The Nature of Human Behaviour
Published in Social Sciences

If you ever get close to a human and human behaviour
Be ready, be ready to get confused
Björk - Human Behaviour
In genetics, we talk about complex traits when we refer to traits that are influenced by a combination of many genes with small effects and the environment. Human behavioural traits are among the most complex phenomena we have encountered, likely impossible to explain in terms of their biological components. That would be like trying to explain a song in terms of the physical components of the musical instruments. Nevertheless, me and professor Karin Verweij just published a paper in Nature Human Behaviour where we describe the steps that need to be taken to improve our understanding of the complex relationships between DNA and human behaviour. We acknowledge that describing all the biological, environmental, and social pathways in between our DNA and our behaviour is a daunting task that is not likely to be completed anytime soon. Even though we are not likely to entirely close the gap between DNA sequence and behaviour in our lifetime, it is both worthwhile and necessary to invest our time and resources in better understanding the nature of our behaviours.
The ability of living organisms to develop, grow, function, and reproduce is encoded in the self-replicating DNA molecules they carry around in every cell of their bodies. These genetic instructions have gone through gradual mutational changes as organisms reproduced throughout the last four billion years. While these mutations are random, only mutations that increase chances of survival and reproduction stand the test of time, making different organisms slowly adapt their behaviours to a variety of environmental circumstances on this planet.
After about three billion years of these gradual changes, some organisms started to evolve brains, which helped them to adapt more quickly to their environment by rapidly adjusting their behaviours to immediate circumstances. Brains also allowed many animals to replicate beneficial behaviours from other members of their species. Humans evolved brains with the capacity to share ideas and opinions about their own behaviour and the behaviour of the world around them. This drastically increased the impact that humans can have on behaviours of their own and of their surroundings. Human populations have the ability to collectively understand, control, and create environments that exert selection pressures on the DNA of plants, animals, and people around them. Consequently, the earth and the plant and animal life it carries have gone through unprecedented changes since the recent explosive increases in human population sizes.
It took about four billion years for DNA to develop creatures that figured out how evolution works and are able to read out and directly influence their own DNA. On an evolutionary timescale, we are in the process of detonating another evolution revolution akin to the Cambrian explosion. The explosion of new species during the Cambrian was driven by the advent of multicellular life. The next one will likely be driven by our ability to consciously tamper with our genetic make-up. Currently, however, our ability to tamper with our genetic make-up is progressing more quickly than our ability to understand how our DNA sequence influences our behaviour.
For two decades, we have been able to read the sequence of large amounts of human genomes all across the world, and more recently even those of many of our ancient ancestors. We have learned much in these two decades about who we are and where we came from. The vast majority of genetic data, however, is still just a massive collection of unintelligible sequences of the nucleotides A, G, T, and C. One of our main strategies in decoding these sequences is the so-called genome-wide association study (GWAS), in which we quantify the relationships between millions of single nucleotides and complex traits using large datasets of DNA from hundreds of thousands to millions of people.
One of the main lessons learned from the thousands of GWAS publications so far is that the vast majority of complex traits are influenced by a very large amount of genetic variants, each with a very small effect (i.e., these traits are highly polygenic). We are making increasingly better estimates on how big (or small) the effects are on behavioural outcomes for each of these genetic variants. This is an incredible scientific accomplishment, but we still understand very little about the nature of these associations. Our Nature Human Behaviour article reviews steps we need to take in order to better understand where these signals come from.
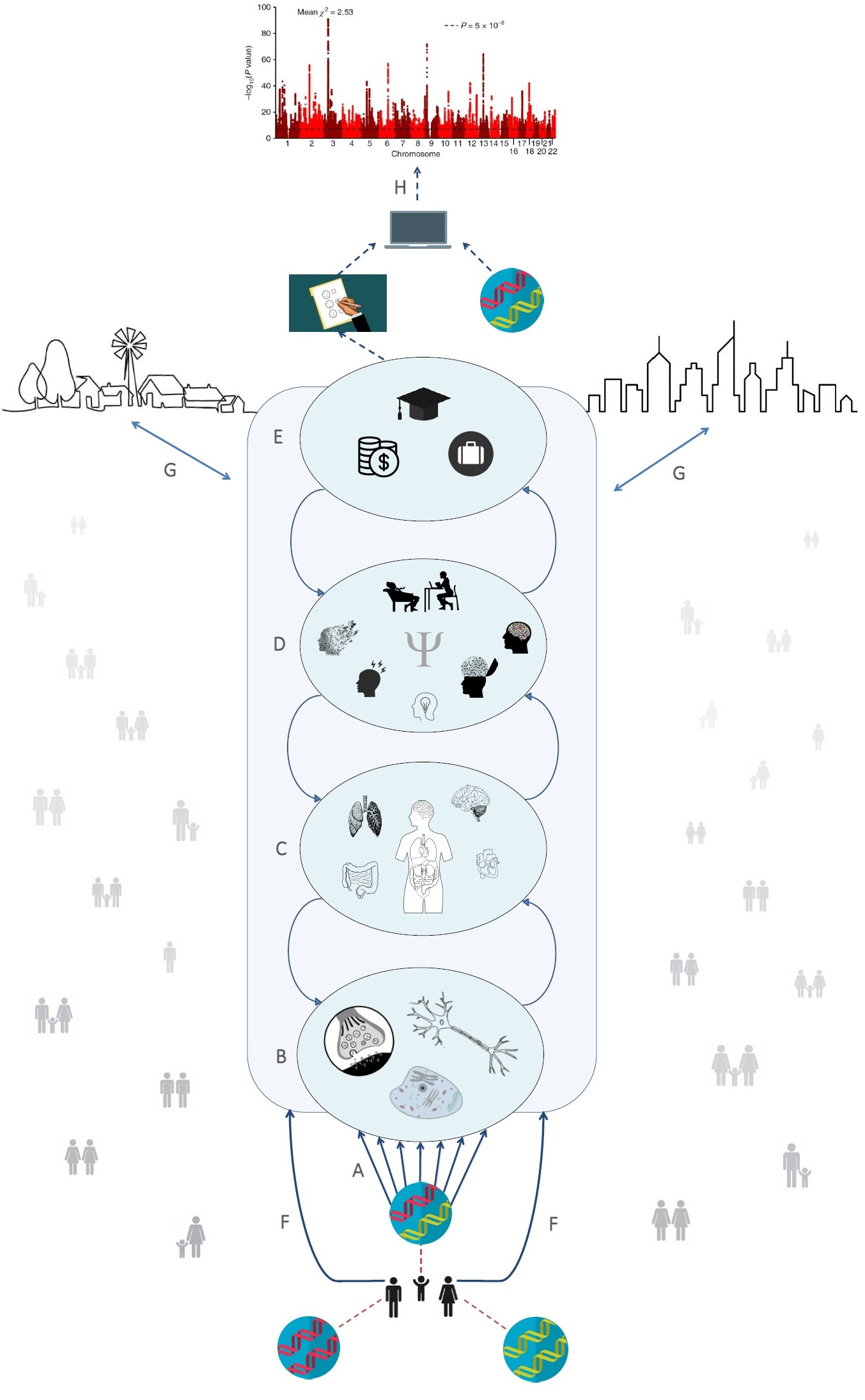
Figure: Polygenic effects (A) travel through many cascades of biological processes that react to environmental stimuli to end up associated with behaviour. Proteins built as a result of instructions in the DNA perform vital functions on the cellular level (B). Complex networks of cells make up organs, complex network of organs make up the human body (C), and complex network of humans make up society (G). Individual differences in biological make-up (B, C) in combination with environmental differences in exposures from family (F) and society (G) explain individual differences in behaviour (D). Behaviour plays an important role in determining one’s place in society, which is often measured through socio-economic outcomes such as education, occupation, and income (E). Modern behavioural genetics research has so far focused largely on associations between genetic variants and complex behavioral outcomes (H, which shows GWAS results for educational attainment based on DNA from 1.1 million people). In our Nature Human Behaviour article we outline how to disentangle the processes in between.
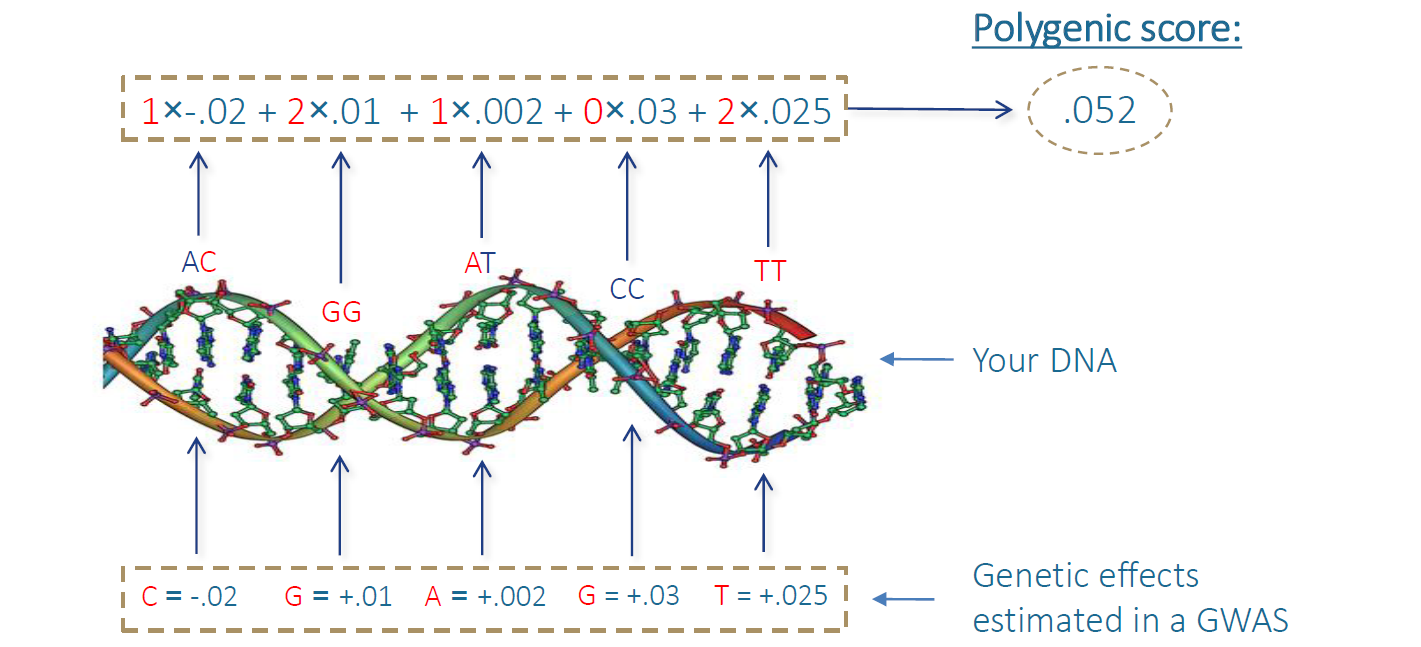
We do not need to understand anything about the biology underlying these genetic associations to use them in practice. If I know which genetic variants you carry, and I know how strong their effect is on a certain heritable outcome, I can improve my estimation of your chance of expressing that heritable outcome without knowing anything about the underlying biological processes. GWAS results are usually made public, which makes it quite easy for anyone to build so-called polygenic scores that can be used to predict someone’s genetic predisposition to heritable traits, without having to understand anything about why exactly they are predictive.
The ease with which these scores can be made increases the urgency with which we need to investigate the nature of their predictive ability. As the GWAS datasets grow, the predictive power of polygenic scores increases, which increases their potential utility. A powerful polygenic score could, for example, be used to improve the efficacy of treatments and diagnoses in the clinic by predicting someone’s risk for certain mental or physical health problems. They could however also be used to try to reduce the risk for health problems for people that have not yet been born, with the potential to impact the genetic make-up of a population.
Recently, a new biotech company was founded that offers to make polygenic scores for your embryo’s and select the best one for an IVF treatment, which was widely condemned by the scientific community. The company uses polygenic scores for ten complex traits: heart disease, stroke, atrial fibrillation, breast cancer, prostate cancer, type 2 diabetes, type 1 diabetes, and inflammatory bowel disease, Alzheimer's Disease, and schizophrenia. A major problem of this approach is that the predictive ability of these polygenic scores is not purely a reflection of the genetic risk for these complex diseases alone.
As we explain in our Nature Human Behaviour article, GWAS results contain signals from other traits than the trait or disease of interest and from environmental influences. Genes that increase the risk for schizophrenia for example are known to also be associated with creativity. We showed previously in Nature Neuroscience that polygenic scores for schizophrenia are predictive for whether someone holds a creative job. Physical diseases are often also influenced by genes that are associated with behaviours. Heart disease and diabetes, for example, are strongly influenced by lifestyle and access to healthy foods and exercise. We showed in another Nature Human Behaviour article that genes that have an effect on educational attainment influence where you live; living in richer neighbourhoods means living with fewer fast-food restaurants, higher fruit and vegetable consumption, more physical activity, and therefore lower rates of heart disease and diabetes. More recently, we indeed showed that GWAS results of a wide range of physical and mental health outcomes contain signals of genetic effects on educational attainment that can be reduced by controlling for geographic location.
It is very hard to select for genes for a certain complex trait without unintentionally selecting for other traits as well. When we domesticated the tomato, we selected them on traits that would improve production, like fruit size and shelf-life, which unintentionally led to a significant loss of their taste. To return to the song-analogy, imagine only allowing musical instruments to survive that are associated with hit-songs in the current cultural context? Luckily, current polygenic scores have relatively small effects when applied to embryo selection, largely because the embryo’s come from the same two parents. A much larger effect could potentially be reached when polygenic scores are used to select offspring genes before any embryos are formed, namely during mate selection. The use of dating apps has rapidly increased in recent times, and it would not be hard to extend the increasingly sophisticated dating apps to only show potential partners with a certain genetic make-up that combines with yours to minimize the disease risk of your potential offspring. All the problems described above would apply, probably more strongly, to such an intervention on the course of our evolution.

The ease with which rapidly improving genetic effect estimates for behavioural traits can be accessed and utilized will result in opportunities to monetize polygenic scores that will be very hard to resist, risking unpredictable and irreversible consequences to our genetic make-up. This increases the urgency with which scientists need to invest in clarifying what the genetic effect estimates that will be used for these kinds of endeavours exactly mean. We describe in our Nature Human Behaviour article how we can improve our methods and data collection strategies to achieve a better understanding of the associations between our DNA and human behaviour.
Follow the Topic
-
Nature Human Behaviour
Drawing from a broad spectrum of social, biological, health, and physical science disciplines, this journal publishes research of outstanding significance into any aspect of individual or collective human behaviour.
Related Collections
With Collections, you can get published faster and increase your visibility.
Digital Media and Mental Health
Publishing Model: Hybrid
Deadline: Oct 30, 2026
Basic Psychological Needs and Well-Being
Publishing Model: Hybrid
Deadline: Nov 27, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in
I really enjoyed reading this, thank you for sharing! Exciting to see how studies of this kind evolve in the future.