Written by Eva Zanditenas, edited by Dr. Ilana Kolodkin-Gal and Prof. Serge Ankri
Entamoeba histolytica is a human parasite responsible for the disease amoebiasis. The parasite enters the human body through contaminated food or water, and, when in the colon, feeds on gut microbiota bacteria, intestinal mucus, and finally epithelial cells. After degrading the epithelial cell, it reach the blood stream and the parasite can cause liver or lung abscesses and even death (100,000 death every year worldwide, [1]).
The feeding of the parasite on gut bacteria is crucial for its virulence. [2]. In addition, the parasite’s virulence varies according to the species on which it feeds. To study relationship between E. histolytica and bacteria is therefore crucial to understand the behavior and dynamics of this parasite in the human colon.
However, all previously reported studies of this interaction use the planktonic form of bacteria, meaning free-living and independent bacteria. In the colon, bacteria assemble to create aggregates protected by a self-generated matrix, forming a biofilm [3].
Amoeba have been shown to interact with biofilm, mostly in aquatic environments. However, our article presents the first study on the interaction between a human amoebic parasite and a bacterial biofilm. The parasite’s transcriptomic response to the biofilm differs completely compare to its response to planktonic bacteria. This difference demonstrates a specific mechanism developed by the parasite to feed on biofilms that may occurs in the infected human colon. Another key difference between planktonic form and biofilm form is the attachment mechanism of the parasite on its target. To attach to planktonic bacteria, the parasite uses the Gal/GalNac lectin receptor to bind to galactose and N–acetyl-galactosamine residues on the bacterial surface [4]. At least for the bacteria species study in this work, this receptor is not paramount for the attachment of the parasite on biofilms.
When E. histolytica first makes contact with the biofilm, it attaches to the biofilm and begins degrading the extracellular matrix produced by the bacteria. The parasite is able to degrade the biofilm of several bacterial species, including Gram-positive species like B. subtilis or gram-negative species like E. coli. The class of enzyme responsible for the attachment and matrix degradation are the cysteine proteinase, a well-known class of enzymes that play an important role in the virulence of E. histolytica because they are responsible for mucus degradation and epithelial cell death [4]. The cysteine proteinase genes are also upregulated in E. histolytica incubated with B. subtilis biofilm compared to their planktonic counterpart. In this study, one particular cysteine proteinase located on the surface of the parasite, called EhCP5, was genetically downregulated. Parasites with downregulated EhCP5 were significantly less able to degrade B. subtilis biofilm, providing evidence for the role of this specific cysteine proteinase in the biofilm degradation.
We studied more specifically B. subtilis biofilm interaction with E. histolytica, and we discovered that TasA, a major B. subtilis matrix protein and an efficient adhesin [5], is the target of the cysteine proteinase. For E. coli, we were not able to discover the target of the parasite but we know that it is not curli, a major E. coli matrix component. Interestingly, TasA serves in a dual role during parasite-biofilm interactions, as it also facilitates the adhesion of the parasite to the biofilm. As a result of the biofilm matrix degradation, bacteria in the biofilm are more exposed and more sensitive to antibiotics like sodium hypochlorite or β-lactam antibiotic ampicillin.
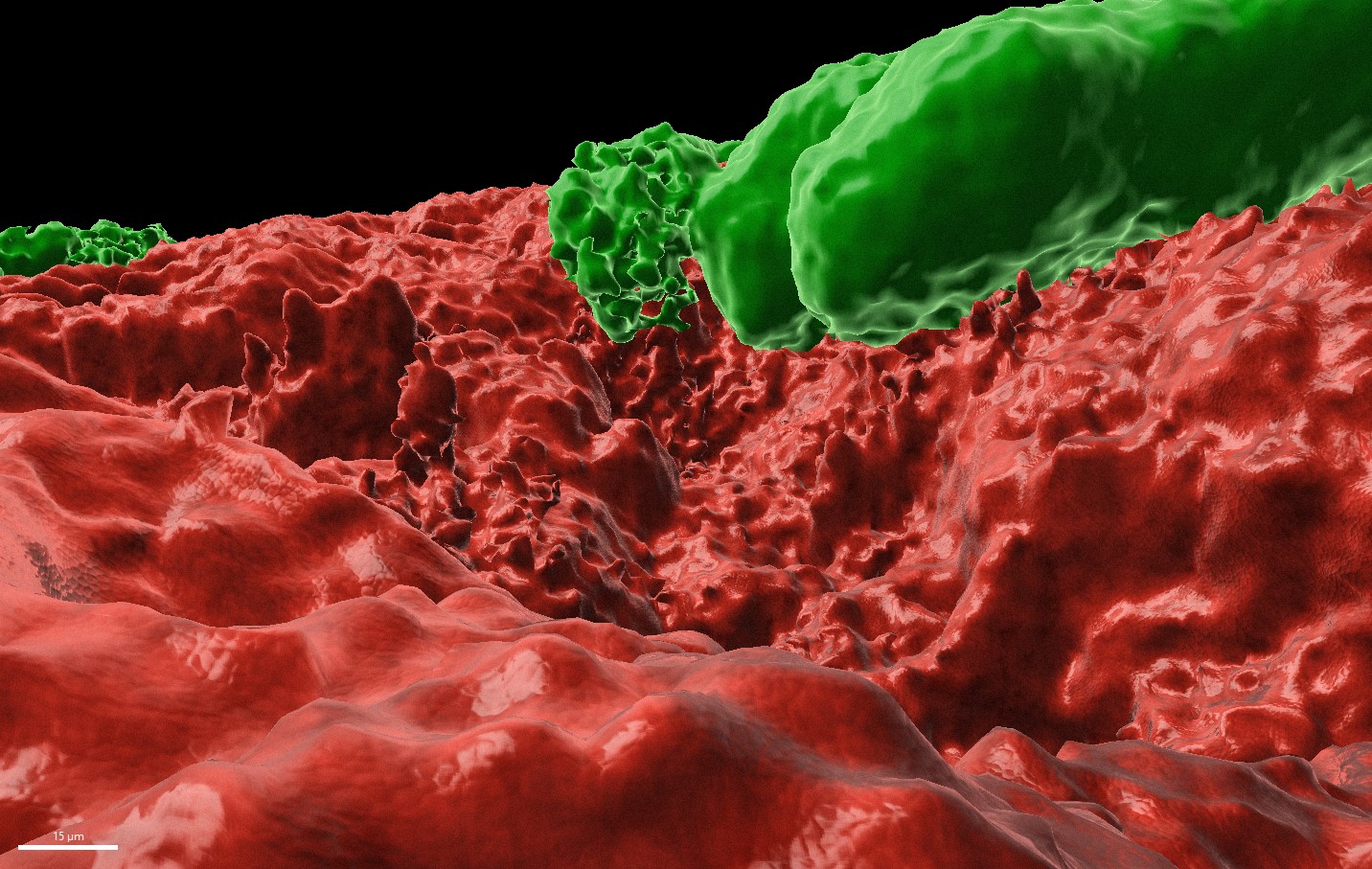
B. subtilis biofilm is a robust and thick biofilm with an impressive 3D structure containing an oxygen gradient. The more deep in the biofilm the less oxygen is present. Microscopy images clearly indicate that E. histolytica penetrates into the biofilm. When the parasite is in the biofilm, it is protected from the environmental oxidative stress compared to when the parasite is incubated with the planktonic form of the same bacteria, as indicated by downregulation of genes related to the parasite response to oxidative stress
To conclude, the parasite demonstrate a specific response in the presence of a biofilm compared to planktonic bacteria. These results are imperative to unravel the ecology of the mammalian microbiome, as well as many additional habitats, where protist predators interact with structured bacterial communities. A specific mechanism was identified that involves cysteine proteinase, which degrade the biofilm matrix protein TasA present in B. subtilis biofilm. When E. histolytica penetrates the biofilm, it is protected against oxidative stress. Indeed, the biofilm being degraded is more sensitive to antibiotic treatment. This research demonstrates that E. histolytica has adapted to feed on biofilms. Indeed, E. histolytica represent a gold mine of anti-biofilm molecules.
For more details on this work, visit our recent publication in npj Biofilms and Microbiomes: https://www.nature.com/articles/s41522-023-00444-x
Legend for the poster image: The image was acquired using the inverted fluorescent confocal microscopy. The 3D model (built with the imaris software) shows B.subtilis biofilm (in red) and some E. histolytica trophozoites (in green) grazing on biofilm. This picture is unique as it shows the parasite in action of breaking the biofilm and to capture bacteria as its prey.
References:
[1] Stauffer W, Ravdin JI. Entamoeba histolytica: an update. Curr Opin Infect Dis. 2003 Oct;16(5):479-85. DOI: 10.1097/00001432-200310000-00016
[2] Phillips B, Wolfe PA, Rees CW, Gordon HA, Wright WH, Reyniers JA. Comparative results of the intracecal inoculation of germfree, monocontaminated, and conventional guinea pigs with Entamoeba histolytica. Am J Trop Med Hyg. 1955;4(4):675–692. doi:10.4269/ajtmh.1955.4.675.
[3] Sauer K, Stoodley P, Goeres DM, Hall-Stoodley L, Burmølle M, Stewart PS, Bjarnsholt T. The biofilm life cycle: expanding the conceptual model of biofilm formation. Nat Rev Microbiol. 2022 Oct;20(10):608-620. doi: 10.1038/s41579-022-00767-0. Epub 2022 Aug 3. PMID: 35922483; PMCID: PMC9841534.
[4] Gilchrist, Carol A, and William A Petri. “Virulence Factors of Entamoeba Histolytica.” Current Opinion in Microbiology 2, no. 4 (August 1, 1999): 433–37. https://doi.org/10.1016/S1369-5274(99)80076-9.
[5] Steinberg, N. et al. ,The extracellular matrix protein TasA is a developmental cue that maintains a motile subpopulation within Bacillus subtilis biofilms. Sci. Signal.13,eaaw8905(2020).DOI:10.1126/scisignal.aaw8905
Follow the Topic
-
npj Biofilms and Microbiomes
The aim of this journal is to serve as a comprehensive platform to promote biofilms and microbiomes research across a wide spectrum of scientific disciplines.
Related Collections
With Collections, you can get published faster and increase your visibility.
Microbial endocrinology
Publishing Model: Open Access
Deadline: Oct 21, 2026
Microbiome and energy metabolism
Publishing Model: Hybrid
Deadline: Dec 06, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
Great piece of work. Thanks for sharing.
Please find my work here on biofilms:
https://go.nature.com/3RduzBM