When a new genome is assembled, it often feels like the bulk of the work is done – certainly, assembly is no small task – but in reality, the work of interpretation is just beginning. An assembled genome sequence gives us the order of the DNA bases, and the set of chromosomes to which they belong, but to make use of this information we need to annotate it – to determine which stretches of DNA correspond to genes, the biological units of meaning in the genome.
Gene annotation may sound like a routine step, but it is one of the main things that determines how useful a genome will be for later research – a consequence of which is that ‘model’ genomes (like those of humans or laboratory mice) have been annotated to far greater levels of detail (e.g. more precisely dissecting the internal structures of their genes) than those of many other species. In general, if annotations are incomplete or inaccurate, virtually all downstream analyses become more difficult.
DNA sequencing technologies have become faster, cheaper and more accessible over the years, to the extent that genome assemblies are now piling up in public databases – but annotating these assemblies remains a task far harder to scale. As such, there is an increasingly pronounced ‘annotation gap’ between the number of assembled genomes and the quality of genome annotations. This gap has widened in recent years as exceptionally large-scale sequencing endeavours – such as the Earth Biogenome Project, and the Darwin Tree of Life, both ambitious ‘moonshot’ initiatives to sequence every eukaryote in an area – have started to produce assemblies for many more species than before.
This was the starting point for our new gene annotation tool, ANNEVO. We began developing ANNEVO because we felt there was also still a gap between what genome sequencing projects require – accuracy, speed, and convenient ‘out of the box’ use, with minimal dependencies – and what existing annotation methods have come to provide. While current methods have undoubtedly benefited the field enormously, they almost invariably come with trade-offs – some depend on additional sources of sequencing evidence (such as transcriptomic or proteomic data), which for non-model species may simply be unavailable, some require elaborate species-specific setups, and some are so computationally demanding they do not easily scale up to process large numbers of genomes with any rapidity.
With ANNEVO, we asked whether it was possible to build an ab initio method of annotation – one that took as input only the genome assembly, and no other information – that was both accurate and practical across a wide range of species. We faced a number of challenges in doing so. Most notably, gene structures vary considerably across the evolutionary history of eukaryotes – and so, a method of annotation that works well for one group of species may not be as effective for another. Some genes are short and with relatively simple structures (and so, easier to annotate), while others – such as the intimidatingly huge TTN, almost quarter of a million bases long – are lengthy, complex, and challenging to predict accurately.
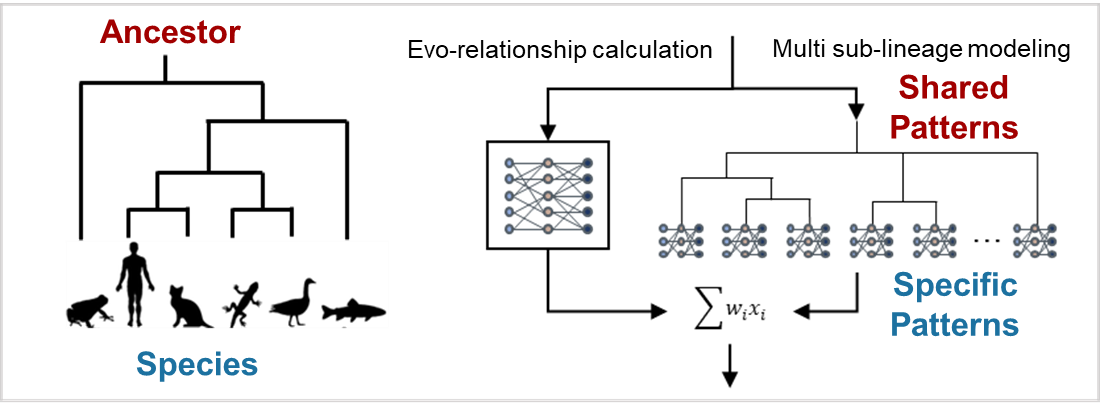
As the project developed, we kept returning to the same idea: if we wanted a method that would be useful beyond the narrow set of model (better-annotated) genomes that could serve as its benchmark, it had to accommodate biological diversity and the complexity of real genomes from the outset. That naturally led us to think about the evolutionary tree itself. Biology, famously, gives us this branching structure: species are related, but not identical, and common gene features are often found more within particular lineages (that is, individual branches of the tree) than across all of them. Tree-like structures have already inspired many ideas in computation too, and for us they suggested a fruitful direction — instead of forcing one single model to treat all species in exactly the same way, could we design a system that allowed different parts of the model to handle different evolutionary groups, each perhaps better at one group than the other? This line of thinking eventually led us to the mixture-of-experts framework implemented in ANNEVO (Fig 1).
One of the most encouraging parts of the project was seeing that ANNEVO did indeed work well across very different groups of species. We were also encouraged to find that its ab initio approach could, in some cases, perform strongly even when compared to the latest, evidence-heavy, annotation tools. We interpret this not as meaning those external sources of evidence (the transcriptomic and/or proteomic data upon which other tools rely) are no longer important — they certainly are — but that annotating the genome using the genome alone can produce just as accurate an outcome.
At the same time, we were very aware of another temptation in AI-driven gene annotation: simply making the model larger. In many areas of AI, increasing model size is often the first instinct, but we considered that for gene annotation – especially in the service of large genome projects – this would not be the correct approach. While a larger model has the potential to improve some results, we reasoned it could also make training slower, inference heavier, and real use much harder. From the outset, we wanted ANNEVO to remain controlled in size rather than becoming a very large model that only works well when abundant computational resources (which impose a cost and limit accessibility) are available. In this respect, practicality – in particular, saving the user’s time – shaped the design from the beginning. Thus, our aim was for ANNEVO to be a lightweight, simple tool, one which did not burden the user with excessive requirements. We were pleased to find that ANNEVO was fast. We could annotate a complete human genome in an hour and a half, maize in half an hour, and Arabidopsis thaliana, a model plant, in three minutes. For many species, this makes genome annotation a task now achievable in times somewhere between a coffee break and a lunch break.
Consequently, we believe the task of annotation is no longer only about whether one genome can be annotated well. In many cases, the outstanding challenge is whether dozens, hundreds, or even thousands of genomes can be processed within a reasonable amount of time. A new method that produces slightly better results but takes much more time to do so may not actually help large-scale sequencing projects very much. So, when developing ANNEVO, we were not solely thinking about accuracy – we were also thinking about whether the method could finish in a timeframe that makes sense for real projects and real users.
Nevertheless, outstanding challenges remain for genome annotation. No single method (even with a mixture of experts) solves every case. Biology is an intrinsically messy discipline, genomes remain diverse, and exceptional cases are not uncommon. ANNEVO does not provide a solution to these problems. Instead, we see ANNEVO as one step towards making annotation simultaneously more accurate, scalable, and broadly accessible. More generally, this project was shaped by a change happening across genomics. The field is moving from a world centered on a relatively small number of model organisms to one that includes a much larger share of life’s diversity. This is an exhilarating time in which to work, but it also means our computational tools must adapt. Contemporary methods have to accommodate broader evolutionary distances and work under a far more varied set of real-world conditions.
Looking back, we think one of the clearest lessons from this project is that useful methods are not defined only by their best result. They are also defined by whether they can meet the needs of the field at the moment they are built. For gene annotation, those needs now include scale, robustness, and the ability to work across many different types of genome. Ultimately, that is the space in which ANNEVO was developed. We wanted to build a method that, without compromising accuracy, could make gene annotation a more practically achievable task – and as an ever larger number of genome assemblies become available, we hope tools like ANNEVO can help researchers move more effectively from raw sequence to actionable biological insight.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in