Surveying the chemistry of pest-controlling fungi.
Published in Chemistry, Microbiology, and Plant Science
Why did we choose Hypocreales?
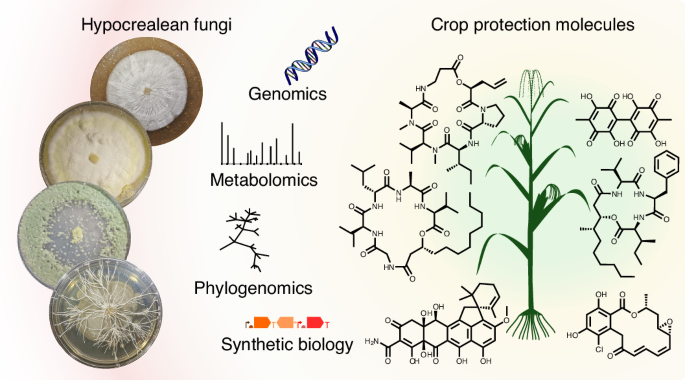
Our group is fascinated by fungi and the wide variety of molecules they produce. These tiny chemists (Figure 1) are able to produce molecules with applications in medicine, agriculture, and materials science. We aimed to explore the chemical potential of fungi to produce molecules that could be used in sustainable solutions and novel materials. Inspired by this diversity, we sought to harness fungi’s biosynthetic machinery through heterologous expression, turning their natural chemistry into accessible, scalable compounds for research and application.
So then, where did we decide to start this journey from? Early on, we became particularly intrigued by fungi in the order Hypocreales, a diverse group that includes insect pathogens, plant-associated endophytes, and other ecological specialists. Many species interact with insects, sometimes killing them or manipulating their behavior, as in the well-known “zombie ant” phenomenon. These interactions raised a compelling question: what molecules underlie such biological effects, and how are they encoded in the genome? There was not much in the literature about the chemistry and genomes of these species, leaving room for us to investigate them. This curiosity was the starting point for a large-scale exploration of Hypocreales chemistry and genetics. Our lab is now focus on using the biosynthetic catalog resulting from the project to improve pest-controlling microbes.
From a single fungus to 82 genomes: scaling up our search for molecules
Our goal was simple in principle but ambitious in practice: to understand what these fungi are capable of producing. We started with a single species, Tolypocladium inflatum, cultivating it under many different conditions and analyzing its metabolome. Even at this early stage, we detected a surprising range of compounds, including molecules not previously described in Hypocreales, a clear sign that these fungi had far more chemistry to reveal.
Encouraged by these results, we expanded the project to a collection of 82 Hypocrealean strains spanning a broad taxonomic range, combining publicly available strains with our own isolates. To keep the study systematic and manageable, we decided to cultivate the entire collection under five different media conditions.
Beyond the chemistry, we also enjoyed simply observing the fungi themselves. Across strains and conditions, they grew in remarkably different shapes and forms, offering small pleasures along the way.

Figure 1: Trichoderma atroviride growing on Potato Dextrose Agar
We sequenced genomes, profiled metabolomes, and applied metabologenomics approaches to link biosynthetic gene clusters to detected molecules. By integrating phylogenomics, we grouped these pathways into gene cluster families and studied their distribution across species.
One particularly interesting observation from the genomic data was that a large fraction of non-ribosomal peptide synthetases (NRPSs) in these fungi are small, often mono- or bimodular enzymes. This was exciting because our lab has expertise in expressing such enzymes in yeast. We selected several uncharacterized NRPS-like genes and expressed them heterologously, allowing us to access their products directly, including compounds that may not have been previously described in Hypocreales.
From raw data to discovery: making sense of thousands of genes and molecules
A project of this scale required a truly collective effort. As a lab, we curated more than 5,000 biosynthetic gene clusters, most of which are associated with unknown products. Annotating these clusters and matching them to metabolites was both the central challenge and one of the most rewarding aspects of the work.
On the metabolomics side, we relied on public natural products databases for spectral matching, carefully evaluating each annotation to ensure confidence in compound identification. Before that, we had to preprocess large volumes of mass spectrometry data, making sure that consistent parameters were applied across all samples.
When we began organizing the data, even simple tasks felt daunting. Starting the spreadsheet to match compounds to biosynthetic gene clusters (Figure 2), it was easy to feel that it might never end. The dataset was so rich that we could have kept exploring indefinitely, each pathway leading to another interesting question, each molecule opening a new direction.
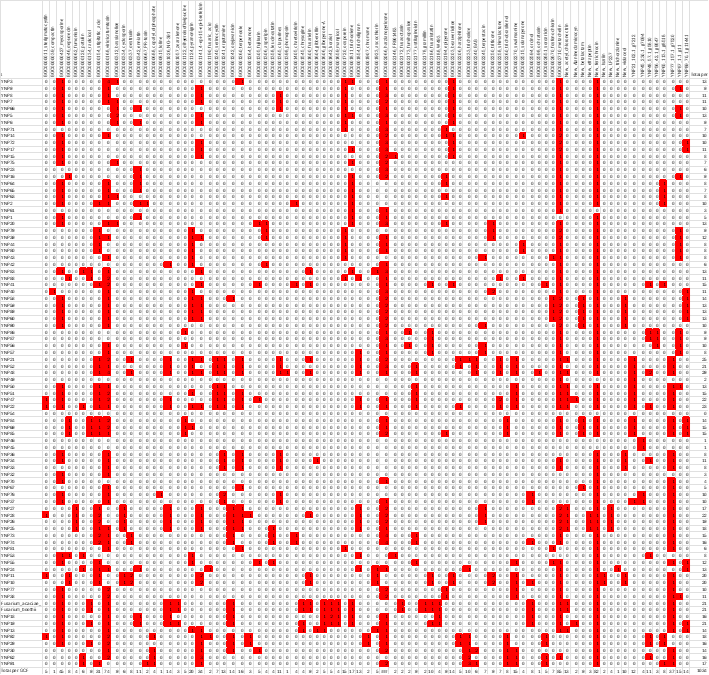
Figure 2: Draft version of a presence–absence matrix showing gene cluster families across Hypocreales strains. Rows represent strains and columns represent Gene Cluster Families (GCF); The numbers in the table indicate the number of compounds putatively associated with that GCF.
At one point, we had to make a conscious decision as a team: to stop analyzing and start writing. It was not easy. The temptation to keep digging into new biosynthetic pathways was strong, and we genuinely worried that we might never “close” the project because there was always something new and exciting to explore.
Linking metabolites to gene clusters required us to dive deeply into biosynthetic logic, comparing gene architectures and building hypotheses for how specific molecules could be assembled. At times, the scale of the data felt overwhelming. But each successful match, each moment where a molecule could be connected to a gene cluster was genuinely exciting and kept us moving forward.
More unknown than known: what surprised us most
Although we are still far from fully characterizing the chemical and genetic landscape of Hypocreales, this work represents an important step. Brick by brick, we are building a more complete understanding of these microorganisms and their capability.
Our results highlight just how much remains to be discovered. We found that around 80% of biosynthetic loci could not be assigned to known gene cluster families, and only a small fraction could be linked to known molecules detected under our experimental conditions. This suggests that Hypocreales harbor a vast, untapped reservoir of chemistry.
At the same time, even linking known compounds to their corresponding gene clusters has significant value. It opens the door to producing these molecules through heterologous expression, which we have already demonstrated in our lab. This is a step toward developing scalable, sustainable production methods for biocontrol agent.
More broadly, this study reinforces the idea that fungal biocontrol has enormous potential for sustainable agriculture. Crop losses due to fungal pathogens and insect pests remain substantial worldwide, despite heavy reliance on chemical pesticides. By uncovering the molecular basis of fungal biocontrol, we hope to contribute to the development of more environmentally adequate alternatives.
From discovery to what comes next: building a roadmap for fungal biocontrol
Looking back, this project was as much about building a resource as it was about answering a specific question. We set out to explore the chemistry of Hypocrealean fungi, but along the way, we created a framework for connecting genes, molecules, and evolution in a systematic way. By mapping the biosynthetic potential of these fungi and understanding how it is distributed across species, we can begin to guide the chemical characterization of this agriculturally important group and identify promising biocontrol strains and compounds for safer, more efficient, and more sustainable pest control.
At the same time, this work highlights how much remains to be done. Cultivation-based methods often yield very small amounts of product, making it difficult to fully characterize many of the molecules we identified. Developing high-throughput heterologous expression systems will be essential to access and study this hidden chemical diversity at scale. In our group, we have already taken steps in this direction by expressing uncharacterized biosynthetic gene clusters, including those encoding pyridones, one of the most abundant classes across our strains.
We also hope that by making our genomic and metabolomic datasets publicly available, other researchers will build upon this work, matching orphan gene clusters to newly discovered molecules and expanding our collective understanding of fungal chemistry. Along the way, we came to appreciate how important it is to share data: advancing knowledge in this field is truly a community effort, where each contribution builds on the last. By continuing to bridge genes and molecules, we move closer to unlocking the full potential of fungal chemistry for sustainable biocontrol.
If there is one takeaway, it is that we are only beginning to understand the chemical capabilities of fungi. The tools are now in place, the datasets are growing, and the possibilities are vast.
And for us, this is just the beginning.
This post was Written by Ana Calheiros de Carvalho with Minimal edition of PCM (figure 3)

Figure 3. The authors of the post during the 2023 Christmas analytical chemistry workshop.
Follow the Topic
-
Nature Chemical Biology
An international monthly journal that provides a high-visibility forum for the chemical biology community, combining the scientific ideas and approaches of chemistry, biology and allied disciplines to understand and manipulate biological systems with molecular precision.




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in