Caenorhabditis inopinata: the unexpected sibling species of C. elegans
Published in Ecology & Evolution

The full paper in Nature Communications can be found here: https://www.nature.com/articles/s41467-018-05712-5
The nematode Caenorhabditis elegans is extensively used as a model organism in biology. One of C. elegans closest known relatives, C. briggsae, is widely used in comparative studies, but these two species are as phylogenetically distant as humans are to mice1. Because of this, a sibling species of C. elegans has been long sought. Our paper introduces C. inopinata, a newly discovered sibling species of C. elegans.
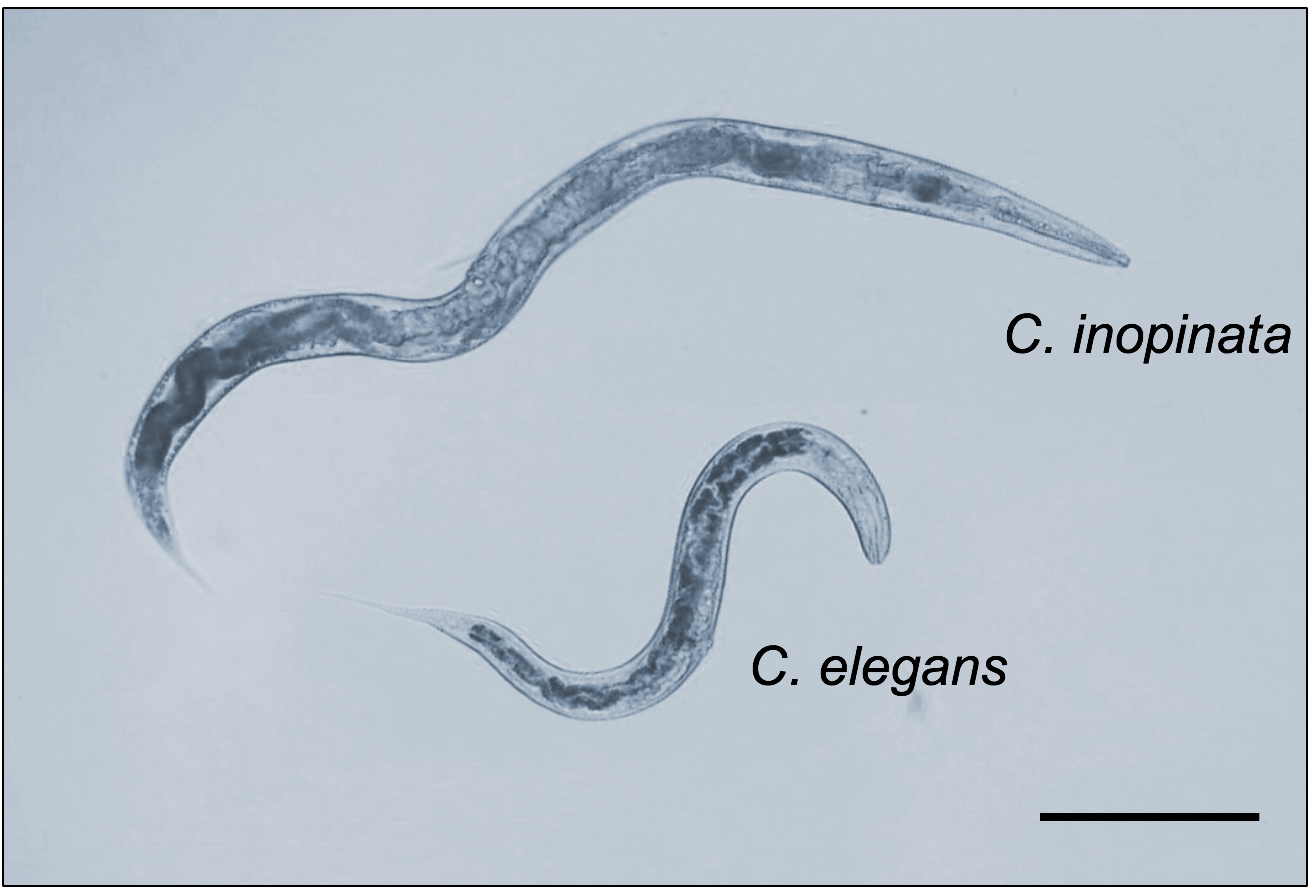
Since 2006, we have sampled nematodes from tropical and subtropical regions of the world. We were particularly interested in figs, which provide a stable and nutrient-rich habitat for nematodes, because nematode-fig associations have independently evolved at least 10 times2,3. It is interesting to note that fig-associated nematodes have distinct morphological characteristics e.g. a long and slender body, when compared to close relatives which do not inhabit figs, which likely reflects their adaptation to the fig environment.
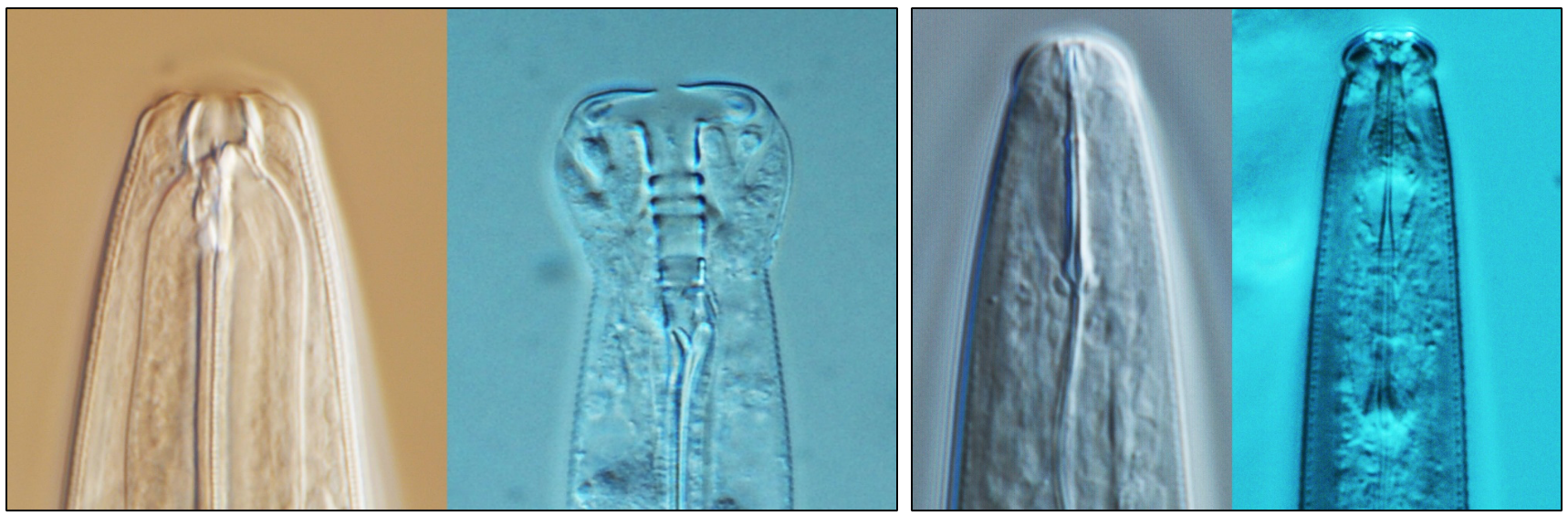
During our sampling project, we isolated a large, slender nematode from the fig Ficus septica in the Ishigaki islands of South-Eastern Japan. At first, we thought this nematode could belong to a new genus because of its distinct morphology, for example, the adults are twice the size of C. elegans adults. However, our molecular phylogenetic analysis revealed that this newly discovered species is the closest known relative of C. elegans. We named this species C. inopinata - ‘inopinata’ from the Latin for surprising or unexpected, because the researchers were surprised to discover a C. elegans sibling species with morphological and ecological characteristics so divergent from C. elegans, including the fresh fig habitat of C. inopinata which is in contrast to the rotting fruit environment commonly associated with C. elegans8.
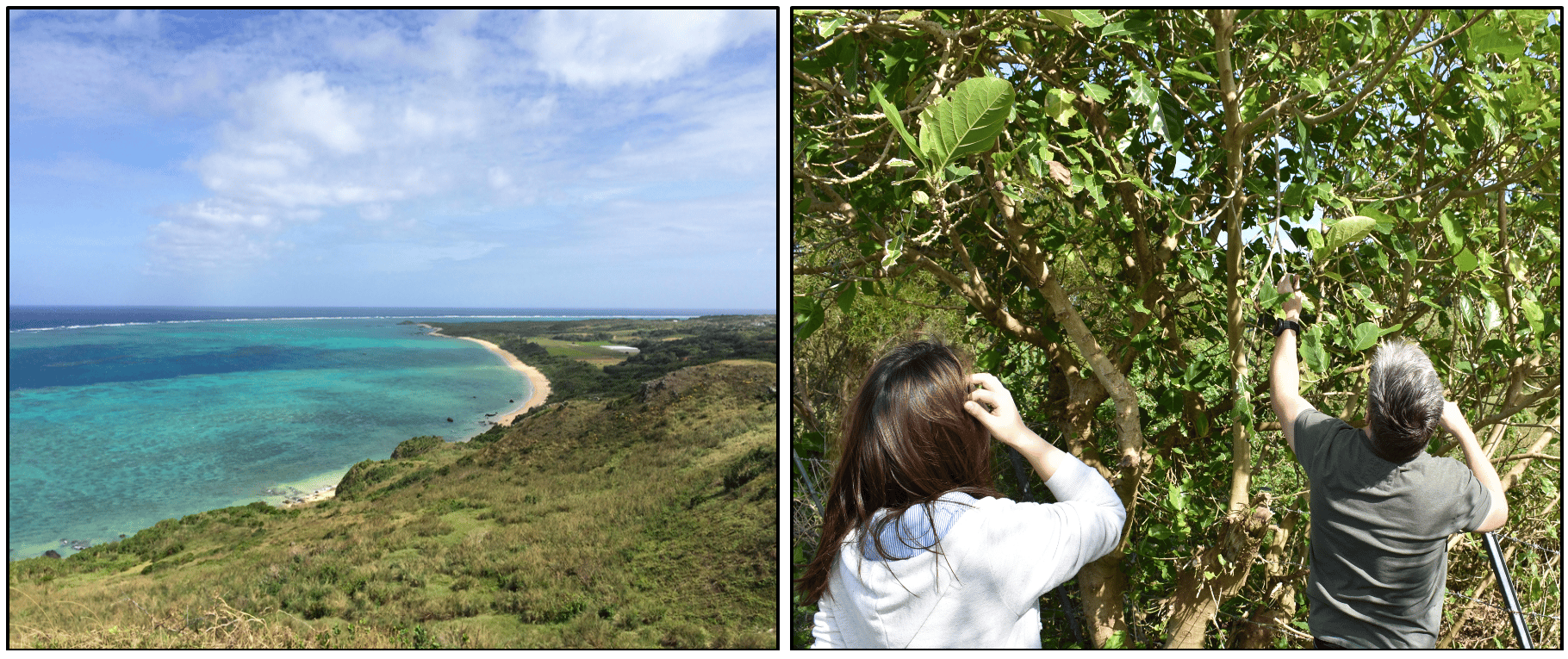

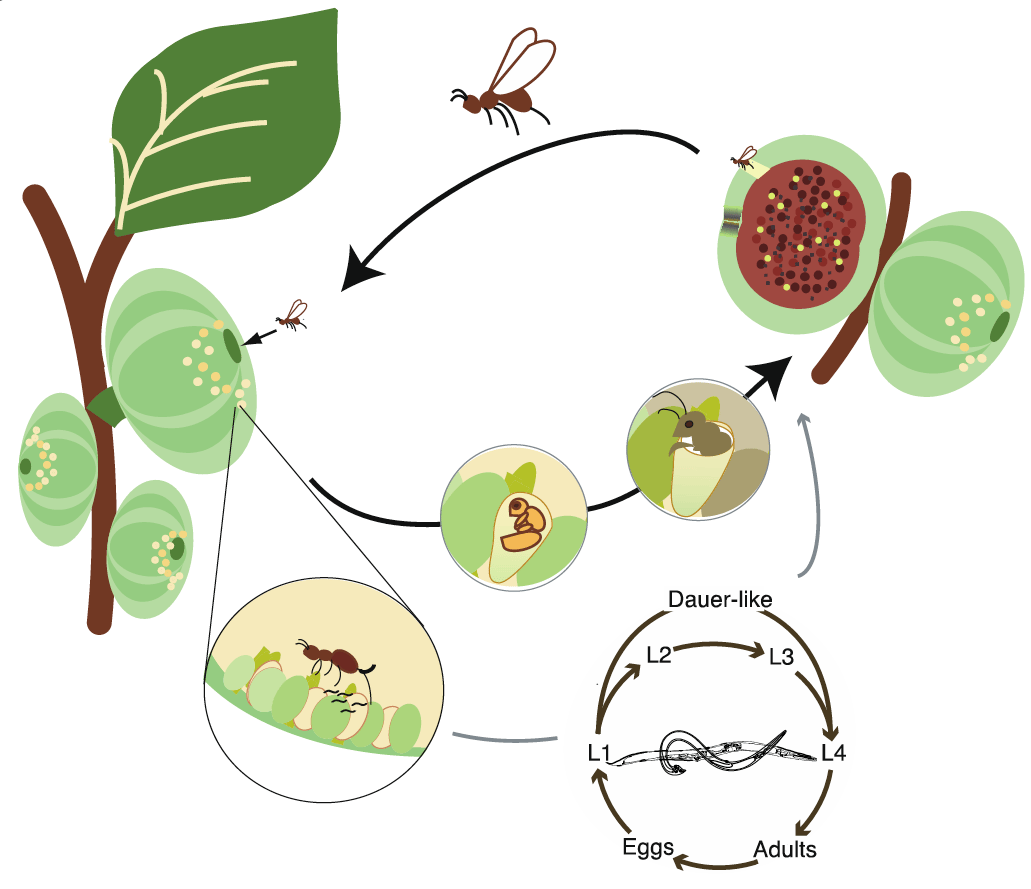
C. inopinata is vectored by a fig-pollinating wasp which has a mutualistic relationship with F. septica figs. The nematodes multiply inside the fresh figs and when the wasps emerge from the fruit as adults and disperse to new fig trees, where nematodes are transferred to fig fruits during wasp oviposition.
For C. inopinata to be a valuable laboratory model for comparative studies with C. elegans it is important that the genomic and genetic data, and laboratory techniques available for C. inopinata are comparable to those available for C. elegans. To achieve this, we sequenced and assembled the 123 Mb C. inopinata genome into chromosome-sized scaffolds (six autosomes and one X chromosome) and identified ~21,000 protein-coding genes. Although much of the genome and genes in C. inopinata and C. elegans are conserved, we identified some key differences between the two species. For example, the C. inopinata genome has an expansion of transposable elements but has lost many chemoreceptor-coding genes, and genes coding for key proteins in the ERGO-1 small RNA pathway.
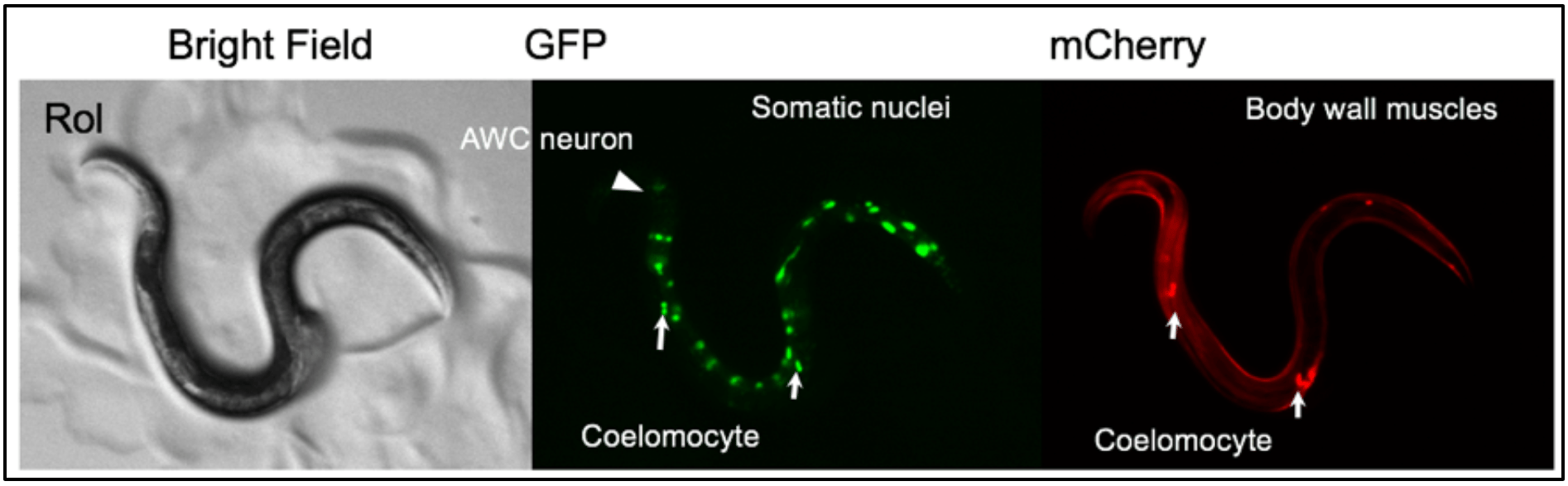
C. inopinata can be cultured using the same methods used for C. elegans. Furthermore, the high level of genetic similarity between C. elegans and C. inopinata allows the application of many laboratory techniques that are well-established in C. elegans to also be applied to C. inopinata. For example, establishing transgenic lines, manipulation by RNAi, and immunofluorescence using antibodies against C. elegans proteins can also be successfully carried out with C. inopinata. Thus, there is great scope for comparative studies on the biology, genetics, ecology and evolution of C. inopinata with C. elegans and other nematodes species.
Poster image: An artistic interpretation of C. inopinata biology by Abe Mana, University of Miyazaki.
References:
1. Cutter, A. D. Divergence times in Caenorhabditis and Drosophila inferred from direct estimates of the neutral mutation rate. Mol. Biol. Evol.25,778–786 (2008).
2. Susoy, V. et al.Large-scale diversification without genetic isolation in nematode symbionts of figs. Sci. Adv.2,(2016).
3. Davies, K. A. et al. A review of the taxonomy, phylogeny, distribution and co-evolution of SchistonchusCobb, 1927 with proposal of Ficophagus n. gen. and Martininema n. gen. (Nematoda: Aphelenchoididae). Nematology17,761–829 (2015).
4. Kanzaki, N., Tanaka, R., Giblin-Davis, R. M. & Davies, K. A. New plant-parasitic nematode from the mostly mycophagous genus Bursaphelenchus discovered inside figs in Japan. PLoS One 9,(2014).
5. Woodruff, G. C., Tanaka, R. & Kanzaki, N. Teratodiplogaster variegatae n. Sp. (Nematoda: Diplogastridae) isolated from the syconia of Ficus variegata Blume on Ishigaki Island, Okinawa, Japan. Nematology16,1153–1166 (2014).
6. Kanzaki, N., Giblin-Davis, R. M. & Ragsdale, E. J. Allodiplogaster josephi n. sp. and A. seani n. sp. (Nematoda: Diplogastridae), associates of soil-dwelling bees in the eastern USA. Nematology 17,831–863 (2015).
7. Kanzaki, N., Aikawa, T., Maehara, N. & Thu, P. Q. Bursaphelenchus kesiyae n. sp. (Nematoda: Aphelenchoididae), isolated from dead wood of Pinus kesiya Royle ex Gordon (Pinaceae) from Vietnam, with proposal of new subgroups in the B. fungivorus group. Nematology18,133–146 (2016).
8. Kiontke, K. C. et al. A phylogeny and molecular barcodes for Caenorhabditis, with numerous new species from rotting fruits. BMC Evol. Biol.11,(2011).


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in