A surprise in a museum storeroom: A 700-year-old secret hidden in a Bolivian mummy
Published in Microbiology, Genetics & Genomics, and Biomedical Research

Imagine a dusty museum storeroom in the heart of La Paz, Bolivia. For decades, shelves filled with ancient, naturally mummified remains have been resting in silence. They are the guardians of a forgotten era, but their stories, the stories of the people who lived, loved, and died in the Andes centuries before the Inca rose to power, remained locked away.
This was the starting point for our journey, a project that would eventually lead us to a discovery that rewrites a small but significant chapter of medical history. This is the story of the Bolivian Mummy Project (1), a tale not just of ancient DNA, but of resilience, interdisciplinary detective work, and a 700-year-old bacterium with a surprisingly modern face.
The Collection that almost wasn't
Our story begins at The National Museum of Archaeology (MUNARQ) in La Paz, home to one of Bolivia's most important mummy collections. These individuals from the Late Intermediate Period (1100 – 1450 AD), lived in the aftermath of the great Tiwanaku Empire. They were laid to rest in Chullpas, the funerary towers spread in the Bolivian Altiplano (2). For a long time, this collection was left behind in storage. It was a sleeping treasure with a scientific potential unrecognized due to a lack of resources and, perhaps, a clear vision.
Years ago, the museum began a monumental task: recovering and cataloging these ancestral remains. Much of their contextual information – the "where" and "how" they were found – had been lost over time, a common situation and a big challenge in modern archaeology (3). But what remained was invaluable: the people themselves.
Our initial idea was straightforward. We had secured the invaluable support of the Marie Curie – Seal of Excellence grant, the Province of Bolzano, and the Bolivian Ministry of Cultures that allowed us to launch an ancient DNA project at the Eurac Research, Institute for Mummy Studies in Bolzano, Italy. We wanted to see if we could extract genetic information from these fragile human remains. But as we stood in the museum, looking at these mummies, we realized we had a chance to do something far more meaningful. We decided to abandon a simple, narrow approach for a holistic one. This wouldn't just be an aDNA project; it would be a bioarchaeological deep-dive (4).
We asked ourselves: What could the new bioarcheological approach (e.g. CT scans) tell us about their lives and health? What diseases might they have carried? What ancient and modern populations from South America are genetically closely related to these mummies? And, critically, how could we ensure the long-term preservation of these ancestors for future generations in Bolivia? This last question was fundamental, leading us to carefully navigate the ethical landscape of working with human remains, ensuring our approach was respectful, collaborative, and responsible (5).
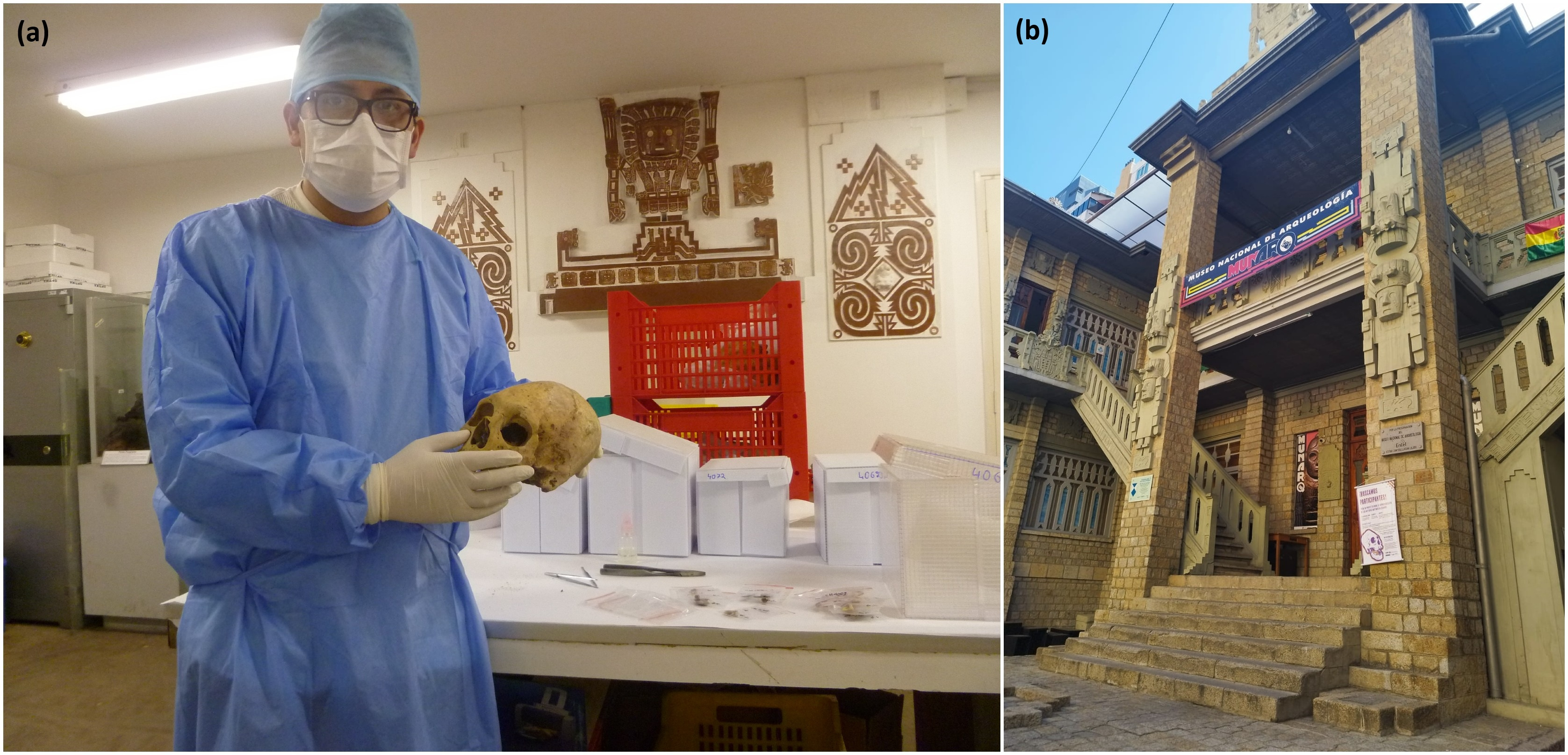

(b) The MUNARQ houses the most important collection of pre-Columbian human remains from Bolivia (Credits: G. Valverde)
The Final Day: A tooth that changed everything
The first step was a leap of faith. In June 2018, we began a sampling campaign to test the DNA preservation. I still remember those days vividly. We worked side-by-side with our colleagues from the museum, and we were all aware of the privilege we had been given. Securing the permits to sample such precious, irreplaceable material was no small achievement. Our guiding principle was simple: do no harm. Every action, every touch, was measured to avoid any potential damage to the integrity of these priceless individuals.
The days were exhausting. We moved methodically from one individual to the next, carefully collecting small samples, always mindful of the weight of our work. By the final day, we were running on fumes. It was late in the evening, and our eyes were heavy with exhaustion. As we were about to pack up, we spotted one last individual, a mummy that had been patiently waiting its turn. We looked at each other. Should we push forward?
Something told us we had to make one final effort. As we examined the remains, we noticed a loose tooth. It was a gift. Without causing any damage, we were able to collect it, a small, unassuming sample that almost didn't get taken. We packed it away with the others, not knowing that this last-minute decision would become the core of this research. The rest, as they say, is history.
Searching for everything and nothing
After the sampling campaign, we began screening for DNA preservation at the Institute for Mummy Studies in Bolzano. After centuries in the Chullpas, decades in museum storage, and countless environmental shifts, we expected the genetic material to be degraded beyond use. To our surprise, it wasn't. The DNA was remarkably well-preserved, whispering stories we were only beginning to learn how to hear.
With renewed excitement, we scaled up. We processed over 300 samples from 60 mummies. This gave us a vast ocean of genetic data, but it was an ocean filled with the DNA of everything from the human host to soil microbes and contamination most likely. Finding a needle in a haystack would be an understatement. This is where modern bioinformatics became our treasure map. We used sophisticated computational tools designed to surf through millions of genetic sequences and flag anything that didn't belong to the human host.
And then, we saw it!
A clear, unmistakable signature emerged from the noise: Streptococcus pyogenes, the bacterium responsible for everything from strep throat to more severe invasive diseases. It wasn't just a faint trace; we had enough data to reconstruct a near-complete genome of this ancient pathogen, applying a unique approach called de-novo assembly, a method to put all DNA sequences without a genome reference (6). We felt a rush of adrenaline. The "easy part" – the discovery – was done. But the real work, the part that would consume us for a couple of years, was just beginning.
_ab2.jpg)
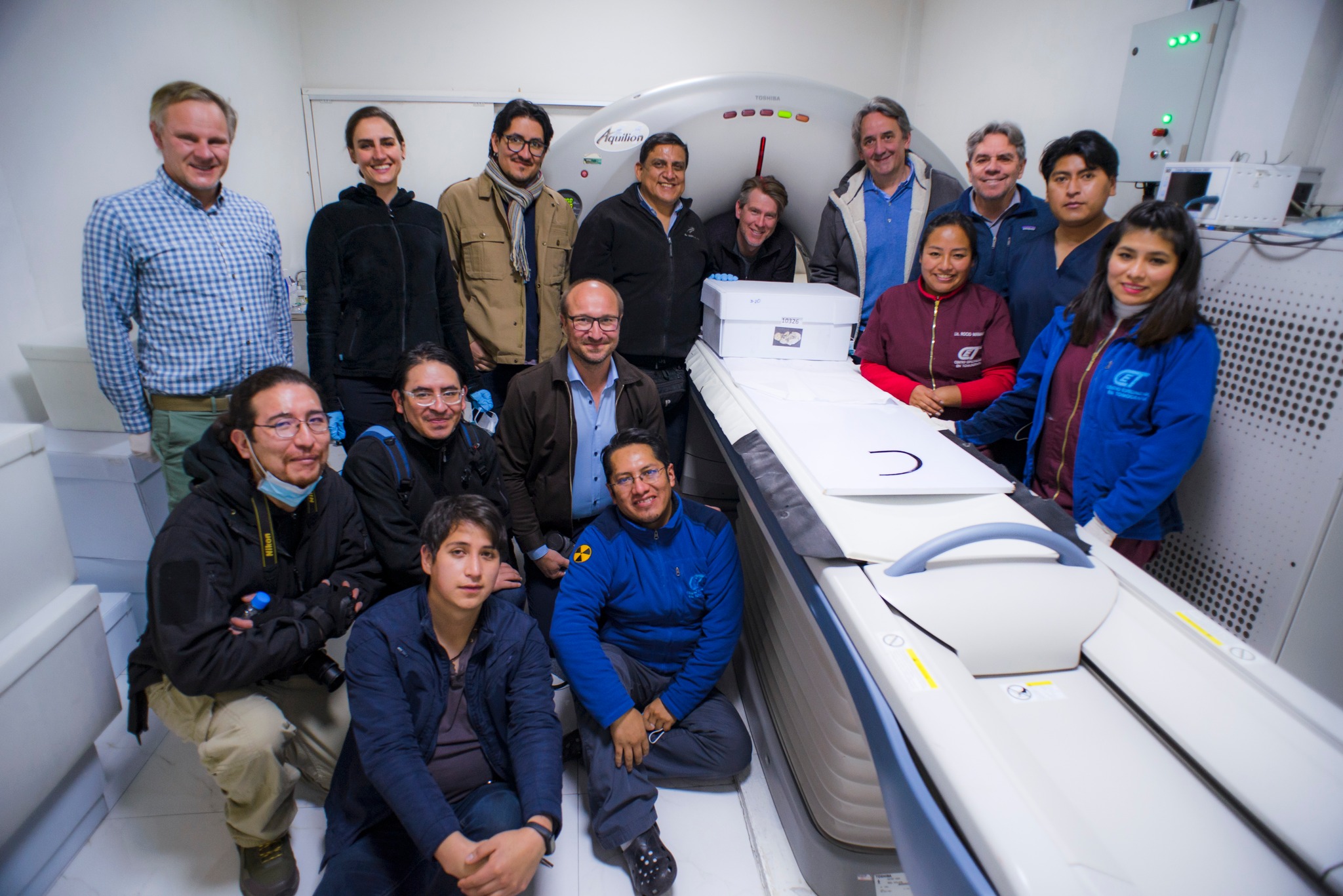
(b) Diving deep into the project’s genetic ocean, where the data is endless and the coffee is essential (Guido Valverde, Frank Maixner, Mohamed S. Sarhan) (Credits: G. Valverde)
Unraveling the secrets of an ancient microbe
Now the challenge started. We had successfully reconstructed the genome of an ancient pathogen, but what did it mean? This is where the story became a fascinating puzzle.
Importantly, we had to first check the time frame of this individual who lived in the Andean region. Radiocarbon dating came back, confirming the individual's pre-Columbian time period and placing it firmly in the Late Intermediate Period. That was a deeply satisfying moment – science validating science. We then continued with the stable isotopes analyses, which results helped us to understand the type of diet this individual has had in the past, basically relying on tubers, like potato and quinoa, but mostly maize, and camelid meat, specific food resources in the Andean highlands of Bolivia.
Going back to the genes, our analysis revealed that this 700-year-old bacterium carried the core genes that make S. pyogenes so virulent today. However, there was a critical difference. Modern strains often carry "prophages" – viral elements integrated into the bacterial genome – that act as delivery systems for enhance their virulence. Although the ancient strain surprisingly harbored also these prophages, they lacked those particular toxin-carrying profile.
But the most profound revelation came when we placed this ancient genome on the family tree of S. pyogenes. It settled at the very base of the tree, near the common ancestor of all modern strains. This suggested that the lineages of this pathogen we see today, causing millions of infections annually, diversified relatively recently.
To truly understand the timescale, we performed one more sophisticated analysis: the molecular clocking. Using complex simulations, we were able to estimate when this ancient bacterium circulated. The results gave us the overall glimpse we were hoping for, suggesting a global diversification within the last 5,500 years – a blink of an eye in evolutionary terms.
_ab.jpg)
(b) Ancient DNA lab work in a fre-contamination room at the Institute for Mummy Studies (Credits: G. Valverde)
Rewriting a chapter of pre-Columbian history
This one loose tooth, from one mummy sampled on a tired final evening in a museum in La Paz, has essentially changed our understanding of this particular disease in the pre-Columbian Americas. It pushes back the confirmed presence of Streptococcus pyogenes on the continent by several centuries. But the deeper implication is even more significant. For a long time, the narrative of pre-Columbian health has been overshadowed by the devastating epidemics brought by European colonizers. This discovery proves that S. pyogenes was already here, circulating among Indigenous populations long before the European upheaval. It was a part of their disease landscape, a silent companion that likely caused illness and, in some cases, death, in a world we often mistakenly imagine as a pristine paradise.
Our work is still ongoing, as we have a couple of extraordinary findings in the pipeline. We are filling a critical gap in South America by reporting such ancient pathogen in Bolivia, and we hope this finding can provide a better understanding of outbreaks in the Andean region. It sets the stage for future discoveries that can illuminate the health conditions, outbreaks, and resilience of Indigenous populations.
For us, this project was not just about a genome. It was about the incredible potential of that museum collection in La Paz, which sat waiting for decades. It was about the power of looking beyond a single discipline to embrace bioarchaeology, ethics, conservation and genomics. It was a testament to the idea that the past is not a dead place, but a living archive, waiting to reveal its secrets to those who approach it with patience, respect, and a willingness to push forward – even on the final, exhausting day of sampling.
The mummies from Bolivia are no longer silent. They have been waiting for centuries to be listened to. And the story they are telling is just the beginning.

REFERENCES
- Valverde, G., Niederkofler, J., Paladin, A., Samadelli, M., Castedo, L., Maixner, F., & Zink, A. (2022). The Bolivian Mummy Project: Different lines of scientific evidence for the study of pre-Columbian mummies at Museums. In 10th World Congress on Mummy Studies, Bolzano.
- Ponce Sanginés, C. & Linares Iturralde, E. (1966). Comentario Antropológico acerca de la determinación paleoserológica de grupos sanguíneos en momias prehispánicas del Altiplano Boliviano, La Paz: Academia Nacional de Ciencias de Bolivia.
- Arriaza, B. T., Cartmell, L. L., Moragas, C., Nerlich, A. G., Salo, W., Madden, M., & Aufderheide, A. C. (2009). The Bioarchaeological Value of Human Mummies Without Provenience. Chungará (Arica), 40(1), 55–66.
- Valverde, G., Paladin, A., Samadelli, M., Conti, V., Castedo, L., Zink, A., & Maixner, F. (2025). Bioarchäologie in Bolivien - Die Mumien aus dem Museo Nacional de Arqueología en La Paz. Antike Welt: Zeitschrift für Archäologie und Kulturgeschichte, 2025(1).
- Conti V., Liu J., Valverde G., Castedo L., López L., Mamami R., Noriega I., Maixner F., Zink A., Wilkinson C., Paladin A. (2025). Integrating participatory praxis for ethical museum dissemination of ancient human remains: The case of the Bolivian National Museum of Archaeology-MUNARQ Mummy Collection, 11th World Congress on Mummy Studies, Cusco, Peru.
- Valverde, G., Sarhan, M. S., Cook, R., Rota-Stabelli, O., Adriaenssens, E. M., Zink, A., & Maixner, F. (2026). An ancient genome of Streptococcus pyogenes from a pre-Columbian Bolivian mummy. Nature Communications. https://doi.org/10.1038/s41467-026-71603-9
Follow the Topic
-
Nature Communications
An open access, multidisciplinary journal dedicated to publishing high-quality research in all areas of the biological, health, physical, chemical and Earth sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Women's Health
Publishing Model: Hybrid
Deadline: Ongoing
Biosensing
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in