AAV- homologous recombination based therapeutics
Published in Bioengineering & Biotechnology
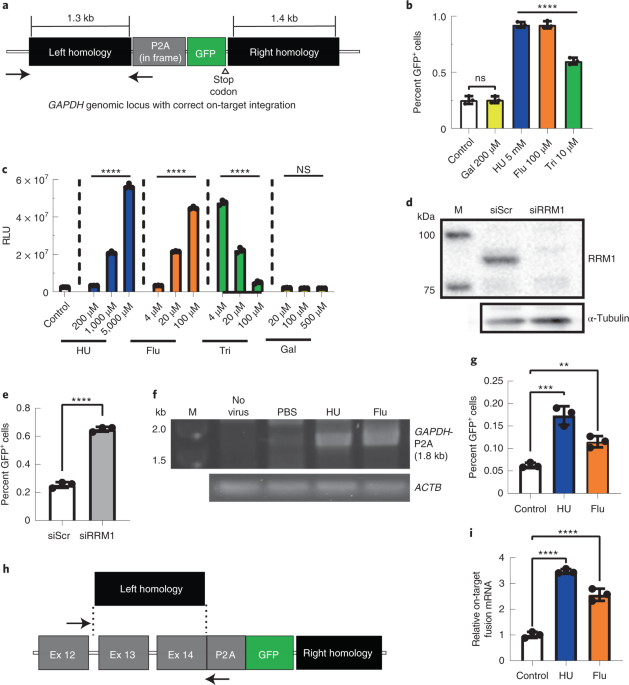
Recombinant AAV vectors have been the primary gene transfer vector for direct in vivo administration of therapeutic genes into humans. Due to the mostly episomal nature of AAV, the vector genomes are quickly lost in non-quiescent or growing tissues. The development of a strong immune response against the capsid makes vector re-administration extremely ineffective. Envisioning treating genetic diseases right after birth and maintaining life-long genetic correction, we had previously developed a genome editing approach (Barzel et al., Nature doi:10.1038/nature13864), which does not require the use of designer nucleases (e.g. CRISPR/Caspase9, zinc-fingers, Talens, meganucleases). The single-stranded nature of the AAV genome makes those vectors a suitable template for homologous recombination. This type of vector is designed in a way that a therapeutic transgene with a ribosomal skipping fragment (e.g. P2A) fused to its 5’ end is flanked by homology arms of around 1 kb each. The homology arms are designed to match sequences near the end of a targeted gene in the host genome that allows homologous recombination to occur just 5’ of the a gene’s translation termination signal which will be deleted after successful vector integration. Transcription of the edited endogenous gene will result in production of a chimeric mRNA that produces both the endogenous protein as well as the therapeutic protein due to ribosomal skipping. This strategy is a non-disruptive gene editing approach. In our first study we designed an AAV-homologous recombination (AAV-HR) vector targeting the Albumin locus and delivered it by simple intravenous infusion to the livers of mice. The Albumin gene was selected as a target because of its high transcription rate. The AAV-HR vector inserted the human factor IX coding sequence just downstream of the Albumin gene and we showed that transgene expression levels were sufficient to correct the bleeding diathesis in hemophilia B mice. Subsequently, several mouse models of human disease including Crigler- Najjar, methylmalonic acidemia, tyrosinemia, and ZZ alpha-1-antitrypsin deficiency have been treated using this approach. Currently this approach is being tested in humans to treat methymalonic acidemia.
While this work represented an important proof-of-concept, the limitation is the relatively low rate of homologous recombination in the liver estimated to occur in about 0.1 to 1% of hepatocytes in mice. Thus, the approach is useful for treating diseases where a relatively few cells can make enough of the protein to correct the deficiency and/or conditions where corrected cells offer a selective advantage and re-populate the liver over time. Currently, this approach will not work well in cases where there is no selective advantage and a higher proportion of corrected cells require the missing protein.
During our studies evaluating the mechanism of AAV-based homologous recombination, we discovered that ribonucleotide reductase inhibitors administered temporarily at the time of vector infusion enhanced AAV-HR rates by almost half a log. There are several ribonucleotide reductase inhibitors approved for use today, such as the purine analog fludarabine phosphate. Fludarabine is an antineoplastic commonly used in combination with other drugs in the treatment of leukemias and lymphomas. In addition, fludarabine is one drug in a cocktail used in a variety of ablation protocols for bone marrow transplantation, including the transplant of gene modified autologous bone marrow-derived cell transplantations. In our protocol, mice underwent a brief regimen of relatively low-dose fludarabine treatment without any additional drugs, thereby avoiding any overt toxicities or side-effects common observed for approved indications. In addition to the observed improvement for nuclease-free AAV-HR, we also found increased levels of HR when using the GeneRide vector in conjunction with CRISPR/Cas9. Our work provides an important example of using a FDA approved drug in combination with nucleic acid based therapeutics to improve its efficacy. Such combinational approaches may become more common place in clinical gene therapy applications. We also found that the improvement in AAV-HR was not related to an increase in DNA replication in targeted cells. In fact, the majority of cells that underwent AAV-HR did not undergo cell division. This finding was unexpected as it was widely believed that active DNA replication as it is occurring in mitotic cells was required for homologous recombination. Our studies raise additional questions regarding the mechanism of AAV-HR. Unraveling the underlying mechanisms of AAV-HR may enable the development of approaches that further enhance the efficacy of the GeneRide technology and opens up further possibilities for future genome editing based therapies.
Follow the Topic
-
Nature Biotechnology
A monthly journal covering the science and business of biotechnology, with new concepts in technology/methodology of relevance to the biological, biomedical, agricultural and environmental sciences.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in