How can a seemingly strict mating signal evolve?
Published in Ecology & Evolution and Microbiology
Sexual communication often depends on precise molecular recognition. One cell or organism releases a signal and another receives it. A successful interaction then determines whether mating occurs. At first glance, such systems appear fragile; if the signal changes, the recognition fails. If recognition fails, reproduction stops. How do these systems evolve?
This question has been central to my work on mating pheromones in fission yeast.
In the fission yeast Schizosaccharomyces pombe, mating depends on two sex pheromones and their receptors. M-type (Minus) cells secrete a short lipid-modified peptide called the M-factor, which is recognized by the receptor Map3 on P-type (Plus) cells. P-type cells secrete P-factor, which is recognized by a different receptor, Mam2. Although this appears to be a symmetric signaling system, our previous work showed that it is not equally constrained in nature. The M-factor/Map3 pair is strikingly conserved, whereas the P-factor/Mam2 pair is more variable [Seike et al., PLoS Biol, 2019]. This asymmetry suggests that one side may act as a stricter gatekeeper for species recognition, while the other allows for more evolutionary flexibility.
This raises a more profound question. If M-factor recognition is so strict, how can it change during evolution without breaking the system?
In our earlier study, we showed that prezygotic isolation can be achieved experimentally by altering the M-factor/Map3 pair [Seike et al., PNAS, 2015]. Importantly, that study altered only the M-type pheromone and its receptor; the P-type pathway was left untouched. This demonstrates that rewiring only one side of the pheromone system is sufficient to create reproductive isolation. However, this also left an important question unanswered: In nature, how can a pheromone change while preserving recognition?
This was the starting point for the present study.
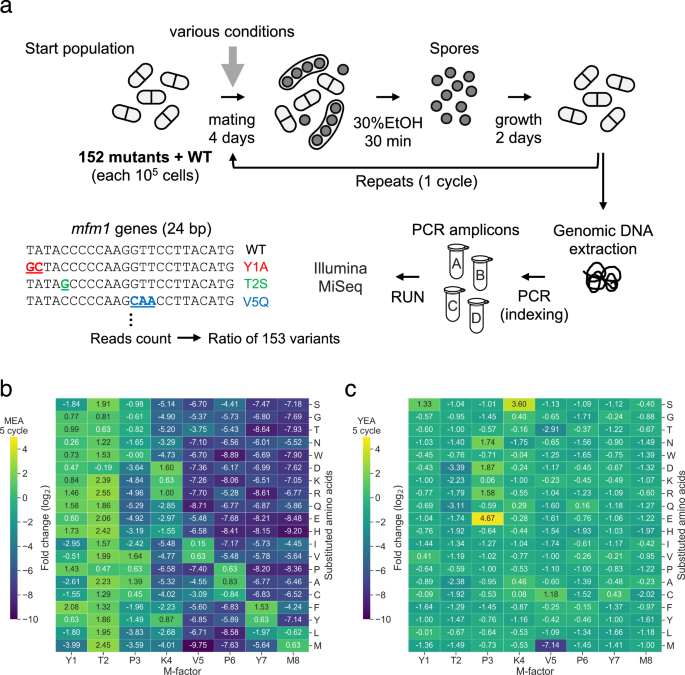
Rather than testing a few mutations individually, we adopted a comprehensive approach. Previously, we constructed a library containing all possible single-amino acid substitutions across non-cysteine residues of the tiny nine-amino acid pheromone M-factor [Seike et al., Genetics, 2012]. This resulted in 152 mutant versions and the wild type. We mixed them together and allowed them to compete through repeated mating cycles. We then determined which variants survived in different environments.
This design was exciting because it transformed the abstract evolutionary question into something visible. Instead of asking only whether a mutation is “functional” or “nonfunctional,” we could ask whether its value changes with context. A variant that appears useless in one condition may become surprisingly successful in another.
This aligned precisely with the observed outcome.
Some variants behaved as expected; changes at certain residues severely damaged mating performance and were quickly lost. However, other results were much more surprising. A subset of variants that appeared sterile or nearly sterile under standard laboratory conditions was much more successful under different pH conditions. In particular, one variant (P6H), in which the sixth residue was changed from proline to histidine, was almost inactive under our usual conditions, but became strongly activated at a higher pH.
This was one of the most exciting moments of the project. A mutation that would normally be dismissed as deleterious turned out not to be simply “bad,” but rather condition dependent. In the right environment, it works very well.
This observation changed my thoughts about pheromone evolution. If the environment can dramatically reshape the activity of a signaling peptide, variants that appear nonfunctional in the laboratory may still matter in nature. A mating system that appears extremely strict under one condition may become more permissive in another. In heterogeneous habitats, this could open temporary windows in which novel variants remain reproductively competent for long enough to persist, spread, or combine with other changes.
The breakthrough came from treating the competition assay not just as a screen for the “best” variants, but as a way to search for hidden conditions under which unusual variants come alive. Once we observed hints that some mutants behaved differently in different media, changing the environmental pH became the next natural step. This decision ultimately allowed us to identify the conditions under which strains that seemed sterile could become fertile.
Another important aspect arises from a different residue in the M-factor. While P6 highlights environmental activation, substitutions at T2 indicate another route of evolutionary change. The T2Q substitution, in which the second residue was changed from threonine to glutamine, was especially interesting because it improved mating performance under some conditions but also imposed a cost during vegetative growth. Even more intriguingly, it could buffer the effects of otherwise deleterious mutations at other positions. In other words, one mutation could make the other mutations more tolerable.
This kind of permissive mutation is a powerful concept in evolutionary biology. It offers a possible route across a rugged fitness landscape, a pathway along which a molecule can change without passing through a completely nonfunctional intermediate. In our case, this also provided clues to a longstanding puzzle. Schizosaccharomyces octosporus, a closely related fission yeast species, carries three differences in its M-factor sequence relative to S. pombe. On their own, some of these changes are harmful in the S. pombe Map3 context. However, if a permissive change, such as T2Q substitution, occurs early, later changes may become evolutionarily accessible. This provides a plausible scenario for how a short peptide pheromone can diverge while maintaining system functionality.
Taken together, this study reveals two complementary ideas. First, environmental changes can uncover latent functions in variants that otherwise appear sterile. Second, permissive mutations can buffer harmful effects and open new evolutionary paths. Both mechanisms help explain how a small, highly constrained peptide can still evolve.
This was the most satisfying aspect of my work. The original question was almost paradoxical: how can a strict mating signal change without losing recognition? Our results suggest that evolution can proceed through context-dependent activation, mutational buffering, or a combination of the two.
Much remains unknown, including the molecular basis of the effect of pH and how often such environmentally responsive variants occur in natural populations. Natural isolates of S. pombe studied so far—approximately 150 strains—show complete conservation of the M-factor. This suggests that relevant ecological niches are still undersampled. Insects such as Drosophila spp. are especially intriguing candidates because they disperse yeasts and may expose them to chemically distinct environments.
More broadly, this work is a reminder that even a nine-amino acid peptide can encode surprisingly rich evolutionary logic. What appears to be a broken key may simply be waiting for the correct working conditions to work.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Related Collections
With Collections, you can get published faster and increase your visibility.
From RNA Detection to Molecular Mechanisms
Publishing Model: Open Access
Deadline: May 05, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in