The intrinsically disordered proteins (IDPs) attracted more and more interested in the last decade. Many puzzles are still along with this kind of proteins. Why these highly dynamic proteins are indispensable in the nature? What are the roles of the abundant IDPs in the higher species such as human? IDPs are usually binding with partners to carry out their biological functions. It is still unclear how IDPs could adopt both high specific and efficient binding ability. In addition, the functions of IDPs are always regulated by the post-translational modifications (PTMs). More than two-thirds of phosphorylation sites are located in the intrinsically disordered proteins or intrinsically disordered regions (IDRs).It remains great challenges to understand the regulation mechanism of PTM sites on the IDP.
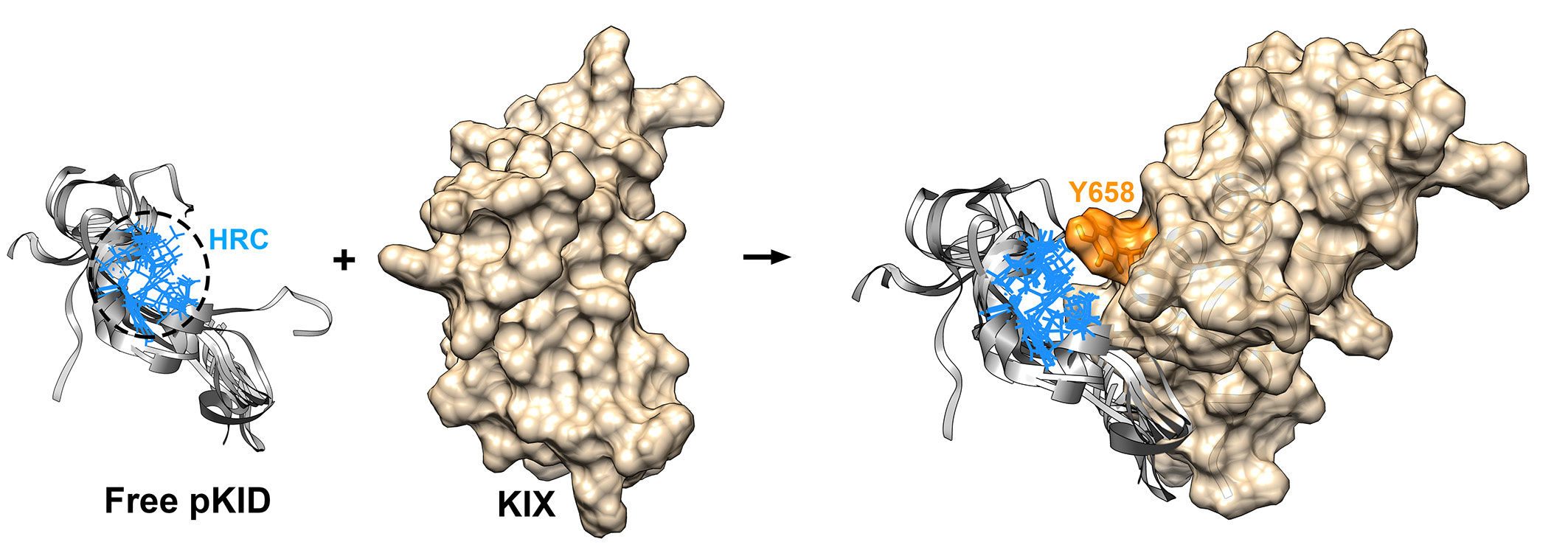
The kinase-inducible domain (KID) is an expressive example to study the structure regulations induced by phosphorylate modifications. Upon phosphorylation at Ser133, pKID undergoes disordered conformations to helical structure transition upon binding to KIX. The bound structure of pKID is formed by two α-helices, i.e. αA (from residues 120 to 129) and αB (from residues 132 to 144). The phosphorylation on Ser133 is critical for the binding between pKID and KIX, which would increase the affinity by almost two orders of magnitude.
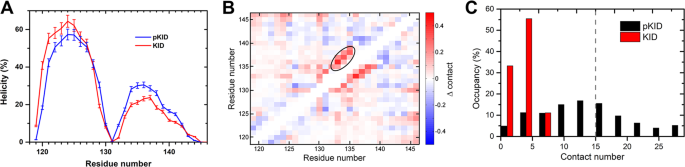
Here, we report the flexible conformational selection mechanismon the IDP binding induced by the phosphorylation. More hydrophobic interactions are formed in pKID, which promote the formation of the special hydrophobic residue cluster (HRC). The pre-formed HRC promotes binding to the correct sites of KIX and further lead the folding of pKID. Consequently, i.e., the substrate protein specifically binding with the IDP with some special locally structures, meanwhile, most of the rest regions on the IDP are flexible and dynamic. The flexible conformational selection model proposed in this work give the explanation of the high specific and efficient of IDP binding. The binding mechanism revealed in this work provides new insights into the dynamic interactions and phosphorylation regulation of proteins.
The details of this work can be found at: https://www.nature.com/articles/s42004-020-00370-5
Follow the Topic
-
Communications Chemistry
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the chemical sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Chemical modification of proteins
Publishing Model: Open Access
Deadline: Jun 30, 2026
Sustainable waste management through polymer upcycling
Publishing Model: Open Access
Deadline: Aug 31, 2026





Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in