ESP: A Machine Learning Model for the Prediction of the Substrates of Enzymes
Published in Computational Sciences
In our recent paper, we developed a model for predicting substrates for enzymes with yet unknown function; we named this enzyme-substrate prediction model ESP. In this blog post, we aim to describe the components of ESP in an easily understandable way. First, we will briefly describe what enzymes are and what is known about them. Second, we will focus on two important ingredients for developing a prediction model: generating training datasets and numerically representing information about the prediction task. Third, we describe the prediction model and its capabilities, before finally focusing on potential applications of our model.
What are enzymes and how do they function?
Enzymes play a vital role in all forms of life on Earth, ensuring that essential chemical reactions occur at the right rate. They are a type of protein that acts as a catalyst, speeding up chemical reactions without being altered or consumed. Each protein consists of a long sequence built from 20 different amino acids; on average, each sequence consists of a few hundred amino acids that are arranged in a precise order. These sequences fold into 3D structures that determine the function of the proteins.
How do enzymes catalyze reactions? Most enzyme-catalyzed reactions can occur even without enzymes, but at a much slower rate and with much less control. To understand the mechanism by which enzymes catalyze reactions, we can take a simplified view and imagine a lock-and-key model. Enzymes have a specific pocket in their 3D structure called the active site that acts like a keyhole. The substances consumed in the chemical reaction are called substrates, and they fit into the active site much like a key fits into a lock. This substrate-enzyme binding greatly accelerates the conversion of substrate molecules into product molecules, up to a million times faster than spontaneous rates in the absence of enzymes. When the reaction is complete, the products are released, and the enzyme is ready for another cycle of catalytic activity.
What do we know about the function of enzymes? Typically, each enzyme has evolved to catalyze a specific chemical reaction. Consequently, most organisms contain thousands of different enzymes to facilitate a large number of different reactions. Unfortunately, we do not know the function of the vast majority of enzymes: Although millions of different enzyme sequences have been identified, the exact function for more than 99% of them is unknown. Prediction models that are capable of identifying candidate substrates for enzymes with yet unknown function are therefore highly desirable.
Numerical representations for enzymes and substrates as input for machine learning models
Since machine learning models are mathematical functions, they require numerical inputs with information about the prediction task. Thus, in our efforts to develop a powerful machine learning model capable of predicting substrates for enzymes, we faced the challenge of representing complex biological information numerically: we needed to encode both enzyme and substrate information into numerical vectors.
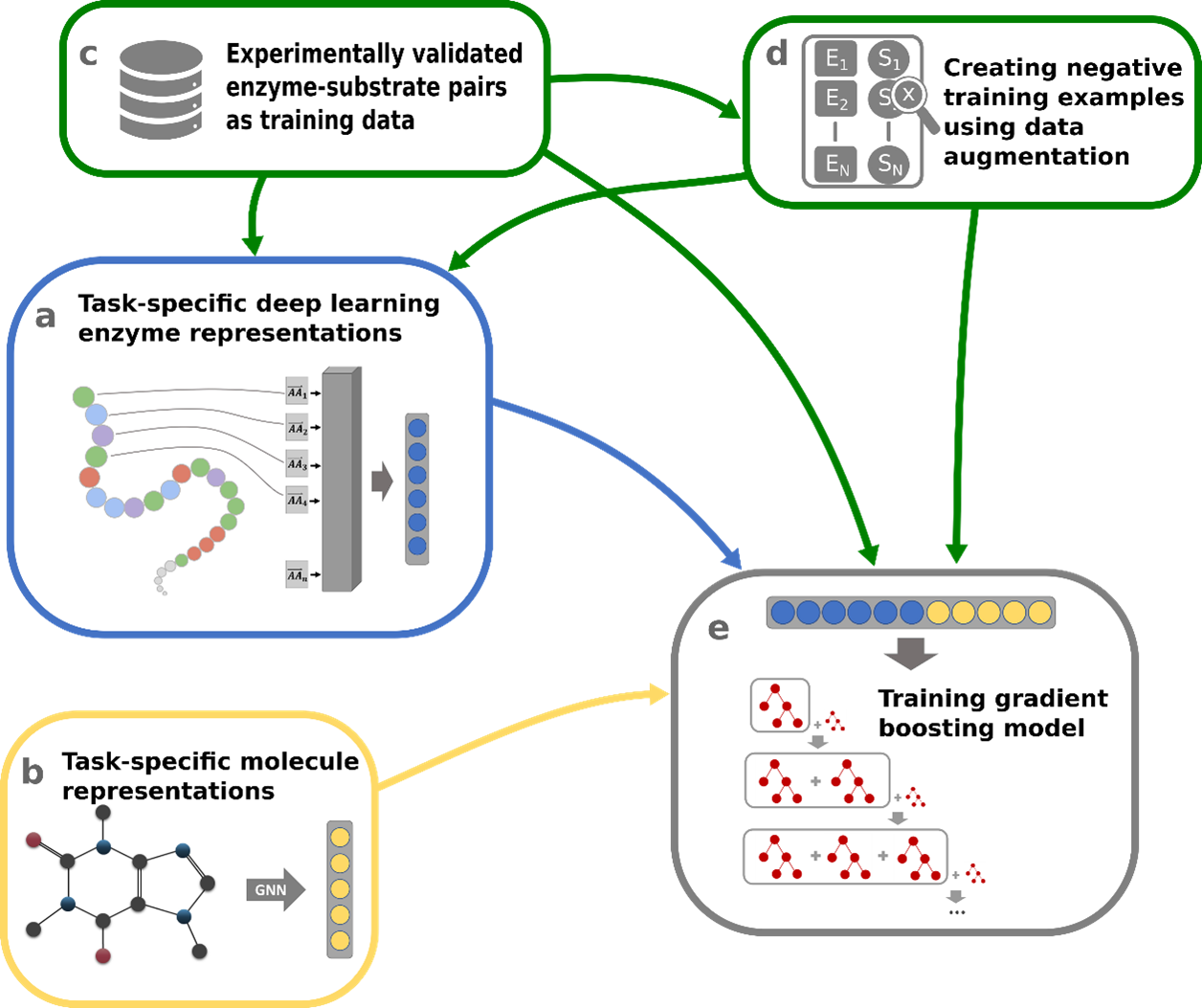
Numerical encoding of enzymes: Given the wide availability of amino acid sequences for millions of enzymes, we decided to use this information to numerically encode enzyme properties. We used Transformer Networks, which were originally developed for Natural Language Processing (NLP) applications such as language translation or text generation. This model architecture also proved to be highly effective when applied to the amino acid sequences of proteins, allowing us to encode valuable information about the structure and function of the protein. Specifically, we used the ESM-1b protein Transformer Network - a Transformer Network trained by the Facebook AI Research (FAIR) team on nearly 30 million different protein sequences – and we fine-tuned it for our prediction task (Figure 1a).
Numerical encoding of small molecules: Substrates of enzymes are typically small molecules composed of atoms connected by chemical bonds. To capture the structural information of small molecules numerically, we represented each substrate as a graph. Graphs consist of nodes connected by edges, making them an ideal way to capture the structural arrangement of atoms and bonds within a molecule: the atoms of a molecule were interpreted as nodes of a graph, and the bonds of a molecule were interpreted as edges of a graph. We converted this graph information into a single numerical vector with substrate information. This conversion was done using Graph Neural Networks (GNNs), deep neural networks specialized for processing graph-structured data (Figure 1b).
Data for training a machine learning model
While we have successfully generated numerical representations for both enzymes and potential substrates, we still lack a critical ingredient: large amounts of high-quality data to train our machine learning model. Training a reliable model requires a diverse dataset that includes many known enzyme-substrate interactions as well as negative enzyme-small molecule pairs, where the small molecule is not a substrate for the enzyme. We successfully extracted about 15,000 experimentally validated enzyme-substrate pairs from the GO Annotation database (Figure 1c). However, information on negative enzyme-small molecule pairs is rarely published in protein databases, and thus our dataset lacks such negative data.
Leveraging the specificity of enzymes: To overcome this obstacle, we took advantage of the fact that the vast majority of small molecules in living cells do not act as substrates for a particular enzyme. We used this information to our advantage: by sampling small molecules from a set of over 1,400 different small molecules that are typically present in biological cells, we generated negative enzyme-small molecule pairs that have a high probability of actually being true negatives (Figure 1d). This approach allowed us to accumulate a significant amount of negative data points, enabling our model to learn from both positive and negative instances.
Performance and limitations of the enzyme-substrate prediction model
We used this compiled dataset with a total of 70,000 data points together with the numerical representations of the enzyme and the potential substrate to train a machine learning model - the Enzyme-Substrate Prediction model, which we named ESP. Let's take a look at the architecture of the ESP model, its performance, and its limitations.
The power of Gradient Boosting models: At the heart of ESP is a gradient boosting model, a state-of-the-art machine learning technique (Figure 1e). This model is an ensemble of multiple, iteratively built decision trees, each contributing to the final prediction. By combining the predictions of different trees, the model can process complex data to make accurate predictions. We put ESP to the test by evaluating it on an independent test set of thousands of data points that it never encountered during training. ESP achieved over 91% accuracy in determining whether a given small molecule was a true substrate for a given enzyme.
Performance for unseen data: ESP is able to perform well on enzymes that do not closely resemble any of the training enzymes. This ability allows the model to adapt to different enzymes across the biological landscape. While ESP shows high overall accuracy and good generalization capabilities to unseen enzymes, it has some limitations: Currently, its prediction quality is limited when dealing with (potential) substrates that were not part of the training set. Fortunately, our training set includes a large set of over 1,400 different substrates, allowing the model to make good predictions for this substantial set of molecules.
Potential applications for the ESP model
Having a model that can predict potential substrates for enzymes with unknown functions is valuable in several scientific and practical applications.
Drug discovery and development: In the pharmaceutical field, predicting potential substrates for enzymes can be important. Enzymes play critical roles in various biological processes, including metabolism and cell signaling. Identifying small molecules that can interact with specific enzymes as substrates can lead to drug development. For example, these substrates can serve as a starting point for the discovery of drugs that inhibit the protein.
Biotechnology and Enzyme Engineering: Enzymes are the workhorses of many biotechnology processes. Predicting substrates for enzymes of unknown function can expand the toolkit for enzyme engineering and bioengineering. This can enable researchers to optimize enzymes to perform specific tasks, such as producing biofuels, synthesizing valuable chemicals, or degrading environmental pollutants.
Unraveling biological pathways: The metabolic networks of biochemical pathways in living organisms are still not fully understood. Predicting substrates for enzymes of unknown function can provide valuable insights into these complex pathways. By deciphering which small molecules interact with specific enzymes, researchers can gain further insight into metabolic pathways and cellular metabolism.
Follow the Topic
-
Nature Communications

An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Mar 24, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in