Old Molecules, New Rules: Why Science Misses the "Forest" for the Proteins
Published in Physics, Cell & Molecular Biology, and Biomedical Research
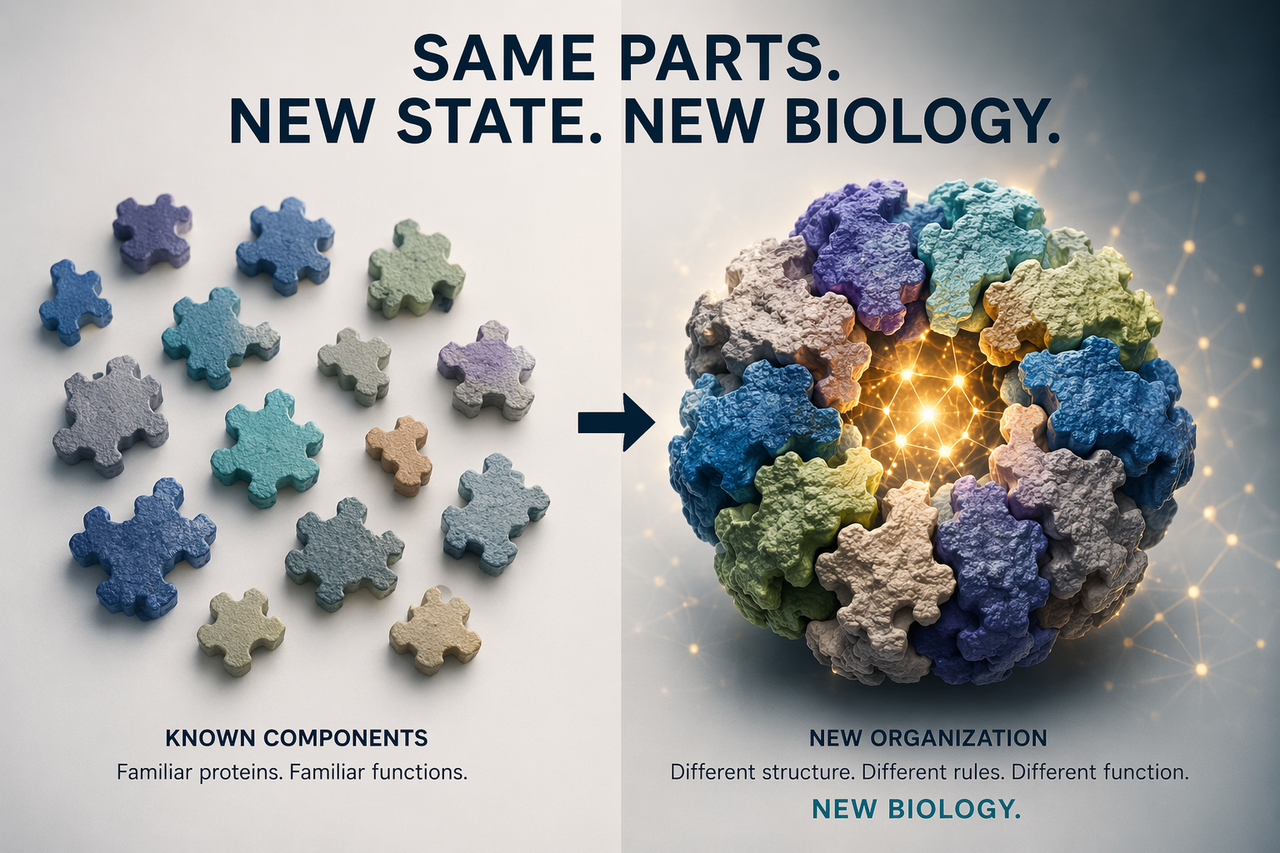

Science is good at recognizing new molecules. It is much worse at recognizing new biological states built from familiar ones.
When the components are already known, the instinct is to assume the biology is already known too. A new assembly is reduced to a variant of an old one. A distinct state is folded back into the dominant framework. What should be examined as new is explained away as familiar.
That reflex is one of the quiet dogmas of modern biology.
Chaperones illustrate the problem well. Because they have long been defined through their canonical roles, there is resistance to seeing that the same molecules might also participate in fundamentally different, regulated assembly states with different functions. Once a field becomes attached to a canonical view, it begins to mistake familiarity of parts for familiarity of biology.
But cells work through organization, not just inventory. The same molecules can be deployed in different states, under different rules, with different consequences. If we do not learn to see that, we will keep missing important biology hiding in plain sight.
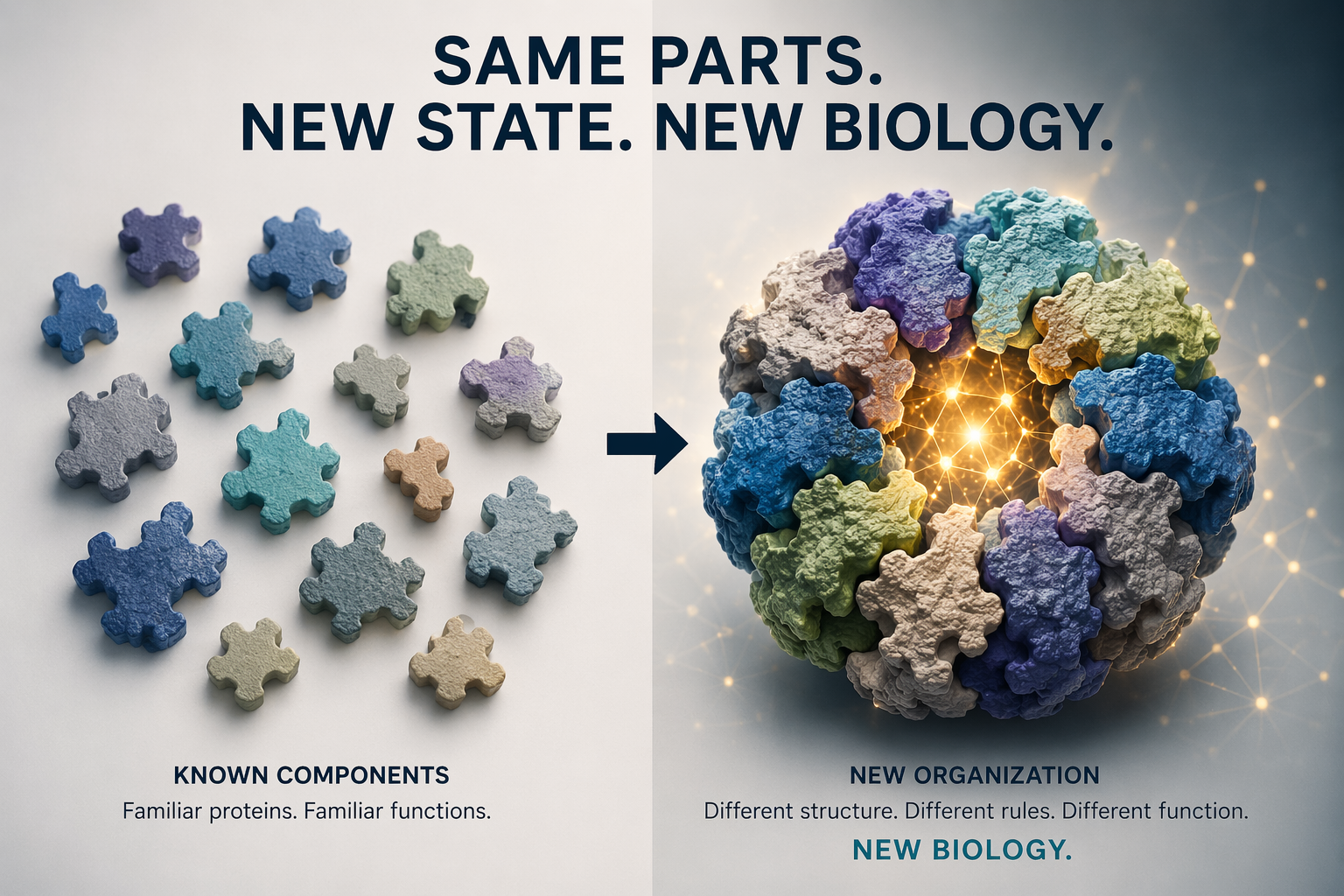

The right question is whether the assembly state is new.
Novelty in biology does not always arrive as a new molecule. Sometimes it arrives when old molecules are assembled into a new state. And those may be the discoveries that are hardest for us to see.
Further reading
Roychowdhury T, McNutt SW, Pasala C, Nguyen HT, Thornton DT, Sharma S, Botticelli L, Digwal CS, Joshi S, Yang N, Panchal P, Chakrabarty S, Bay S, Markov V, Kwong C, Lisanti J, Chung SY, Ginsberg SD, Yan P, De Stanchina E, Corben A, Modi S, Alpaugh ML, Colombo G, Erdjument-Bromage H, Neubert TA, Chalkley RJ, Baker PR, Burlingame AL, Rodina A, Chiosis G, Chu F. Phosphorylation-driven epichaperome assembly is a regulator of cellular adaptability and proliferation. Nat Commun. 2024 Oct 16;15(1):8912. doi: 10.1038/s41467-024-53178-5.
Chiosis G, Digwal CS, Trepel JB, Neckers L. Structural and functional complexity of HSP90 in cellular homeostasis and disease. Nat Rev Mol Cell Biol. 2023 Nov;24(11):797-815. doi: 10.1038/s41580-023-00640-9.
Digwal CS, Wang S, Chiosis G. Embracing diversity: Post-translational modifications, the chaperone code, and the emergence of new chaperone entities. Cell Stress Chaperones. 2026 Mar;31(2):100148. doi: 10.1016/j.cstres.2026.100148.
Chu F, Sharma S, Ginsberg SD, Chiosis G. PTMs as molecular encoders: reprogramming chaperones into epichaperomes for network control in disease. Trends Biochem Sci. 2025 Oct;50(10):892-905. doi: 10.1016/j.tibs.2025.07.006.
Yan P, Patel HJ, Sharma S, Corben A, Wang T, Panchal P, Yang C, Sun W, Araujo TL, Rodina A, Joshi S, Robzyk K, Gandu S, White JR, de Stanchina E, Modi S, Janjigian YY, Hill EG, Liu B, Erdjument-Bromage H, Neubert TA, Que NLS, Li Z, Gewirth DT, Taldone T, Chiosis G. Molecular Stressors Engender Protein Connectivity Dysfunction through Aberrant N-Glycosylation of a Chaperone. Cell Rep. 2020 Jun 30;31(13):107840. doi: 10.1016/j.celrep.2020.107840.
Pasala C, Sharma S, Roychowdhury T, Moroni E, Colombo G, Chiosis G. N-Glycosylation as a Modulator of Protein Conformation and Assembly in Disease. Biomolecules. 2024 Feb 27;14(3):282. doi: 10.3390/biom14030282.
Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in