DNA Star as a Viral Sensor and Inhibitor
Published in Chemistry
Viruses are infectious molecular assemblies with a diameter of 20–250 nm. Many of these viruses exhibit exquisitely symmetric organization and present a unique pattern of capsid proteins on their surfaces [1]. The specific and repeated patterns of capsid proteins facilitate multivalent binding to host cells, resulting in enhanced viral infectivity. Based on this naturally occurring multivalent binding mechanism, synthetic multivalent entry blockers were introduced in the 1990s by Whitesides and other groups [2, 3, 4]. Ligands that bind capsid proteins can be linked with a scaffold to create significantly more potent and soluble multivalent inhibitors. However, existing scaffolds, which include peptides, proteins, polymers, dendrimers, nanoemulsions and inorganic nanoparticles, may be toxic, imprecise in ligand spacing, and limited in scaffold shape and ligand valency.
Previously, we trapped influenza viruses using a dendrimer-based spherical nanoparticle with terminal sialyllactose ligands. Both dendrimer size and ligand spacing distance were found to be critical in inhibiting viral infection. Specially, a spacing of 3 nm between ligands resulted in the strongest binding to hemagglutinin (HA) trimers [5]. However, the synthetic polymer was unlikely to be accepted as a therapeutic because of its possible toxicity. Unlike dendrimers and other polymeric scaffolds, non-toxic, biodegradable, and biocompatible DNA can be customized to possess precise geometric shapes and designed to contain a pattern-recognizing display of multiple ligands.
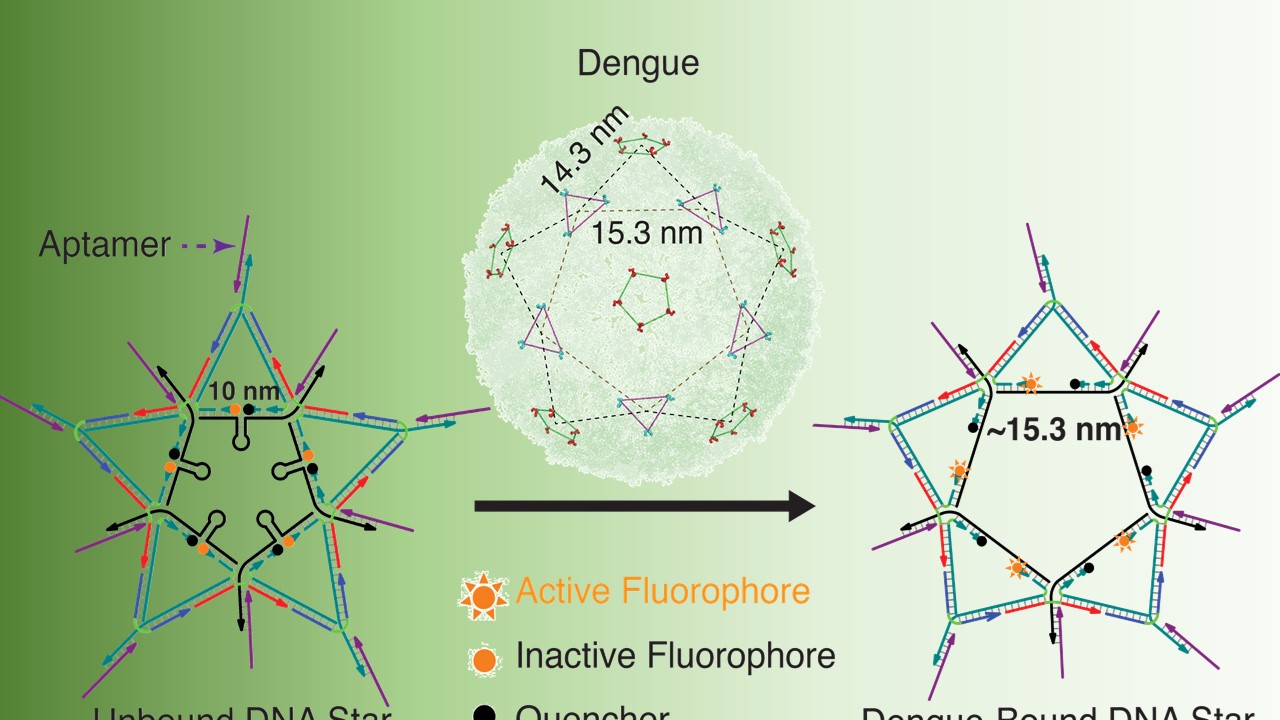
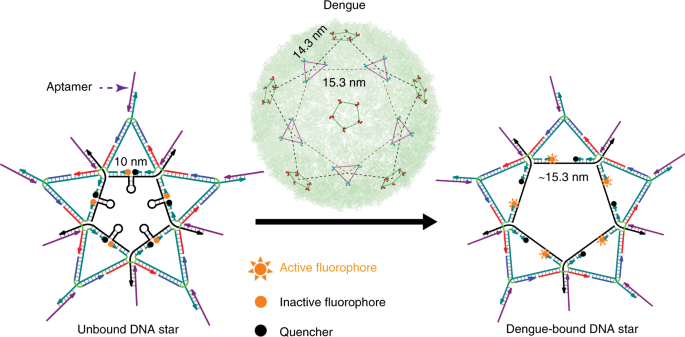
The mosquito-borne dengue virus (DENV) that causes dengue fever presents a complex geometric “star-shaped” pattern” that cannot be addressed by existing scaffolds that are not precise in ligand spacing or provide limited control over scaffold shape and ligand valency. Using DNA nanotechnology, we designed a star-shaped DNA architecture with attached specific aptamers targeting dengue envelope protein domain III (ED3) by superimposing the capsid patterns of the virus. This “DNA star” precisely matched the spatial arrangement of ED3 clusters on the DENV surface, enabling rapid fluorescent detection of specific pattern-matched dengue viruses. Multivalent interactions resulted from the pattern-matched binding and provided high DENV-binding avidity. Hence, the DNA star can be used as a strong multivalent inhibitor, which was approx. 7,500 times more effective than the monovalent inhibitor for the in vitro inhibition of DENV infection in human serum [6].
As a result, we discovered that a pattern-recognizing display of multiple ligands spaced with nanoscale precision on a specific scaffold was critical in designing potent inhibitors and sensors against infectious viruses. Our molecular-platform design strategy could be adapted to detect and combat other disease-causing pathogens by generating the requisite ligand patterns on customized DNA nanoarchitectures.
Follow the Topic
-
Nature Chemistry
A monthly journal dedicated to publishing high-quality papers that describe the most significant and cutting-edge research in all areas of chemistry, reflecting the traditional core subjects of analytical, inorganic, organic and physical chemistry.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
Very interesting paper!
I have two questions about it:
1) Where did the idea of using a star-shaped DNA come from? Did you decide to use a star-shaped structure because the geometry would have allowed you to "multivalently arrange binders in a 2D pattern mirroring the complex spatial distribution of dengue epitopes with nanometre precision"? Exactly which properties of this star-shaped DNA structure allowed you to do so?
2) To which other species of viruses is the application of this technique most amenable? For example, would it be most easily adapted for other flaviviruses? What about viruses of other species?
1) As shown in Figure 1a in the article, we found the star-shaped ED3 sites (target) where the aptamer (ligands) can bind to.
2) You can apply this technology to any viruses.
Dear Kwon:
Great work, which company manufactured the star shape? and did you apply this technology to other viruses? and do you have coming publications on this topic?
take care
1) We designed the star-shaped structure using DNA oligomers.
2) We can design any shaped and patterned 2D and 3D scaffolds. Now, we will design the specific structures for targeting influenza viruses.