Unlocking Genomic Variant Insights: Your Free Resource for Rare Disease (Germline) and Cancer (Somatic) Variants
Published in Protocols & Methods, Genetics & Genomics, and General & Internal Medicine

The new dawn of genomics began this year with the promise of affordable whole genome sequencing. The democratization of genomics globally will require interpretation of technologies that can adopt the mass adaptation of whole genome sequencing in clinics. As part of this new wave of large-scale genomics, GenomeArc opened a free to use ‘HorizonTM’ Variant Search Server (https://variant.genomearc.com/) for rare diseases and cancer. This is an ongoing effort from GenomeArc to connect the global genomic community with its clinical genomics knowledge hub. The variant search server will allow users to search all types of genomic variants (SNVs/Indels and long read compatible SVs) and provides a structured, evidence-backed interpretation. The pathogenicity of each germline or somatic variant is classified following the American College of Medical Genetics and Genomics (ACMG) and Association of Molecular Pathology (AMP) guidelines.
A host of proprietary databases has been integrated into the implementation of clinical genomics guidelines and is thoroughly maintained and updated by the GenomeArc R&D team. These include GenomeArc’s own rare disease and cancer sequencing cohorts, large expert-annotated databases for pharmacogenomic insights, and multiple variant databases from population controls (e.g., Middle Eastern and South Asian genome variants). Our current users from Stanford, Washington University and other global large genome institutions, provide us feedback and we update our proprietary clinical genomics and pharmacogenomics databases every 3 months.
Germline predictions primarily target rare diseases and hereditary cancers. The GenomeArc 'HorizonTM' database comprises over 5,000 rare conditions, annotated with disease inheritance (e.g., recessive or dominant) and gene mutations. The standard model, which implements ACMG guidelines for rare germline variants, evaluates whether variants are present in ClinVar and/or proprietary clinical genomic databases. Regarding hereditary cancer, our algorithm provides a comprehensive pan-cancer pathogenicity assessment. The 'HorizonTM' algorithms have been validated in numerous peer-reviewed journals, achieving a germline CNV prediction accuracy of AUC = 0.96 within known pathogenic and likely pathogenic variant cohorts. Additionally, 'HorizonTM’s application in a complex, consanguineous, rare neurodevelopmental disorder cohort achieved a 37% diagnostic yield.
Finally, somatic variant analysis precisely identifies AMP Tier 1A variants associated with FDA-approved drugs for malignant cancer. The variant.genomearc.com server, provides a comprehensive oncogenicity assessment for somatic variants following AMP guidelines. For any somatic variant, AMP tier insights can be accessed for 26 types of major malignant cancer. Apart from Tier 1A or B, 'HorizonTM' also analyze oncogenicity insights for variants that in other associated AMP tiers. The implementation of AMP tiers was a major integration effort for 'HorizonTM' to provide actionable pharmacogenomic insights at the variant level.
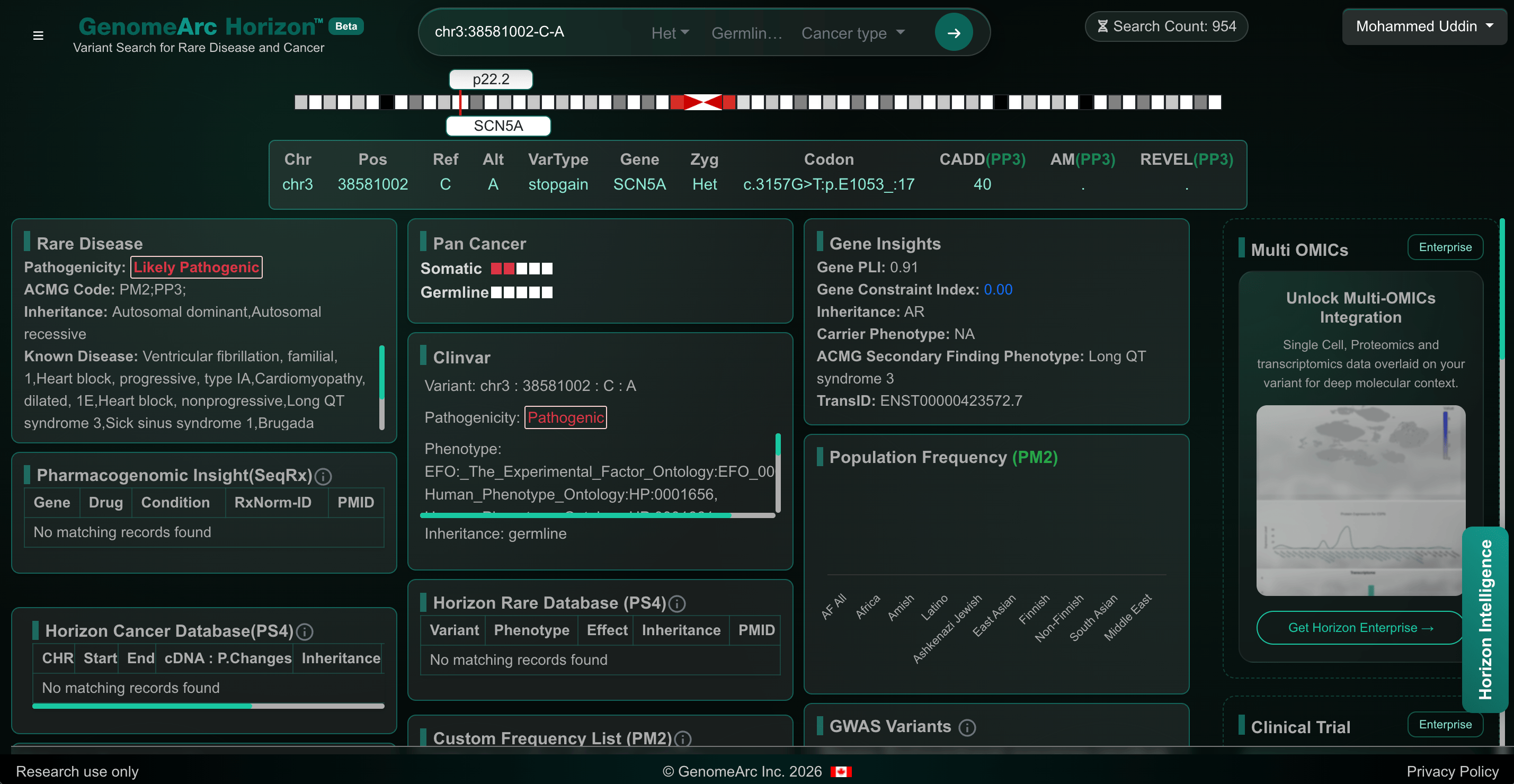 Figure 1. A variant insight snapshot from 'HorizonTM' variant search server.
Figure 1. A variant insight snapshot from 'HorizonTM' variant search server.
The tool automates variant interpretation according to the ACMG/AMP Variant (germline or somatic) interpretation guidelines by systematically evaluating genomic evidence and mapping it to the corresponding ACMG/AMP criteria. For each variant, the platform presents the supporting evidence together with the relevant evidence codes (e.g., PS, PM, PP, BS, BP, Tier 1), enabling transparent and traceable classification. This structured presentation allows clinicians to rapidly assess the rationale behind pathogenicity assignments while providing the evidence needed for reporting support.
Watch our simple tutorial of 'HorizonTM' Variant Server:
The premium version of the 'HorizonTM' software can be accessed through cloud or on prem server installation. It is an enterprise version of 'HorizonTM', that can conduct tertiary analysis of long read whole genome variants (over 5 million SNV/Indels and SVs) within 15 minutes. The somatic variant profiles AMP tiers for 26 major types of cancer, including TMB computation, somatic mutation signature, and clonal tree to trace driver mutations. The software is equipped with multiple AI agents that connect variants with phenotypes and embedded within the platform as a prompt. One of major feature of the premium 'HorizonTM' is the integration of multi-OMICs to provide insights on single cell transcriptome and proteomics. The software is designed to scale and adopt small to large scale clinical labs and research projects.
Reference:
Follow the Topic
-
Human Genetics
This is a leading international journal dedicated to publishing high-quality research in all areas of human genetics.
Related Collections
With Collections, you can get published faster and increase your visibility.
Genetic epidemiology in the era of biobanks and multi-omics: study designs, methods, and translation
Publishing Model: Hybrid
Deadline: Jul 31, 2026
Laboratory Genetics and Genomics Fellowship
The Laboratory Genetics and Genomics Fellowship Training Program is a two-year fellowship program that trains qualified individuals (MD, DO, PhD, or equivalent) to become laboratory directors with the expertise to oversee and interpret cytogenetic and molecular genetic tests important in the diagnosis and management of human genetic disorders.
Publishing Model: Hybrid
Deadline: Ongoing




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in