When a transcription factor refuses to fit the rules
Published in Biomedical Research
In immunology, transcription factors serve as molecular caretakers of cellular identity, controlling which genes are turned on or off, and ensuring developmental programs run as intended. They fit the bill, occupy their assigned rung on the developmental ladder, regulate a defined pathway, and remain largely faithful to closely related cellular identities, often performing similar functions across those lineages. However, GFI1 defies this conventional logic.
First identified in the early 1990s in screens for genes associated with growth factor–independent growth in T cell lymphoma models1, GFI1 emerged as a transcriptional repressor linked to cytokine autonomy rather than orderly lineage specification. Subsequent work established GFI1 as a key regulator of myeloid development, where it promotes granulocyte differentiation while repressing alternative lineage programs, thereby shaping myeloid cell fate decisions.
Over the last several years, our work, and the work of many others, kept bumping into GFI1 in very different immune contexts. In the thymus, during T cell development, it follows a logic familiar from myeloid systems, it blocks alternative fates and promotes T cell development2. However, in mature CD8+ T cells, it helps maintain stemness and long-term persistence3. In innate lymphoid cell populations such as ILC2s, it shows up again, this time regulating differentiation and development in yet another way. Each system seemed to reveal a different “version” of GFI1.
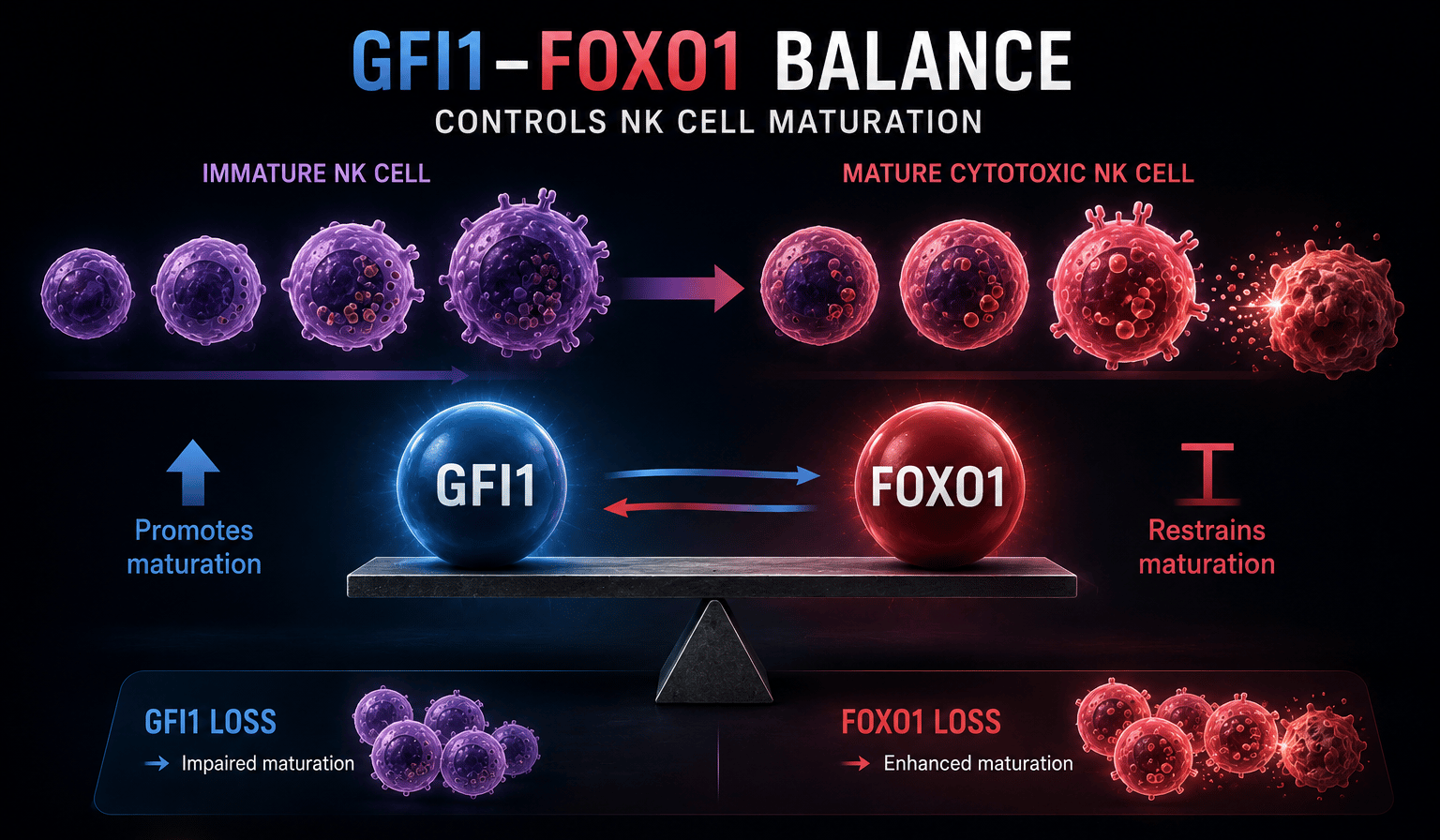
Rather than behaving like a lineage-defining transcription factor, we were intrigued by the flexibility of GFI1 functions which vary according to the biological context it finds itself in. These observations became the starting point for our recent natural killer (NK) cell study4. GFI1 proved essential for NK cell development and for the transition from immature precursors to fully functional cytotoxic effectors, shaping both their transcriptional program and epigenetic landscape. Loss of GFI1 stalled this maturation process, leaving NK cells locked in a less differentiated state with reduced effector capacity4.
NK cells rely on tightly coordinated developmental transitions to acquire cytotoxic function. Several transcription factors governing this process are well established, particularly EOMES, T-BET, and FOXO1. Among these, FOXO1 is widely recognized as a suppressor of NK cell maturation. Yet despite how frequently FOXO1 appears in NK cell biology, we were surprised to find that a comprehensive molecular footprint of FOXO1-deficient NK cells did not really exist, nor did a clear view of how it integrates into broader regulatory networks. That gap became an important turning point in the project. Instead of treating GFI1 in isolation, we started thinking in terms of transcriptional balance and co-regulation.
From there, the project expanded into something larger than we initially anticipated. By combining paired single-cell transcriptomic and epigenetic profiling, we could track how GFI1 reshapes both gene expression and chromatin accessibility during NK cell maturation. What emerged was a striking reciprocal relationship between GFI1 and FOXO1: loss of GFI1 pushed NK cells toward an immature state, while deletion of FOXO1 drove the opposite phenotype. Removing FOXO1 in GFI1-deficient NK cells substantially restored differentiation and anti-tumour capacity. At that point, the biology started to make conceptual sense. GFI1 was not simply “driving” NK cell maturation in a linear way. Instead, it functioned more like a regulatory checkpoint, calibrating transcriptional programs, and the balance between EOMES, T-BET, and FOXO1.
One of the most technically challenging aspects of the study was separating direct from indirect regulation. GFI1 is fundamentally an epigenetic repressor, yet the tools to define its chromatin-level interactions remain frustratingly limited for in vivo research. High-quality antibodies suitable for mechanistic assays are hard to obtain, and many standard approaches for mapping transcriptional repression in primary NK cells are inconsistent at best. In many ways, the project highlighted how rapidly single-cell multiomics is advancing compared to the experimental toolkit available to validate the mechanisms it reveals.
Many questions remain open. We still do not fully understand how GFI1 selectively engages different chromatin landscapes across immune cell types, nor how much of its activity is direct versus indirectly routed through broader regulatory networks. But what is clear is that GFI1 does not behave like a conventional lineage-defining transcription factor. Instead, it appears to tune immune cell fate according to context, stabilising whichever transcriptional program the system requires at that moment. And perhaps the broader lesson is that immune regulation is rarely about one-factor-one-function relationships, but that most immune regulators are less about enforcing identity, and more about keeping it adaptable.
References
1 Gilks, C. B., Bear, S. E., Grimes, H. L. & Tsichlis, P. N. Progression of interleukin-2 (IL-2)-dependent rat T cell lymphoma lines to IL-2-independent growth following activation of a gene (Gfi-1) encoding a novel zinc finger protein. Mol Cell Biol 13, 1759-1768, doi:10.1128/mcb.13.3.1759-1768.1993 (1993).
2 Yucel, R., Karsunky, H., Klein-Hitpass, L. & Moroy, T. The transcriptional repressor Gfi1 affects development of early, uncommitted c-Kit+ T cell progenitors and CD4/CD8 lineage decision in the thymus. J Exp Med 197, 831-844, doi:10.1084/jem.20021417 (2003).
3 Chaudhry, M. Z. et al. GFI1-driven transcriptional and epigenetic programs maintain CD8(+) T cell stemness and persistence. Nature immunology 26, 894-907, doi:10.1038/s41590-025-02151-5 (2025).
4 Huang, Q. et al. The GFI1–FOXO1 axis regulates NK cell maturation and function. Nature Communications, doi:10.1038/s41467-026-72022-6 (2026).
Follow the Topic
-
Nature Immunology
A multidisciplinary journal that publishes papers of the highest quality and significance in all areas of immunology. Priority is given to work that provides translational and/or fundamental insight into the workings of the immune system
Related Collections
With Collections, you can get published faster and increase your visibility.
Immune mechanisms in arthritis
Publishing Model: Hybrid
Deadline: Aug 07, 2026
Cell death and inflammatory signalling
Publishing Model: Hybrid
Deadline: Oct 28, 2026






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in