Go CO2 free
Published in Microbiology and Protocols & Methods
The growth of all bacteria in a laboratory requires specific environmental conditions, including the nutrients in the media, the temperature, and the atmosphere. Much of the media that we use today was created decades ago to provide the needed nutrients to bacteria of interest.
Temperatures for bacterial growth cultures are generally set by the temperature of the natural environment in which the bacteria are found; for this reason, most pathogenic bacteria are grown at human body temperature, 37°C.
Lastly, bacteria need to have the right atmospheric environment to be able to grow in a lab; this also often mimics the conditions where the bacteria is found in nature. Some bacteria grow fine in the lab in a normal atmosphere. Some bacteria require there to be no oxygen, an anaerobic atmospheric environment like would be found in the human gut. Other bacteria need something different, such as the bacteria that cause gonorrhoea, Neisseria gonorrhoeae; these need enhanced levels of CO2 to be able to grow in the lab.

Due to this requirement for CO2, special incubators are often used to grow N. gonorrhoeae that have regulators to add CO2 from gas tanks into the atmosphere within the incubator. Growth like this requires there to be a readily available supply of CO2 gas and for there to be the special incubators able to use this gas.
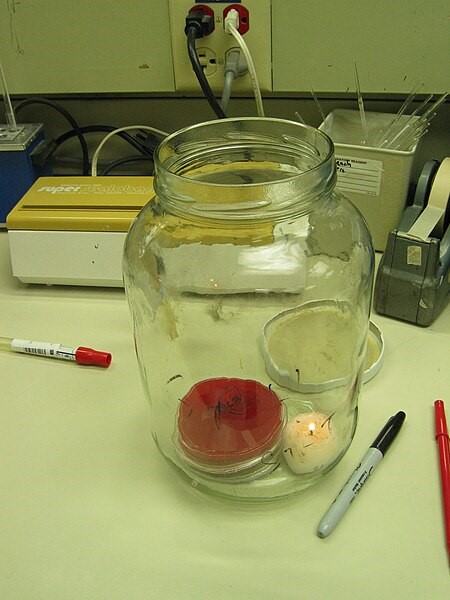
There are some alternatives, such as putting the agar plates with the bacteria into sealed jars with a small candle to deplete oxygen and thus raise CO2 levels in the jar. But, these jars then take up additional space in an incubator, where space can be precious.

It would be so much easier, and more readily accessible to remote and resource limited areas, to be able to grow Neisseria gonorrhoeae and bacteria like it on media that eliminates the need for CO2.
This is what we have done in our paper, ‘Improved Neisseria gonorrhoeae culture media without atmospheric CO2.’ In Applied Microbiology and Biotechnology (10.1007/s00253-025-13449-7).
During the pandemic, there were shortages in available CO2 gas cylinders. Even outside of the pandemic, CO2 cylinders run out of gas, ruining experiments and analysis of samples. In remote areas and resource limited areas, there can be issues trying to grow bacteria such as Neisseria gonorrhoeae that have non-standard atmospheric requirements.
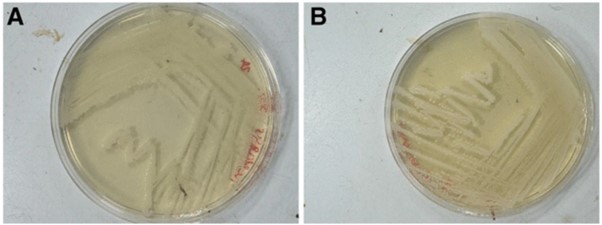
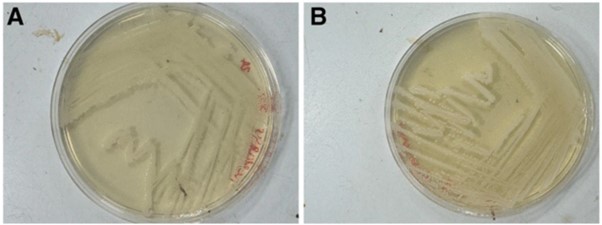
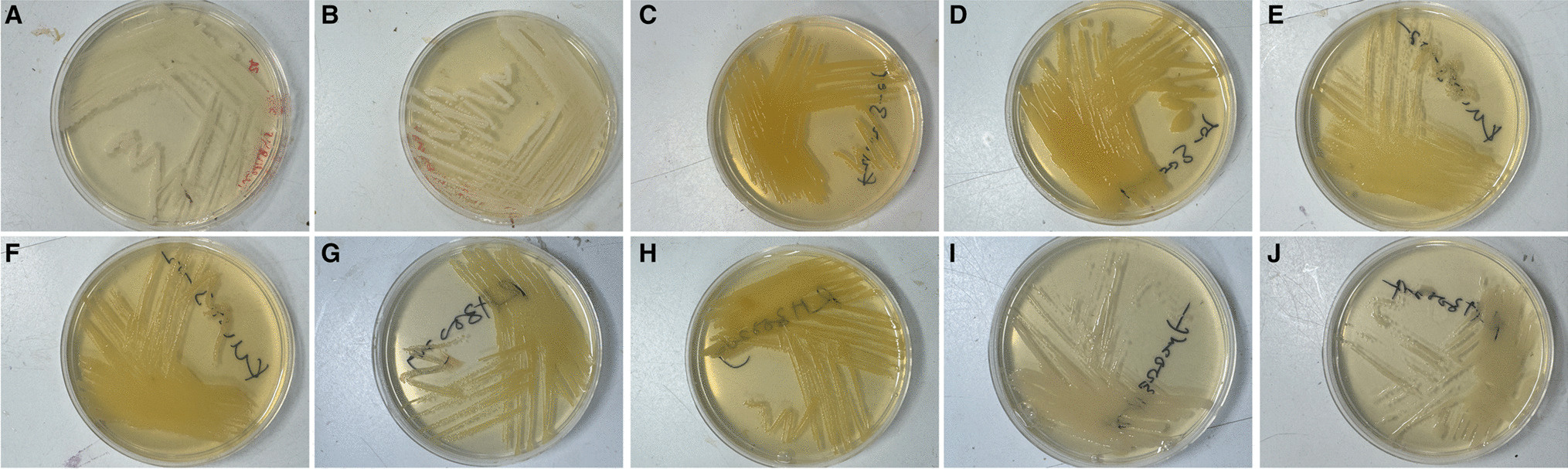
Our modified media solves this problem. By simply adding sodium bicarbonate to the three commonly used growth media for these bacteria, we have demonstrated growth of Neisseria gonorrhoeae in normal, standard atmosphere incubators. The GC agar, Chocolate agar, and Thayer Martin agar growth media with sodium bicarbonate are as effective as growth of the bacteria in a CO2 incubator (shown below, A in CO2, B on our media) . This opens up a world of possibilities about where and when these bacteria can be grown in laboratory conditions.
Follow the Topic
-
Applied Microbiology and Biotechnology
This journal focuses on research regarding prokaryotic or eukaryotic cells, relevant enzymes and proteins, and applied genetics and molecular biotechnology.
Related Collections
With Collections, you can get published faster and increase your visibility.
Cultivation of Microorganisms: New Media and Techniques
Applied Microbiology and Biotechnology invites articles on new approaches to cultivating microorganisms, focusing on novel culture media and techniques that bring previously difficult to culture microbes into the laboratory. Topics of interest include customized genome-informed media, microfluidic and high-throughput isolation systems, co-culture and in situ devices, and advanced bioreactor methods.
Culture media and cultivation methods are key to unlocking microbial diversity for biotechnology. Many microbes remain uncultured, highlighting the need for new approaches. Many organisms cannot grow on standard lab media and instead require tailored formulations mimicking their native habitats. This collection seeks novel methods to culture microorganisms, from optimized growth media to high-throughput platforms, including co-culture and in situ techniques. Emphasis is on expanding culturable diversity and enabling strain development for applied microbiology and industrial biotechnology.
We invite studies on cultivation methods and media that enable growth of previously uncultured or fastidious microbes. Innovative culture media topics include defined media guided by genome data or novel additives to mimic environmental niches. Contributions on advanced platforms – such as microfluidic or droplet-based high-throughput systems, diffusion chambers, and automated culturomics – are welcome. Also relevant are co-culture or continuous culture strategies, anaerobic and extremophile techniques, and improved protocols for culture reproducibility.
Topics of interest include, but are not limited to:
• Genome-informed and environmentally mimetic culture media
• High-throughput cultivation platforms
• Co-culture and community culture strategies to grow interdependent microbes
• In situ cultivation devices and methods
• Continuous and automated culture systems
• Specialized methods for anaerobic, microaerophilic, or extremophilic microorganisms
Publishing Model: Open Access
Deadline: Oct 15, 2026
Emerging Applications of Nanoparticles in Biotechnology and Medicine
Applied Microbiology and Biotechnology invites articles on nanoparticles and nanomaterials in biotechnology and medicine. Topics of interest range from nano-enabled diagnostics and drug delivery to enzymatic catalysts in bioreactors and nanoparticles for environmental and health applications. Medical and health-related articles should specifically address diseases caused by bacteria or viruses, or focus on therapeutics produced through biotechnological or enzymatic processes.
Nanoparticles, with their unique size-dependent properties, are transforming biotechnology and medical research. This collection highlights innovative applications of engineered nanoparticles in diagnostics, targeted drug delivery, biosensing, and catalytic bioprocesses. Contributions may include studies of nanoparticle design and functionalization, nanocarrier systems for drug, gene, or enzyme delivery, and nanoparticles as enzyme-immobilization supports to boost bioprocess efficiency. Emphasis is on how these nanoscale materials can improve processes from bio-production to therapeutics, highlighting the impact of nanobiotechnology on health and industrial biotechnology.
We invite contributions on novel nanoparticles and functionalized materials with biotechnological or medical applications. Potential topics include nanoparticle synthesis and engineering for targeted functions, nano-carriers for drug, gene or enzyme delivery, and nanomaterials for biosensors and diagnostics. Studies of nanoparticle applications in bioprocesses (e.g. catalysis in bioreactors, fermentation enhancement) and environmental biotechnology (e.g. pollutant sensing or degradation) are welcome. Works on antimicrobial or antiviral nanomaterials, nano-vaccines, and the safety/toxicology of nanotechnologies are also encouraged.
Topics of interest include, but are not limited to:
• Nanoparticle design and functionalization
• Nanocarriers for targeted drug, gene, or enzyme delivery
• Nano-enabled diagnostics, imaging, and biosensors for microbial and medical applications
• Nanoparticles in bioprocess engineering
• Antimicrobial, antiviral, and nano-vaccine technologies
• Environmental nanoparticle applications (bioremediation, bioenergy) and nanotoxicology studies
This collection supports and amplifies research related to SDG 3 (Good Health and Well-being).
Publishing Model: Open Access
Deadline: Aug 12, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in