Global predictions for the risk of establishment of Pierce’s disease of grapevines
Published in Ecology & Evolution

Pierce's disease (PD) of grapevines is a vector-borne disease produced by the bacterium Xylella fastidiosa (Xf) and nowadays is considered a potential major threat to winegrowers worldwide.
Although it may seem unbelievable now, this is the only information I knew about Xf and PD at the time of starting this project. The journey began almost two years ago, when I was a recently graduate physicist and had just completed my master in Physics of Complex Systems. Thanks to the "open-mindedness" of biologists like Eduardo Moralejo, entomologists like Alberto Fereres and physicists like Jose Ramasco and Manuel Matías, both professors from the master, this interdisciplinary work was indeed possible.
Perhaps the first thing I learnt from the pathosystem under study was that PD is a thermal-sensitive disease, so that temperature is a range-limitig factor. Indeed, the starting point of the project was some inoculation assays that were performed to 36 grapevine varieties, which were intended to elucidate the effect of temperature on the symptom development process. This, together with previously measured data of Xf growth as a function of temperature, allowed us to develop a mechanistic framework capable to describe the symptom development process only from temperature data. The general idea is that, by knowing how fast bacteria grow at different temperatures, one can compute an approximation to the bacterial load of infected plants over a given time period if temperature data is available. Then, thanks to the inoculation assays, this approximated bacterial load can be translated into a probability of developing symptoms.
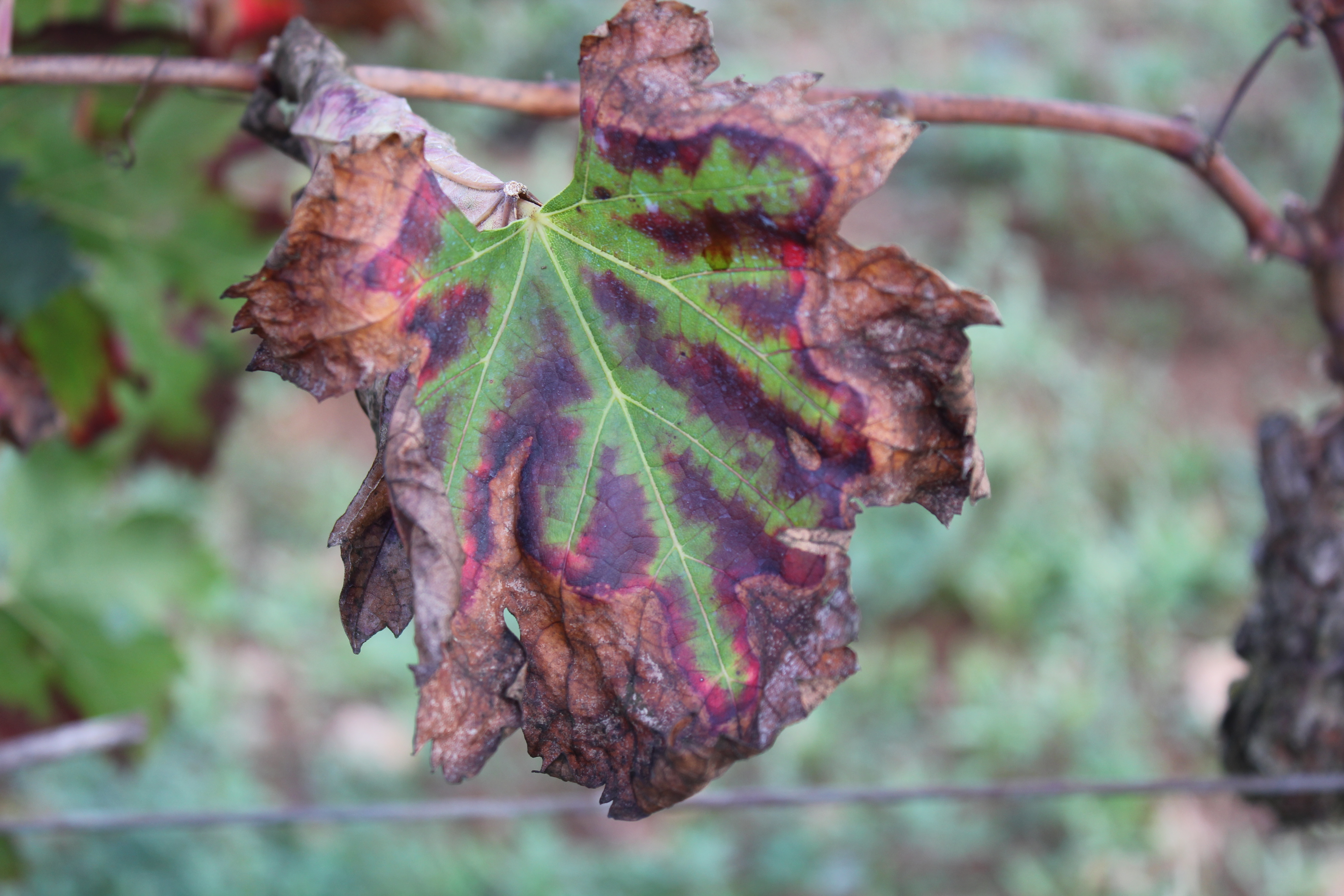

However, this is not the end of the story, as cold enough temperatures can then kill bacteria from infected plants! PD is very rarely found in zones in which the minimum average temperature of the coldest month is below -1.1ºC, limiting its geographic expansion in the US. We then used this information to relate cold accumulation to plant survival, or what is the same, to the probability of recovering from infection.
By combining symptom development and recovery, we finally developed a unified framework to model where PD can thrive or not given a particular weather. In essence, we just built a mechanistic model of climatic suitability for Xf within its grapevine hosts.
Up to this point, it seemed that all was done to model the risk of establishment of PD. However, we still missed something: disease transmission. In short, there will not be any risk of having the disease in a place where transmission from infected to susceptible plants cannot occur, even if the weather is perfectly suitable for the bacteria. To take this into account, we developed a compartmental model of disease transmission, which is customary used in mathematical studies of epidemic spreading. The most important feature of this model is that the more abundant the vector is in a given zone, the easier it will be for the disease to spread.
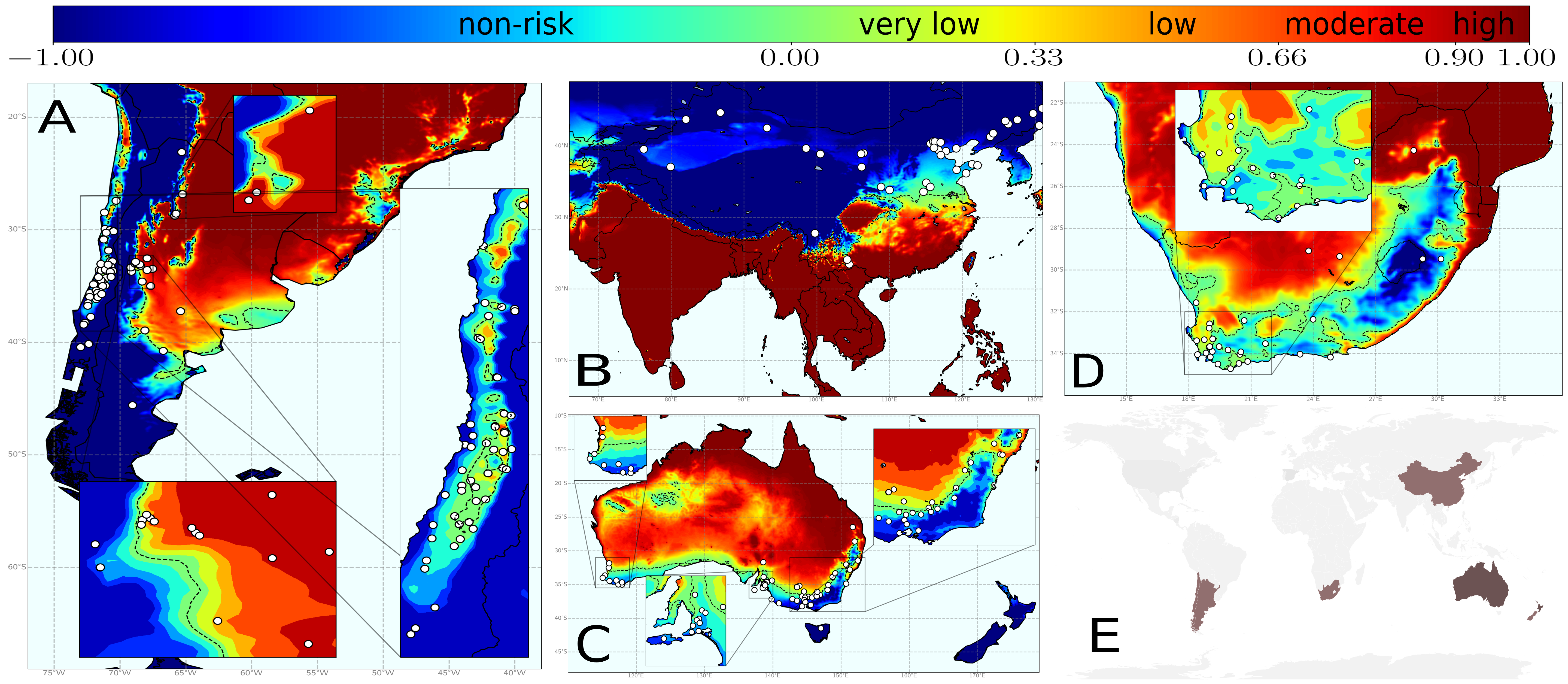
Finally, we had all the pieces to develop our complete model on PD risk. Information on the vector abundance allow us to know how many new infected plants will an infected plant contribute to infect, while temperature data allow us to know how many of these new infections will indeed thrive. So only if this balance is positive, an outbreak will occur. Nevertheless, information on vector abundance was not available for all wine-producing zones worldwide. In such cases, we selected a reasonable transmission scenario as a homogeneous baseline, i.e. the same mean value of the vector abundance is used everywhere (Fig 2).
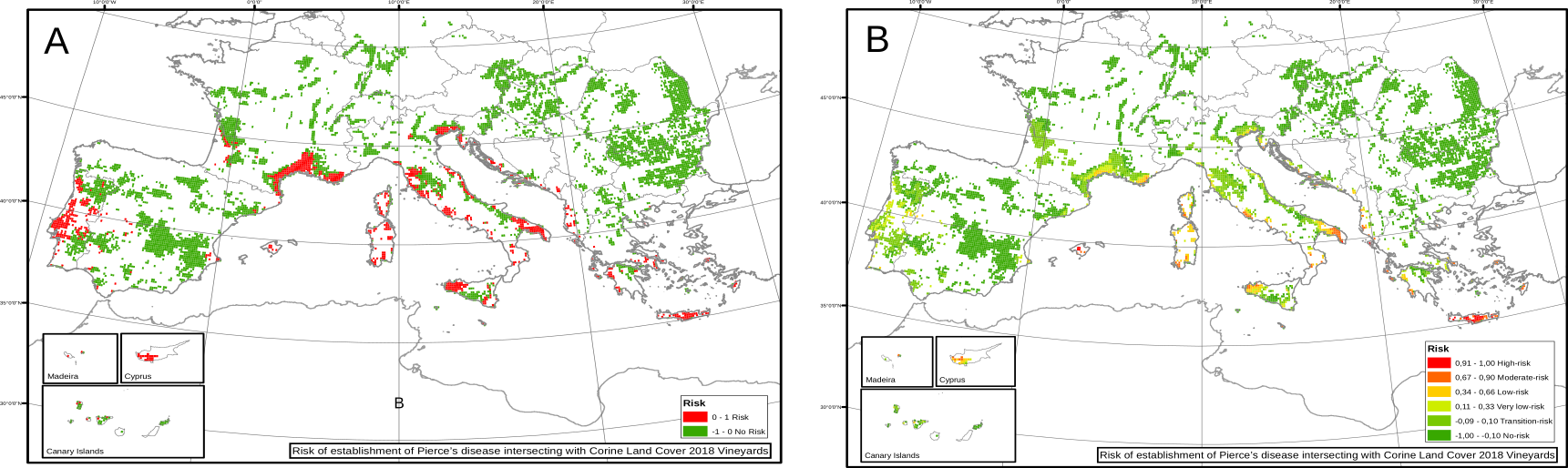
In the particular case of Europe, we had estimations of the spatial distribution and abundance of the main vector, Philaenus spumarius. Results show a drastic lower risk than compared with the homogeneous baseline case. Using the spatial heterogeneous distribution of the vector makes clear that some climatic suitable zones for Xf have not enough vector to allow for its spread (Fig 3).
The global application of our model gave rise to many interesting results. We developed a simple webpage to show all these results that can be obtained with our model in an interactive mode. Go to check it out!
I would like to conclude with some thoughts on which I think is the most interesting feature of our work. Although our results are quite accurate, e.g. precisely representing the actual distribution of Xf-affected zones in Europe, this is not for me the best part of our work. Perhaps as a physicist, I mostly like to admire the theoretical consistency and meaningfulness of some models, which I think is a strong point of ours. Although some other correlative models (e.g. AI) could perhaps equal the accuracy and predicting power of our mechanistic model, the fact that we can explain the fundamental factors that allow or not the expansion of PD is priceless.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
From RNA Detection to Molecular Mechanisms
Publishing Model: Open Access
Deadline: May 05, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in