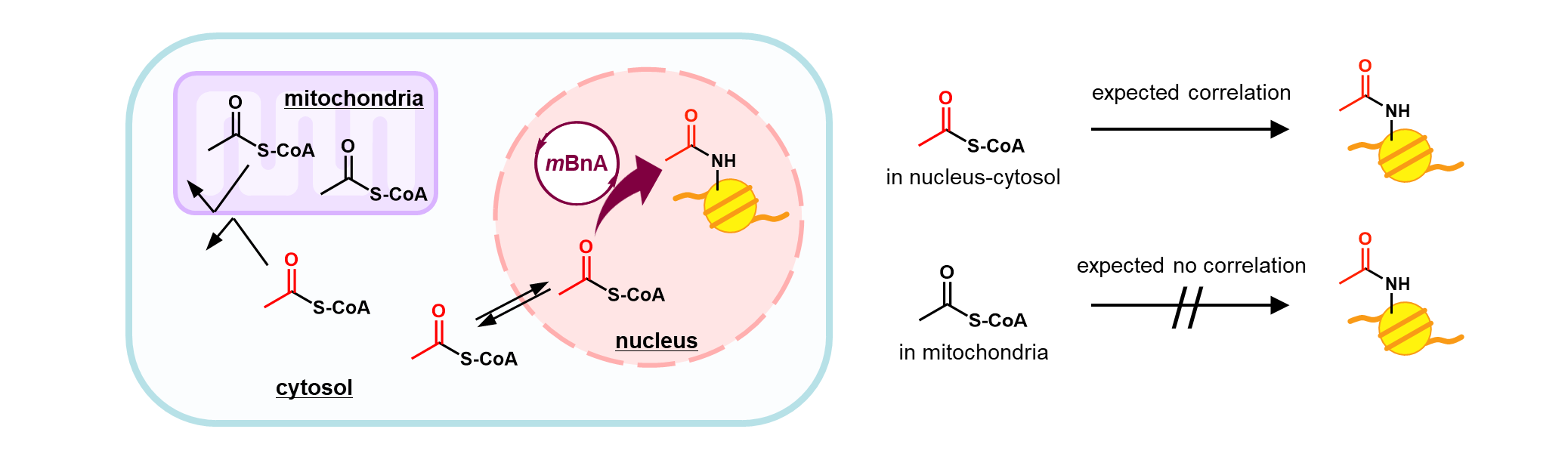
To show that mBnA is a "useable" catalyst, we explored further applications. The ability of mBnA to utilize intracellular Ac-CoA, especially in the nuclear-cytosolic compartment where histones are present, led us to focus on its potential as a chemical probe to detect changes in the nuclear-cytosolic Ac-CoA concentration via histone acetylation. In experiments aimed at increasing the in-cell Ac-CoA levels, we increased the concentration of glucose in the medium, a typical metabolic source of Ac-CoA. Yield of histone acetylation decreased, however, indicating that the in-cell Ac-CoA level decreased. This unexpected result initially puzzled us. After repeating the experiment, we were confident with the result. We realized that the catalytic histone acetylation is detecting the nuclear-cytosolic Ac-CoA level, segregated from the mitochondrial compartment by the lipid membrane. Namely, under glucose-rich conditions, the nuclear-cytosolic Ac-CoA concentration decreases likely because the consumption of nuclear-cytosolic Ac-CoA is increased prior to the supply of Ac-CoA from mitochondria. The ability to probe Ac-CoA concentrations in a specific compartment without technically cumbersome and metabolically disruptive fractionation steps adds another dimension to the value of mBnA.
Prospects
We have developed a lysine acylation catalyst, mBnA, which serves as a surrogate for endogenous histone acetyltransferases by promoting acyl group transfer from cellular Ac-CoA to histone protein lysine residues. This would be a steppingstone to a new therapeutic strategy replacing dysfunctional endogenous enzymes with chemical catalysts to regulate disordered epigenetic histone acetylation in disease. Toward this future vision, research is currently underway to intervene in transcription and phenotype through the catalyst-mediated histone acetylation, or acylation in more general. In addition, the detection of compartmentalized Ac-CoA concentration using the mBnA catalyst may contribute to future biochemical studies on Ac-CoA metabolism.
- Amamoto, Y. et al. Synthetic Posttranslational Modifications: Chemical Catalyst-Driven Regioselective Histone Acylation of Native Chromatin. J. Am. Chem. Soc. 139, 7568–7576 (2017).
- Adamson, C., Kajino, H., Kawashima, S. A., Yamatsugu, K. & Kanai, M. Live-Cell Protein Modification by Boronate-Assisted Hydroxamic Acid Catalysis. J. Am. Chem. Soc. 24, 10–14 (2021).
- Fujiwara, Y. et al. Live-cell epigenome manipulation by synthetic histone acetylation catalyst system. Proc. Natl. Acad. Sci. USA. 118, e2019554118 (2021).
- Fujimura, A. et al. Chemical catalyst/protein hybrid as artificial histone-modifying enzyme for epigenome manipulation. ChemRxiv https://doi.org/10.26434/chemrxiv-2022-xt610 (2022).
- Mizumoto, S. et al. Hydroxamic Acid-Piperidine Conjugate is an Activated Catalyst for Lysine Acetylation under Physiological Conditions. Chem. Asian J. 15, 833–839 (2020).
- Habazaki, M. et al. A chemical catalyst enabling histone acylation with endogenous Acyl-CoA. ChemRxiv https://doi.org/10.26434/chemrxiv-2022-zxn90 (2022).
Follow the Topic
-
Nature Communications

An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Mar 24, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in