Targeting DCPS in myeloid malignancies: The path from biomarker discovery to clinical utility
Published in Cancer, Biomedical Research, and Pharmacy & Pharmacology

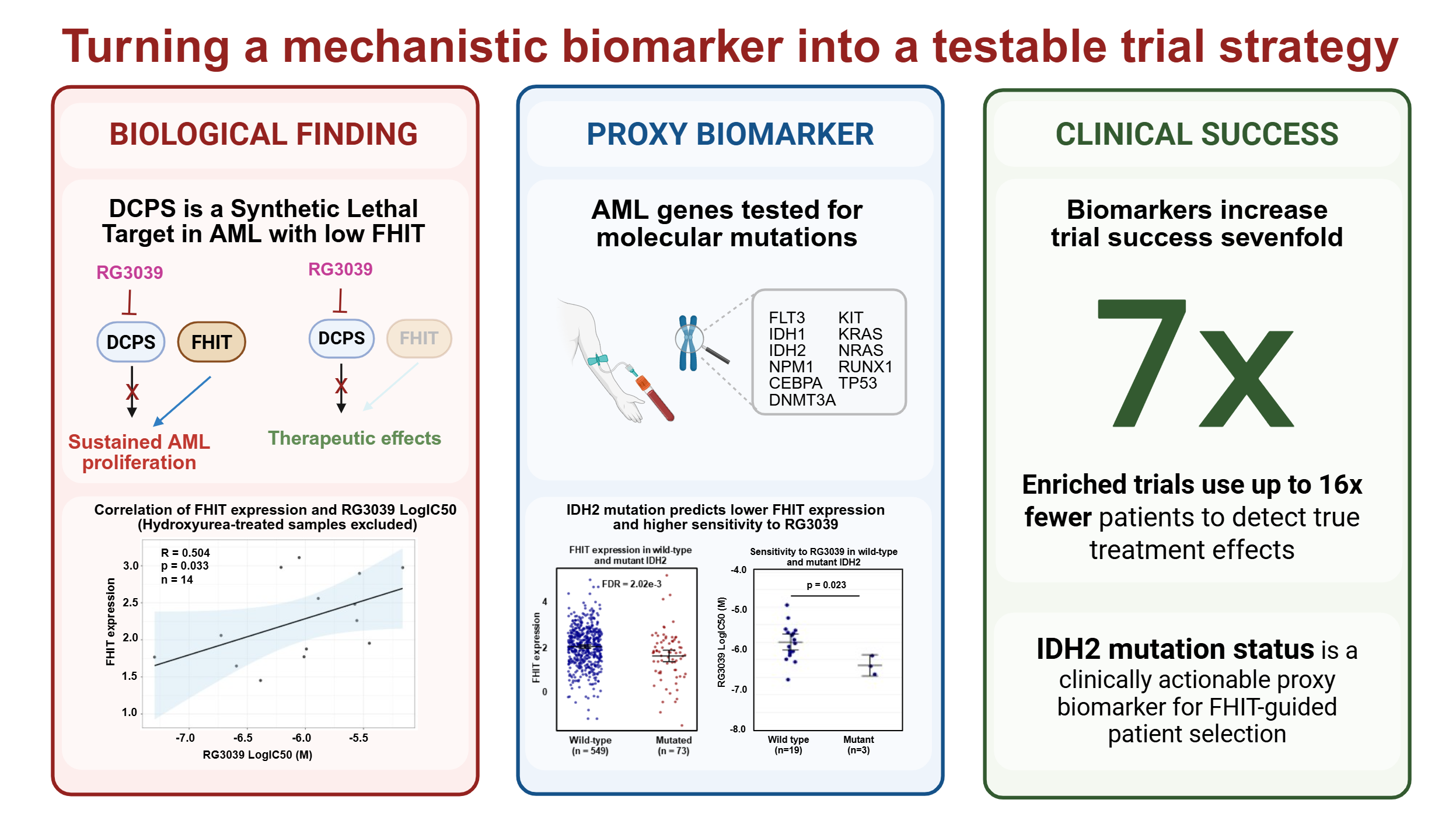
We previously published a link between Acute Myeloid Leukemia cells expressing low levels of the tumor suppressor FHIT and sensitivity to loss of the decapping scavenger enzyme DCPS [1]. In the current study we asked whether that link holds in primary AML samples, how common FHIT-low disease may be across AML cohorts, whether myelodysplastic syndrome (MDS) also fits within this biomarker framework, and whether a more clinically accessible proxy biomarker could be found. The core finding was both encouraging and cautionary: low FHIT expression did predict sensitivity to the DCPS inhibitor RG3039, but hydroxyurea pretreatment confounded that signal in primary AML samples by reducing their sensitivity to treatment. At the same time, the broader cohort analyses showed that FHIT-low biology is relevant in a meaningful subset of disease, including 5–24% of AML, with the highest prevalence reported in pediatric AML (24.4%). FHIT promoter methylation was present in 35.8% of MDS and, notably, remained stable during azacitidine treatment, suggesting that FHIT-low predictive value may be feasible beyond AML. Most importantly, IDH2 mutation status emerged as a clinically accessible proxy biomarker for FHIT-linked DCPS sensitivity in AML [2].
That shift, from an interesting biomarker to a usable one, is really the story behind the paper. Biomarkers matter in oncology because they can reduce biological heterogeneity that can hide a real treatment effect in a clinical study. The numbers back this up: one large analysis of oncology drug-development programs found that biomarker-selected trials succeeded at roughly seven times the rate of unselected ones (10.7% versus 1.6%) [3]. A separate large-scale analysis across several major cancer types reported a nearly fivefold higher likelihood of approval when biomarkers were used [4]. These studies are retrospective rather than predictive for any single program, but they make a clear case for why biomarker-guided drug development has become so central in oncology.
However, a biomarker is only helpful if it can be used prospectively. That is where many promising ideas stall. A novel marker may be biologically compelling but practically difficult because it requires a specialized assay, an impractical measurement, or a workflow that most diagnostic labs simply do not run. The FDA’s enrichment guidance puts it plainly: to use a predictive biomarker to select patients who are more likely to respond to treatment, that marker has to work reliably enough to tell responders from nonresponders in a clinical setting [5].
This is where proxy biomarkers become valuable. They do not replace the underlying biology; they provide a more practical way in. FHIT remains the informative signal, but IDH2 mutation offers a cleaner clinical handle because it is already part of routine diagnostic testing. That means easier screening, tighter eligibility, and robust patient selection across multiple sites. In other words, the scientific question stays the same, but the path to testing it becomes less complicated.
This logic also affects trial design and cost. The FDA frames predictive enrichment as a way to improve efficiency by focusing on patients most likely to respond, which means a smaller study can still detect a real treatment effect. The math is striking: if only 25% of patients carry the relevant marker and the rest are unlikely to benefit, an unenriched trial may need up to 16 times more patients than an enriched one [5]. In practice, that translates to fewer patients to screen, fewer sites, faster recruitment, and much cheaper path to a proof-of-concept outcome.
For us, this is what made this paper satisfying. It started with a mechanistic biomarker hypothesis around FHIT and became a much more practical question: how do you carry elegant biology into a real clinical trial? Not by stepping away from the science, but by building a bridge between biology and deployment. IDH2 is that bridge: not a replacement for FHIT, but a clinically actionable way to reach it. In oncology drug development, this is the kind of bridge that often separates an interesting finding from a strategy that can be actually tested.
References
- Grassi F, Singh M, Moussaud S, Vazquez Rodruguez G, Ali Z, et al. DCPS is a synthetic lethal therapeutic target in acute myeloid leukemia expressing low levels of FHIT. Leukemia39, 2021–2025 (2025). DOI: 10.1038/s41375-025-02661-z
- Grassi F, Bast L, Singh M, Tobiasson M, Walfridsson J, de Milito A, et al. Integrated FHIT and IDH2 biomarker profiling predicts lethal sensitivity to DCPS inhibition in Acute Myeloid Leukemia and Myelodysplastic syndrome. Discover Oncology. 2026. DOI: 10.1007/s12672-026-04880-x
- Wong CH, Siah KW, Lo AW. Estimation of clinical trial success rates and related parameters. Biostatistics. 2019;20(2):273–286. DOI: 10.1093/biostatistics/kxx069
- Parker JL, et al. Does biomarker use in oncology improve clinical trial failure risk? A large-scale analysis. Cancer Medicine. 2021;10(6):1955–1963. DOI: 10.1002/cam4.3732
- U.S. Food and Drug Administration. Enrichment Strategies for Clinical Trials to Support Determination of Effectiveness of Human Drugs and Biological Products. Guidance for Industry. March 2019. Official FDA guidance (PDF)
Follow the Topic
-
Discover Oncology
This is a fully open access general oncology journal that aims to provide a unified forum for researchers and clinicians. The journal spans from basic and translational science, to preclinical, clinical, and epidemiology, and welcomes content that interfaces at all levels of cancer research.
Your space to connect: The Myeloid cell function and dysfunction Hub
A new Communities’ space to connect, collaborate, and explore research on Clinical Medicine and Cell Biology!
Continue reading announcementRelated Collections
With Collections, you can get published faster and increase your visibility.
Single-Cell RNA Sequencing in Cancer Immunotherapy
Cancer immunotherapy is a hot area of current oncology research, with its core focus on activating or enhancing the body's immune system's ability to recognize and kill cancer cells. However, cancer cells possess complex heterogeneity and dynamics, which affect the efficacy of immunotherapy in many ways. Single-cell RNA sequencing (scRNA-seq) has emerged as a powerful tool in recent years, providing us with an unprecedented insight into the cellular heterogeneity and dynamics within tumors. This technology has revolutionized our understanding of cancer biology, especially in the context of cancer immunotherapy. By enabling researchers to analyze individual cells, scRNA-seq allows them to identify distinct cell populations, track cellular responses to treatments, and discover new therapeutic targets. This collection aims to compile cutting-edge research in this field and explore the various applications of single-cell RNA sequencing in cancer immunotherapy.
This collection will cover the following topics: 1. The latest advances in single-cell RNA sequencing technology in cancer immunotherapy, including research on technology optimization and data interpretation; 2. Using single-cell RNA sequencing to reveal the characteristics of immune cell subgroups in the tumor microenvironment and their interaction mechanisms with cancer cells; 3. Analyzing the molecular basis of immune therapy response and resistance through single-cell RNA sequencing, exploring new biomarkers and therapeutic targets; 4. Combining single-cell RNA sequencing with clinical studies of immunotherapy to assess treatment outcomes, predict patient prognosis, and optimize treatment plans.
Keywords: cancer immunotherapy, single-cell RNA sequencing, therapeutic targets, tumor microenvironment, treatment response
Publishing Model: Open Access
Deadline: Jun 30, 2026
Computational and Analytical Methods for Multi-omics Approaches in Cancer Research
Exploring the molecular mechanisms, clinical outcomes, and their interplay is crucial for cancer research. Several recent research works have highlighted the importance of understanding inter- and intra- relationship among molecular mechanisms assayed at several layers of biological process, e.g. genomic, transcriptomic, proteomic, metabolomic, and microbiomics. Advanced analytical methods are needed to better understand the crosstalk signals among the biological processes and to discover molecular markers for future therapeutic targeting. Understanding of such molecular processes and clinical outcomes will provide a more holistic approach to investigating cancer biology and therapeutics. Overall purpose of this collection is to provide a platform for discussion on emerging analytical methodologies for studying tumor heterogeneity, diversity, progression, and treatment response with a focus on advancing our knowledge of the complex disease better.
Considering the recent advances in statistical, bioinformatics, and machine learning approaches in cancer research, we propose a collection of the topics aiming to facilitate interdisciplinary dialogue and collaboration among researchers working in quantitative and clinical aspects of cancer biology and therapeutics. Authors are encouraged to submit articles that investigate the molecular mechanisms underlying the disease heterogeneity and progression, as well as studies that explore the potential therapeutic strategies targeting these interactions. We welcome submission of original research articles addressing the following themes but not limited to: (1) new analytic methods covering both traditional statistical, bioinformatics, and advanced machine learning methods, (2) review of the literature and perspectives, (3) tutorial on any recently developed analytical methods, (4) imaging and artificial intelligence analysis methods, etc. Submission of the articles from all types of cancer research using any type of quantitative analysis methods is welcome.
Keywords: tumor heterogeneity, statistical methods, bioinformatics, machine learning, cancer biology
Publishing Model: Open Access
Deadline: Jul 31, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in