A Molecular Shredder at the Source: How Trypanosomes Fine-Tune Their Surface Coat
Published in Chemistry, Microbiology, and Cell & Molecular Biology
A 40-year mystery
To stay one step ahead of the host’s immune system, Trypanosoma brucei wears a "molecular cloak" composed of a single protein: the Variant Surface Glycoprotein (VSG). While its genetic wardrobe contains thousands of these cloaks, the parasite’s survival depends on expressing only one at any given moment while maintaining the ability to periodically shift its disguise.
The parasite’s "printer"—RNA Polymerase I—doesn't produce the instructions for this cloak in isolation. Instead, it generates a massive, polycistronic manual containing the code for the protective VSG alongside several "helper genes" known as Expression Site Associated Genes (ESAGs). These genes encode proteins that mediate critical interactions with the mammalian host, facilitating survival and immune evasion.
A forty-year mystery persisted in molecular parasitology: how does the VSG mRNA reach a 140-fold higher abundance than ESAGs co-transcribed from the same promoter? This striking discrepancy suggests a sophisticated post-transcriptional filter. How does the parasite so effectively prioritise its molecular cloak?
The "Aha!" moment
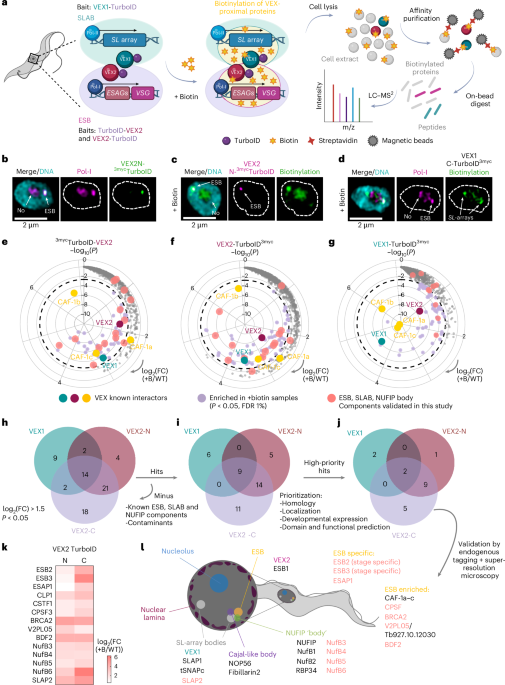
The breakthrough was a slow build of evidence. We began with quantitative proteomics to identify the proteins sequestered in the Expression Site Body (ESB)—the dedicated subnuclear "factory" where the active VSG is transcribed. We identified three novel proteins, but our attention quickly centred on a nuclease we named ESB2.
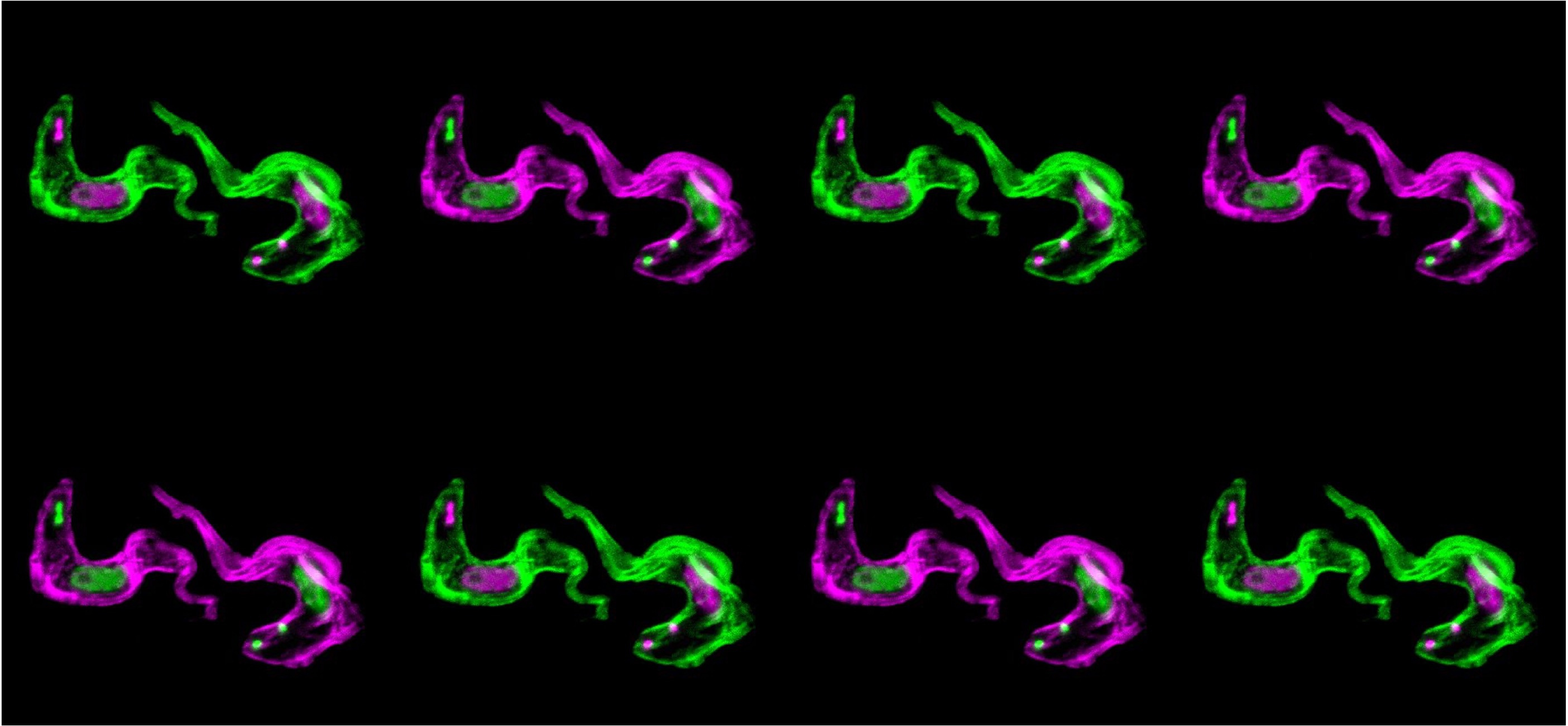
Super-resolution microscopy confirmed that ESB2 was localised exclusively within the ESB. However, the real "Aha!" moment came when we depleted ESB2 using RNAi. The result was startling: the "helper genes" (ESAGs) significantly increased in abundance, while the "cloak" (VSG) remained unaffected.
Suddenly, the pieces fell into place: was ESB2 acting as a specialised "shredder" to redact the genetic manual in real-time? To prove this, we pursued several orthogonal approaches, from using CRISPR/Cas9 to precisely abrogate nuclease activity to in vitro biochemical assays. The results were consistent: the changes in gene expression were tied directly to nuclease activity.
What we observed was entirely novel because of where it was happening: the parasite coupled transcription directly to decay within this specialised compartment. Imagine a molecular shredder stationed right inside the genetic print room, selectively redacting specific pages of a manual as they are being printed. It is a remarkably efficient way to downregulate genes at the source.
The messy reality of discovery
Of course, the final paper looks polished, but the reality was far more gruelling. There were months of failed protein purifications and a myriad of incompatible constructs. The "wet lab" effort was immense—countless transfections, dozens of cell lines, and significant proteomics endeavours.
But we will always remember the feeling of those first images in the microscope room. I still recall when Lianne sent the first results to the lab WhatsApp group. Then came the first transcriptomics data, the moment Leon confirmed the nuclease activity in vitro, and finally, when Htay observed the spatial regulation within the ESB. These are the milestones that make every late night worth the effort.
It takes a village
The asymmetrical expression within the VSG expression site had been a "cold case" in my mind since my postdoc days. But as is so often the case in science, we didn’t start with a perfect roadmap. We didn’t set out to solve this specific mystery; we stumbled into it.
While the field has largely focused on how the parasite starts making RNA (transcription), we looked at the other end: how it destroys it. Notably, this paper is more than a study on RNA decay; it is the "birth certificate" of our research group.
Starting a lab involves a significant push to establish momentum. Beyond the immediate need for results, there is a fundamental drive to define one's unique contribution to the field. It is a beautiful, but often terrifying, experiment. I vividly remember those first months: the empty benches and an intimidatingly blank whiteboard. Every setback feels magnified when you worry about the careers of the people who moved across countries to join your vision. But that shared struggle built our culture.
In the beginning, it was just Lianne and me. She moved from the Netherlands and helped me set up the lab in York from scratch—a far more daunting task than joining an established group. Then came Leon from Germany, our resident biochemist whose "German bluntness" and wit we often mock but (not so) secretly love.
The true test came at the finish line. As I began maternity leave and Lianne left academia for new ventures, Htay joined us from Singapore, bringing vital imaging expertise and a passion for condensates. Despite only a month of overlap, Htay took over with incredible dedication and mastery. Alongside Leon, they were absolute troopers, conducting critical experiments while I was away.

I navigated revisions and complex analyses while caring for a newborn—a challenge I had seriously underestimated. Refusing to sacrifice time with my baby meant working whenever he slept, resulting in even less sleep than a typical new parent gets. This paper is a testament to a team that stepped up when it mattered most, and I couldn’t be prouder of the research culture we have built.
New horizons
Writing this for Nature Microbiology is a surreal, full-circle moment. Even in a field with decades of history, there are still massive secrets hiding in plain sight—waiting for a fresh whiteboard and a bit of serendipity to find them.
By tethering RNA decay to a dedicated subnuclear factory, the parasite manages its RNA levels with incredible precision. This discovery underscores a broader principle of infection biology: pathogens don't just rely on what they "print", but on what they choose to "redact" to stay one step ahead of the immune system. For more details, read our paper in full.
Follow the Topic
-
Nature Microbiology
An online-only monthly journal interested in all aspects of microorganisms, be it their evolution, physiology and cell biology; their interactions with each other, with a host or with an environment; or their societal significance.



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in