Systematically designing combination therapies by targeting coexisting tumor cell states in diffuse midline glioma
Published in Cancer, Protocols & Methods, and Genetics & Genomics
Hidden diversity inside tumors
Intratumor heterogeneity represents, perhaps, the greatest challenge to successful cancer therapy. Growing evidence shows that virtually all tumors comprise multiple cell populations with distinct transcriptional states and equally distinct drug sensitivities. As a result, monotherapies will almost invariably select for resistant states, which—through adaptation and plasticity—will eventually reestablish tumor heterogeneity, leading to relapse.
This challenge is particularly evident in diffuse midline glioma (DMG), one of the most devastating pediatric brain tumors. DMG is genetically simple, but its hallmark mutations in histone H3 at lysine 27 (H3K27) disrupt chromatin regulation, leading to large-scale remodeling of the tumor regulatory landscape and giving rise to multiple transcriptionally distinct, coexisting cell states. These alterations are not readily druggable and, together with the resulting heterogeneity, contribute to the repeated failure of systemic therapies to produce durable responses.
We set out to address this challenge by elucidating the individual cell states that comprise these tumors and the Master Regulator (MR) proteins that implement and homeostatically maintain them, enabling the rational design of combinations of state-specific drugs targeting these dependencies.
Defining cell states through their regulatory programs
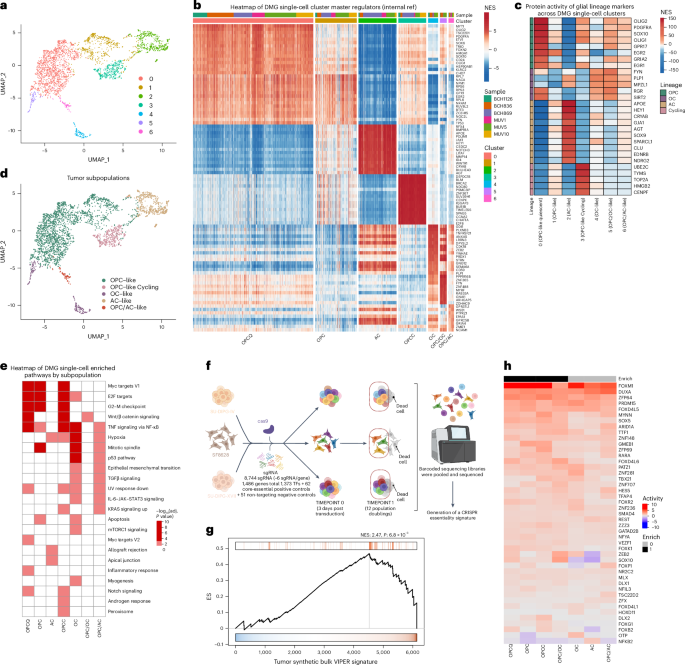
Previous studies had shown that DMG tumors contain diverse transcriptional cell states arising from glial progenitors. Building on these observations, we asked whether single-cell network-based analyses leveraging the ARACNe and VIPER algorithms could further resolve these states and identify the MR proteins that represent their mechanistic determinants and, therefore, their potential therapeutic vulnerabilities. Briefly, VIPER infers protein activity from gene expression data by identifying proteins whose transcriptional targets—as inferred by ARACNe—best recapitulate a tumor cell state gene expression signature.
Single-cell analysis of 14 primary DMG tumors identified seven molecularly distinct, DMG-specific cell states that coexisted in virtually every sample and were reproducible across independent datasets. These included three oligodendrocyte precursor (OPC)-like states, including a highly proliferative population, oligodendrocyte (OC)-like and astrocyte (AC)-like states, as well as transitional states (OPC/OC- and OPC/AC-like), providing a higher-resolution regulatory view of DMG heterogeneity.
This suggests that regulatory canalization constrains tumors to a finite number of states, each controlled by a highly conserved module of MR proteins. This “quantized” view of cancer, in turn, implies that universal, rather than overly personalized, therapies may be achievable by co-targeting the MR proteins of distinct cell states. Pooled CRISPR knockout screens—in three genetically distinct DMG models—confirmed that VIPER-inferred MRs were significantly enriched in tumor-essential genes, thus representing bona fide tumor dependencies.
Prioritizing drugs targeting complementary tumor cell states
We next asked how to effectively target the MR protein dependencies of each individual cell state. We leveraged a mechanism-based approach that prioritizes drugs based on their ability to invert the activity of either individual essential MRs or the entire MR module governing each state, using the OncoTarget and OncoTreat CLIA-compliant algorithms, respectively. For this purpose, we generated transcriptional response profiles for hundreds of clinically relevant oncology drugs in two DMG cell lines selected to optimally recapitulate patient-derived aberrant MR protein activity. Using the VIPER algorithm, we assessed whether each drug could invert the activity of state-specific MR proteins, by comparing drug- versus vehicle-treated samples. The analysis revealed drugs predicted to selectively target the MRs of each tumor cell state, including OPC-like states (avapritinib, dinaciclib, trametinib, mocetinostat, and etoposide), as well as AC-like states (ruxolitnib, venetoclax, and larotrectinib), together accounting for the majority of tumor cells.
In vivo treatment confirms state-specific drug sensitivity
To test these predictions in DMG cells grown in vivo, we used flank xenograft models, which strikingly recapitulated the full repertoire of states observed in patient tumors. This provided an optimal pharmacodynamic system to assess whether selected drugs could effectively and specifically deplete their predicted target states. Single-cell analysis of treated tumors revealed that eight of the nine predicted drugs significantly depleted their corresponding cell states—an almost 90% validation rate for cell state-specific drug sensitivity in vivo. Intriguingly, one drug (larotrectinib) also emerged as a potent inhibitor of the OC-like state in vivo. Taken together, these results provide a mechanism-based rationale for combinatorially targeting all DMG cell states. These experiments also revealed something that is often missed in conventional drug studies: therapies did not simply shrink tumors but reshaped their internal composition. Depleting one cell state enriched others, revealing how selective pressure on distinct cell states may drive drug adaptation and resistance.
Combination therapies targeting complementary tumor cell states
The ultimate test was whether targeting complementary cell states could improve outcomes in tumors growing in their native brainstem environment. Consistent with OPC-like states representing the dominant cell population in DMG tumors, treatment of an orthotopic model with OPC-targeting monotherapy significantly reduced tumor growth and extended survival. However, the most compelling results arose from combining drugs shown to target complementary cell states. Pairing OPC-targeting and AC-targeting drugs resulted in multiple combinations that significantly outperformed their associated monotherapies in vivo. Among the most striking, avapritinib and ruxolitinib extended median survival to 83 days compared to 53.5 days for avapritinib alone and 25 days for vehicle-treated animals. Avapritinib combined with larotrectinib—also found to deplete OC-like cells—was similarly effective. Together with the improved outcomes observed for combination therapies, this suggests that a three-drug combination targeting OPC-, AC-, and OC-like states may further improve outcomes.
Critically, several drugs showing in vivo synergy, such as ruxolitinib, had little or no effect as single agents, because they targeted minority cell states. This suggests that traditional drug screens may miss critical components of potentially effective combination therapies. Finally, BLISS analyses performed in homogeneous cell cultures comprising almost pure individual cell states showed no synergistic effects, confirming that the synergy observed in vivo was not due to targeting synthetic lethal targets in a single cell state.
Looking forward
This work highlights a new concept of quantized cancer biology, in which tumors are comprised of distinct cell states and associated MR proteins that are remarkably conserved across an entire cohort, thus providing a path toward more universal, rather than overly personalized, therapy approaches. These results show that designing mechanism-based drug combinations—to inactivate the MR proteins controlling complementary tumor cell states at the single cell level, as determined by the differential expression of their MR-targets—represents a more systematic, rational, and effective strategy than empirical drug combination approaches.
Using DMG as a proof of concept, this strategy yielded combinations that almost universally outperformed their associated monotherapies and identified actionable therapies for clinical translation. More broadly, this work establishes a tumor- and mutation-agnostic framework for rational combination therapy design in heterogeneous cancers.
Follow the Topic
-
Nature Genetics
This journal publishes the very highest quality research in genetics, encompassing genetic and functional genomic studies on human and plant traits and on other model organisms.





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in