The Forty Year Journey of Cupriavidus smyrnaeus: Honoring the Pearl of the Aegean in Microbial Taxonomy
Published in Ecology & Evolution and Microbiology

Cupriavidus smyrnaeus is an oxalotrophic Gram-stain-negative bacterium with a metabolic profile relevant to biogeochemical carbon and calcium cycling in soil ecosystems. Oxalotrophic bacteria are well recognized for their role in the oxalate–carbonate pathway, a process that mediates long‑term carbon sequestration by transforming oxalate into carbonate minerals.
The species epithet smyrnaeus derives from Smyrna — the ancient Greek name for İzmir, the Aegean city in Türkiye, where the strain TA19 was isolated, and initial characterization experiments were carried out. It is a small tribute to a place that shaped this research, and to the long tradition of embedding geography into microbial taxonomy. Behind every Latin epithet, there is a story.
The journey began in 1980 at the LAMUN in Switzerland. My doctoral co-advisor, Professor Emeritus Dr. A.U. Tamer (Celal Bayar University, Türkiye), began studying the taxonomy of oxalotrophic bacteria (microorganisms that can use oxalate, a simple organic acid found in plants and animals, as a carbon source) under the supervision of Prof. M. Aragno. Strain TA19 was first isolated in 1982 from decaying plant litter near an Arum sp. It was one of the first isolates he obtained after completing his research and returning to Türkiye. He carefully lyophilized (freeze-dried) the strain and preserved it for future study.
The first mention of strain TA19 in the scientific literature came in 1988 as a non-H2-lithotrophic, oxalate-oxidizing bacterium. Years later, when I began my doctoral studies in 1997, I chose oxalotrophic bacteria as my thesis topic, inspired by Dr. Tamer's work. Strain TA19 was included alongside my own oxalotrophic isolates, serving as a reference strain. I grew fond of this isolate—not just for its oxalotrophic metabolism, but because it represented a lineage of research passed down from mentor to student.
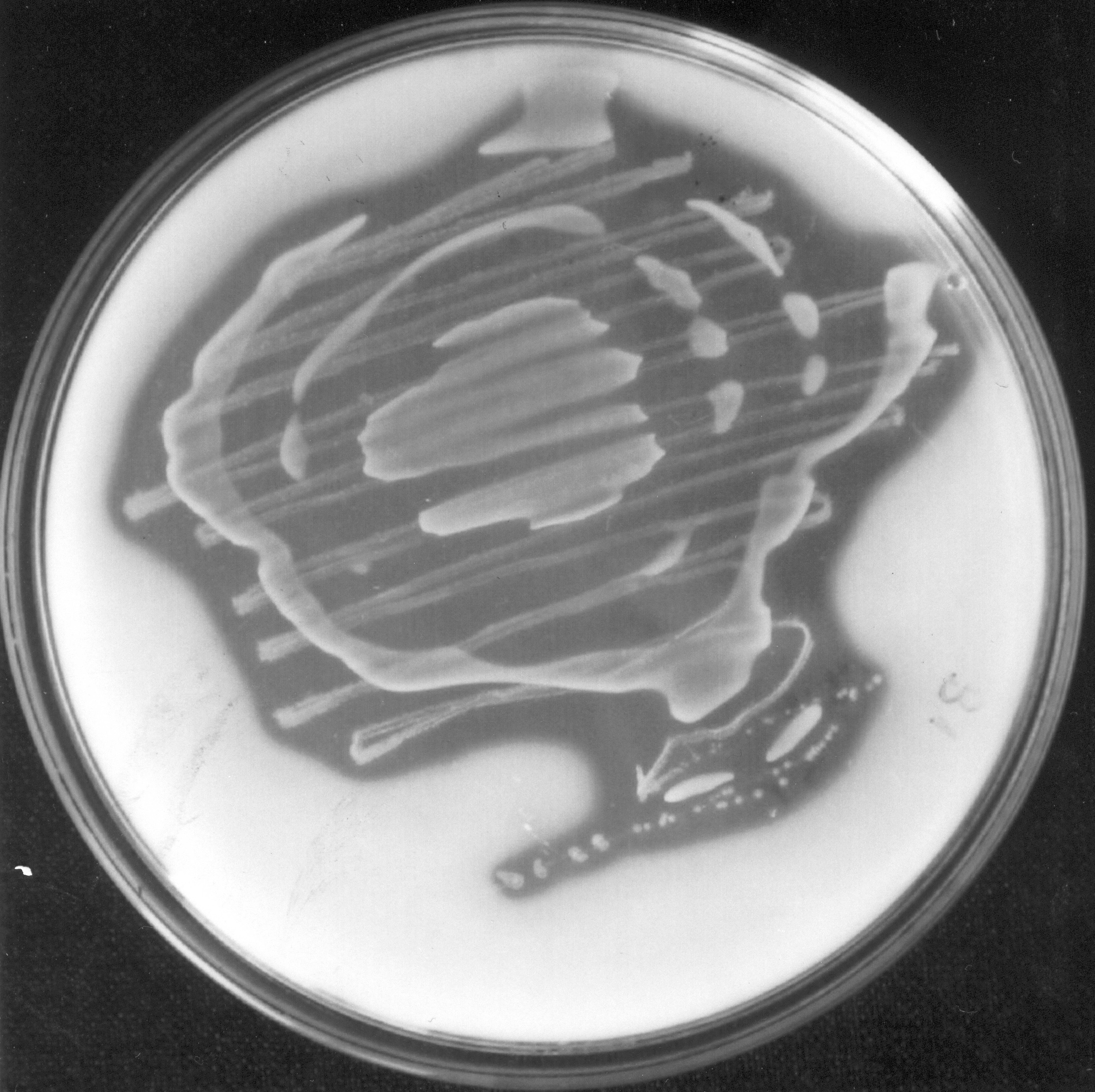
Clearing zones on Ca-oxalate medium
After completing my PhD thesis in 2002, I felt strongly that strain TA19 deserved recognition as a new species. It wasn’t just another strain; it had unique characteristics that set it apart. However, publishing a new species is never straightforward. The gold standard for defining bacterial species was wet lab DNA-DNA hybridization, combined with chemotaxonomic analyses such as fatty acid methyl ester (FAME) profiling and quinone composition. I began working on strain TA19 using these approaches, confident that they would be sufficient.
By 2016, I had prepared a manuscript describing strain TA19 as a novel Cupriavidus species and submitted it for publication. Yet the field was changing: whole‑genome sequencing had become a prerequisite for species descriptions, and editors were increasingly hesitant to accept new species description papers based on single‑strain studies, except in rare cases. As a result, our manuscript stalled, and revisions were suspended.
In 2018, I had the opportunity to go to the Institute of Plant Science and Resources (IPSR), Okayama University, as a visiting researcher under the supervision of Dr. A. Tani, where we sequenced the genome of strain TA19. This collaboration was crucial-it provided the missing piece that systematic microbiology now demanded. The genome data not only validated strain TA19’s uniqueness but also aligned the work with modern standards of species description. This international collaboration underscored how science often moves forward: through networks of researchers, shared resources, and mutual support. Without access to genome sequencing, strain TA19 would have found its way into print eventually — but anonymously, folded into a comparative numerical taxonomy study alongside other oxalotrophic Cupriavidus isolates, its distinct identity forever overlooked.
For me, strain TA19 is more than a bacterium. It represents a bridge between past and present, mentor and student, old methods and new technologies. It reminds us that science is cumulative, built on the work of those who came before us. It underscores the importance of persistence-sometimes the most meaningful discoveries take decades to be recognized.
I hope this inspires others in the systematic microbiology community to value both the old and the new, preserve isolates, and continue the long, sometimes winding journey of discovery. In the end, Cupriavidus smyrnaeus’s story is a reminder that microbes, like humans, have histories. They are discovered, studied, sometimes forgotten, and occasionally rediscovered. By publishing strain TA19 as a new species, we have ensured that its history is not lost. It now has a name, a place in microbial taxonomy, and a story that connects generations of microbiologists.
Science often demands speed, novelty, and immediate impact. However, sometimes the most meaningful contributions come from patience, persistence, and honoring the past. Cupriavidus smyrnaeus is proof of that.
Follow the Topic
-
Current Microbiology
Current Microbiology is a renowned scientific journal committed to advancing Microbiology. It delves into the realms of prokaryotic and eukaryotic cells, viruses, and the intricate interplay among microorganisms, hosts, and their environment.
Related Collections
With Collections, you can get published faster and increase your visibility.
Novel Prokaryotic Taxa
Publishing Model: Hybrid
Deadline: Ongoing


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in