Geothermal Survivor: Bacteria from Mexican Hot Springs Holds Key to Heavy Metal Cleanup
Published in Earth & Environment, Research Data, and Agricultural & Food Science

Adaptive Genomic Features of Raoultella ornithinolytica LAM1
from the Geothermal Site of Los Azufres Reveal Potential
for Heavy‑Metal Bioremediation
https://link.springer.com/article/10.1007/s00284-025-04507-4
Introduction
It's not every day that you scoop a water sample from a steaming, sulfur-scented hot spring and end up with a potential solution for heavy metal pollution. But that's exactly what happened when we visited the Los Azufres geothermal field in Michoacán, Mexico.
This place is extreme, even by microbial standards. Nestled within the Mexican Volcanic Belt, Los Azufres is a landscape of fumaroles, acidic ponds, and waters so rich in toxic elements that they would be fatal to most forms of life. Previous studies had documented alarming levels of arsenic, lead, mercury, and uranium in the area, concentrations far exceeding safety limits for drinking water. Yet, life finds a way. And in one of those metal-laden ponds, we found ours.
We weren't just looking for any bacterium. We wanted to understand how microorganisms survive, and thrive in environments that would kill most others. Could their survival strategies be harnessed to clean up the mess we humans have made with industrial pollution?
That's how we met LAM1.
Isolated from an acidic geothermal pond, this strain of Raoultella ornithinolytica turned out to be something special. Not only did it grow happily in arsenic concentrations that would make us run for the hills, but it also seemed unbothered by a cocktail of other heavy metals. When we sequenced its genome, the secrets began to unfold.
What we found was a genetic toolkit for survival: complete operons for arsenic detoxification, multiple copies of mercury resistance genes, and sophisticated systems for handling copper, zinc, and nickel. It was as if this bacterium had evolved its own miniature wastewater treatment plant.
This is the story of how we discovered Raoultella ornithinolytica LAM1, what its genome revealed about life at the edge, and why we believe this tiny geothermal survivor might hold a big key to bioremediation.
Behind the Paper: What We Found in the Genome of a Geothermal Survivor
When we first isolated Raoultella ornithinolytica LAM1 from a steaming, metal-rich pond in Los Azufres, we knew it was tough, but we had no idea just how well-equipped it really was.
The first clue came from the petri dish. While other bacteria curled up and died at arsenic concentrations above a few hundred ppm, LAM1 kept growing happily at 1500 ppm sodium arsenate, that's 54,000 times the WHO limit for drinking water. We needed to understand how.
So we sequenced its genome.
A Genetic Toolkit for Extreme Survival
The numbers alone were impressive: a 6.01 Mb genome with 5,744 genes. But the real story was in the details. We found 99 genes dedicated to heavy metal resistance: arsenic, mercury, copper, zinc, cobalt, cadmium, nickel, and lead. This wasn't just a bacterium with a single trick; it was a walking, dividing detoxification factory.
The arsenic system caught our attention first. LAM1 carries a complete ars operon (arsR-arsB-arsC-arsA-arsD), but with a twist: three copies of arsC, the gene that reduces arsenate to the more exportable arsenite. Having three paralogs scattered across different contigs suggests gene duplication or horizontal transfer—nature's way of doubling down on what works.
Mercury? Four copies of merA. Copper? A complete system including copA, cueO, and cusA. Nickel? The full nikABCDE-nikR uptake and homeostasis system. Zinc and cadmium? zntA and zntR were right where we expected them.
Surprises Along the Way
Not everything was predictable. When we classified the 99 resistance genes by metal type, zinc-related genes came out on top (27.9%), followed by nickel and copper. But functional enrichment analysis told a different story: arsenic metabolism was the most statistically significant pathway, despite representing only 16.3% of the genes. This duality reflects both the specialized arsenic detoxification needs of geothermal environments and the cross-cation promiscuity of many transporters.
We also found unexpected players: sulfur metabolism genes (dsrB) that likely help LAM1 handle the hydrogen sulfide-rich waters of Los Azufres, and antibiotic resistance determinants that let it grow on CHROMagar ESBL. This co-occurrence of metal and antibiotic resistance genes raises fascinating questions about co-selection pressures in extreme environments.
How Does LAM1 Compare?
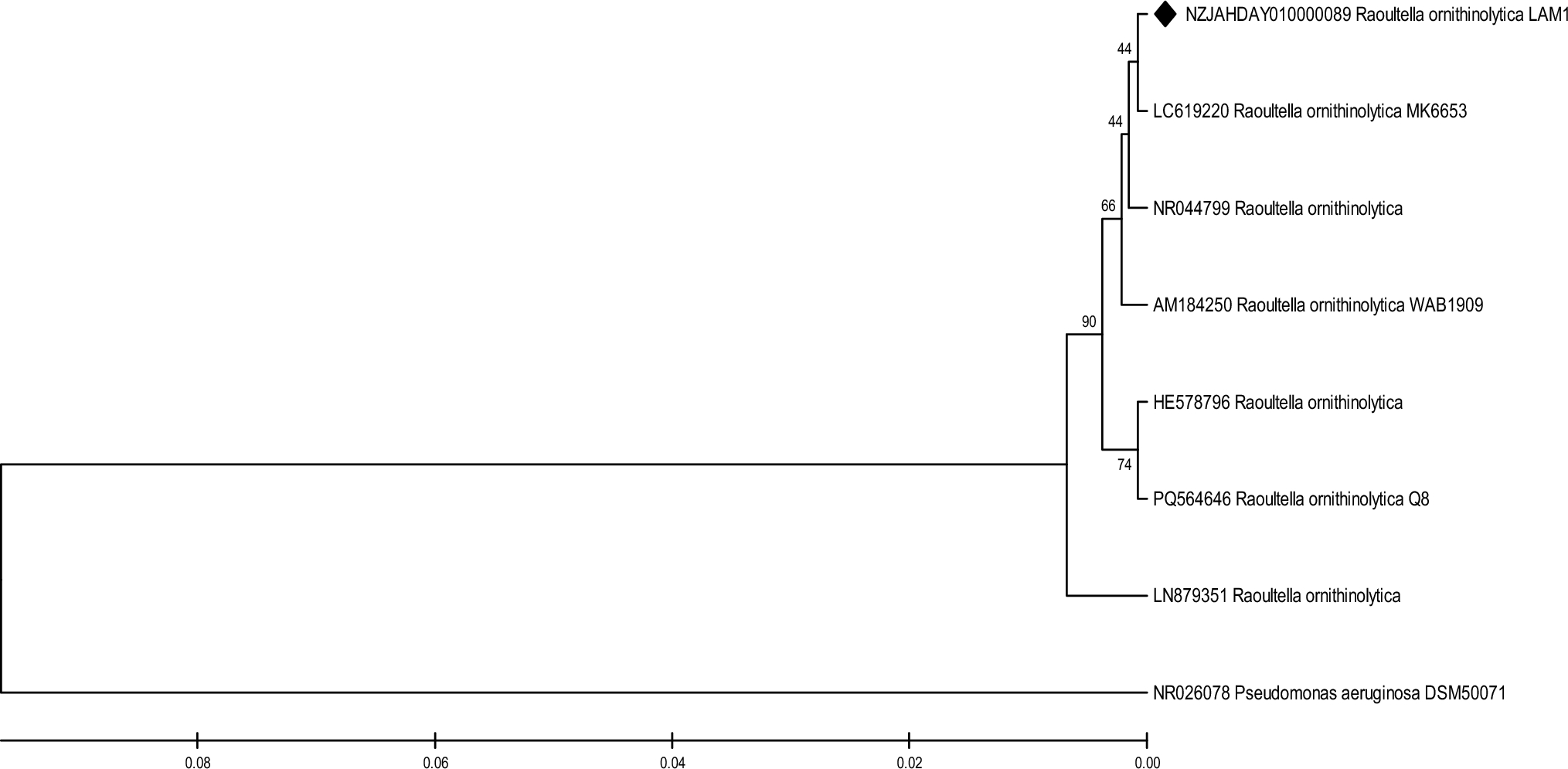
We compared LAM1's genome against 508 publicly available R. ornithinolytica genomes. The differences were striking. Clinical isolates like WM1 carry antibiotic resistance islands but lack heavy metal genes. Wastewater strains like MQB_Silv_108 show some metal tolerance, but none approach LAM1's 1500 ppm arsenic limit. Even lead-resistant R. planticola FACU 3 taps out well before LAM1's threshold.
The message is clear: LAM1 is special. Its expanded ars operon, quadruple merA genes, and geothermal adaptations like dsrB reflect a bacterium that has spent generations evolving in one of Mexico's most chemically challenging environments.
What It All Means
LAM1 isn't just a curiosity, it's a genetic resource. The same genes that help it survive Los Azufres could be harnessed to clean up mining waste, industrial runoff, or arsenic-contaminated water. And as we face growing heavy metal pollution worldwide, having a naturally optimized solution feels like finding a key just when you need it.
But we're not done. Next steps include validating which genes are actively expressed under metal stress, dissecting their regulatory networks, and exploring how LAM1's toolkit might be engineered for real-world applications. From geothermal hot springs to polluted industrial sites and who knows, maybe even beyond, this tiny bacterium has a lot to teach us.
Conclusions
Raoultella ornithinolytica LAM1 stands out as a genomically equipped extremophile, adapted to thrive under the unique physicochemical stressors of geothermal environments. Its genome reveals a suite of specialized traits, including metal resistance, sulfur metabolism, and acid tolerance, that reflect ecological adaptation to the conditions of Los Azufres. Beyond individual resistance operons, the broader genomic landscape exhibits functional redundancy and plasticity indicative of adaptation to chronic environmental pressures. The strain's potential role in nutrient cycling and its apparent cross-resistance to antibiotics raise important ecological and biosafety considerations. Altogether, LAM1 bridges fundamental microbial ecology with applied environmental biotechnology, offering a promising platform for bioremediation and synthetic biology applications under extreme conditions. Looking ahead, LAM1's unique genomic repertoire provides a valuable framework for understanding microbial resilience in chemically dynamic ecosystems and for guiding the development of engineered strains capable of operating in harsh environments, from polluted industrial sites to potential extraterrestrial habitats. As a model for extremophile adaptation, LAM1 may help unravel the evolutionary strategies that support survival under sustained environmental stress, while inspiring novel solutions for environmental restoration and sustainable biotechnologies.
Follow the Topic
-
Current Microbiology
Current Microbiology is a renowned scientific journal committed to advancing Microbiology. It delves into the realms of prokaryotic and eukaryotic cells, viruses, and the intricate interplay among microorganisms, hosts, and their environment.
Related Collections
With Collections, you can get published faster and increase your visibility.
Novel Prokaryotic Taxa
Publishing Model: Hybrid
Deadline: Ongoing


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in