Decoding the Water Lily: A Tale of Mud, Megabytes, and the Tragic Fate of the World's Smallest Lily
Published in Agricultural & Food Science
When most people look at a water lily, they see tranquility. They see Monet paintings, serene koi ponds, and the gentle beauty of nature. When our research team looks at a water lily, we see 50 million years of evolutionary secrets, hundreds of gigabytes of sequencing data, and a truly unreasonable amount of mud.
Recently, our team (Led by Prof. Fei Chen from Hainan University) published a comprehensive genomic study on the genus Nymphaea in Nature Plants. The final paper is neat, tidy, and full of elegant charts. But the journey to get there? That was a beautiful, chaotic, and sometimes heartbreaking mess.
Our aim
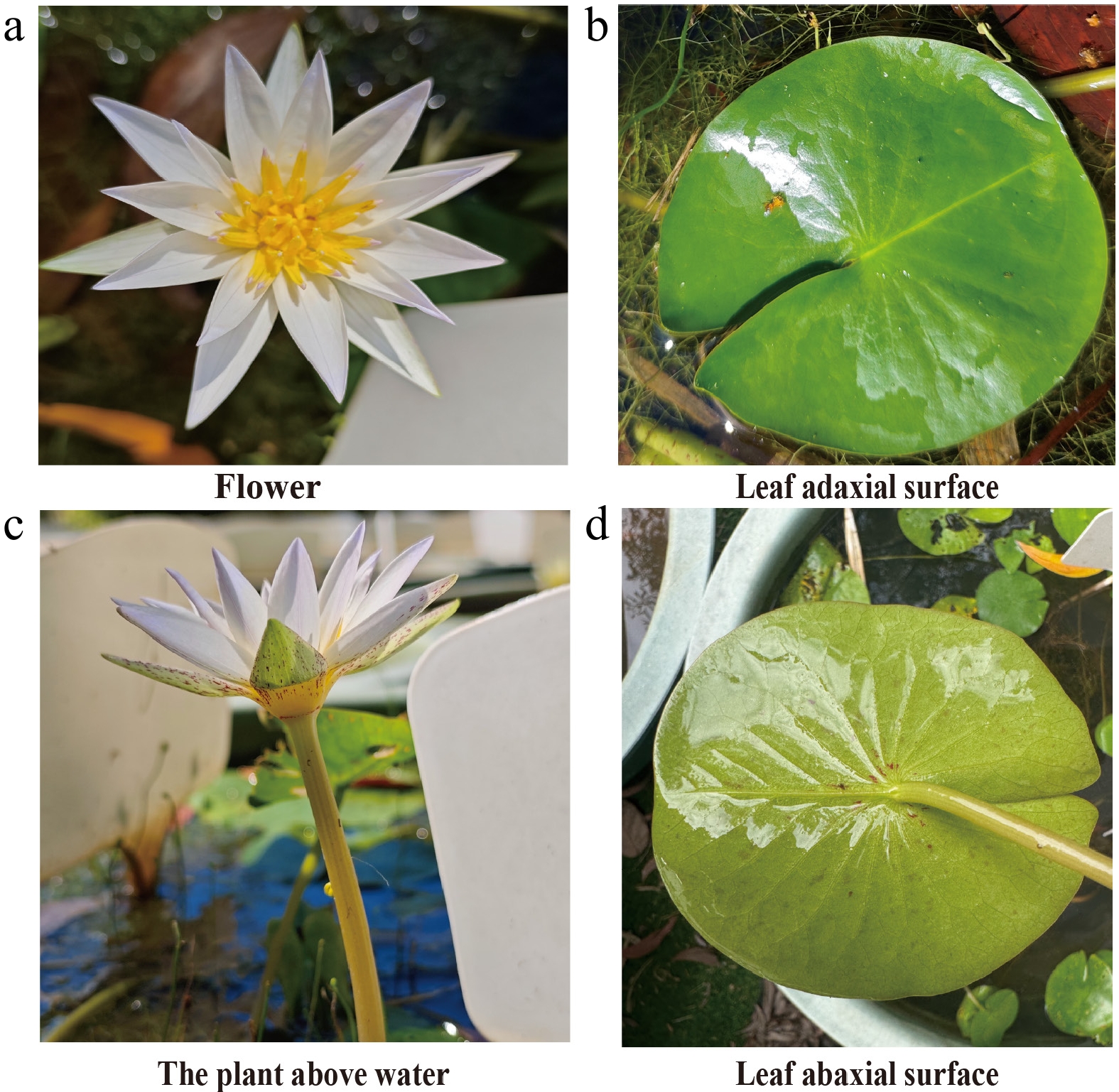
Our primary goal was simple on paper but daunting in reality. Water lilies are among the most basal groups of angiosperms and retain many morphological and physiological traits of early angiosperms, making them invaluable for studying angiosperm evolution, particularly floral organ development. We wanted to understand the molecular mechanisms that drove the rapid diversification of early flowering plants—specifically focusing on how they developed their stunning colors and complex scents.
Our approach: Building the "Holy Grail" of Genomes
To really understand the genetics of these plants, we couldn't just skim the surface. We needed complete, highly accurate genomes. We selected three species as our primary optimal candidates: N. colorata, N. thermarum, and N. caerulea.
To make this happen, we didn't just grab plants from a local pond. The genomes of N. colorata, N. caerulea, and N. thermarum were derived from highly homozygous inbred lines through multi-generational selfing. Then came the heavy lifting. We utilized PacBio HiFi sequencing, Oxford Nanopore Technologies (ONT) ultra-long reads, and Hi-C data.
If you aren't a genomicist, just know this: it is an absolute ocean of data. We are talking about mapping billions of letters of DNA. After rigorous filtering and scaffolding, we achieved something incredibly rare and satisfying: gap-free genome assemblies for the three water lily species. No missing pieces. No mysterious black holes in the data. Just pure, continuous genetic code.
An unexpected finding: The Angiosperm "Pollen Drill"
Once we had the genomes, we started comparing them to other plant lineages. We discovered 885 orthologous gene families exhibiting significant expansion in angiosperms. But one finding stood out.
Unlike gymnosperms where pollination occurs via liquid droplets, angiosperm fertilization requires the growth of pollen tube through the pectin-rich extracellular matrices of stylar tissues. How did early flowers pull this off? We found an angiosperm-exclusive pectin lyase gene specifically expressed during pollen tube elongation. Essentially, flowering plants evolved a microscopic, enzymatic "drill" that facilitates this process through targeted degradation of middle lamellae, thereby enabling efficient pollen tube penetration.
Solving the Mystery of the Blues
Have you ever wondered why blue flowers are relatively rare in nature? In the relatively uncommon case of blue-purple flowers, pigmentation is primarily attributed to derivatives of delphinidin and cyanidin. We wanted to find the master switch for this color.
Through a mix of targeted metabolomics and transcriptome-wide network analysis, we hit the jackpot. We identified the transcription factor NcolMYB75-like as a master regulator of blue anthocyanin biosynthesis. When we overexpressed this gene in mutant Arabidopsis plants, it significantly promoted anthocyanin accumulation. We literally found the genetic paintbrush that gives the water lily its iconic blue hues.
The Heartbreak: The Scent of Extinction
But research isn't all "aha!" moments and pretty colors. Sometimes, the genome tells a tragic story.
Floral scent is a critical adaptive innovation in angiosperm pollination systems. We found that the expansion and diversification of the O-methyltransferase (OMT) gene family drive the synthesis of species-specific floral scent volatiles. For example, N. colorata and N. caerulea use these OMT genes to emit methyl phenols, which attract specific pollinators.
Then there is N. thermarum, a diminutive and highly endangered water lily. When we profiled its floral scent, we found zero methyl phenols. Why? Our biochemical tests revealed a heartbreaking genetic reality: the OMT expressed in N. thermarum lacks catalytic activity, indicating that the loss of OMT function is the key reason for the absence of methyl phenols in its floral scent.
Because it lost its signature perfume, the plant couldn't attract the insects it needed. The absence of methyl phenol in the floral scent of N. thermarum may contribute to its limited pollinator visits, leading to a reliance on self-pollination. While self-pollination works in a pinch, it is an evolutionary dead end in the long run. The reliance on self-pollination in N. thermarum led to decreased genetic variation, ultimately contributing to their extinction in the wild.
It’s a sobering reminder hidden in the data. A single inactive enzyme, a lost fragrance, and a species quietly fades from its natural habitat.
Looking Forward
Our gap-free genomes and findings on floral traits provide a genomic framework for plant breeding and ecological conservation. We hope this data will not only help create more resilient crops and beautiful ornamentals but also aid in conserving fragile species like N. thermarum before it's too late.
So, the next time you admire a water lily, take a moment to appreciate the complex genetic machinery, the millions of years of evolutionary trial and error, and the sheer volume of muddy boots and late-night data processing it took to understand it all.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in