Advances in infectious disease diagnostics – how can they help us achieve SDG3.3?
Published in Microbiology and Sustainability
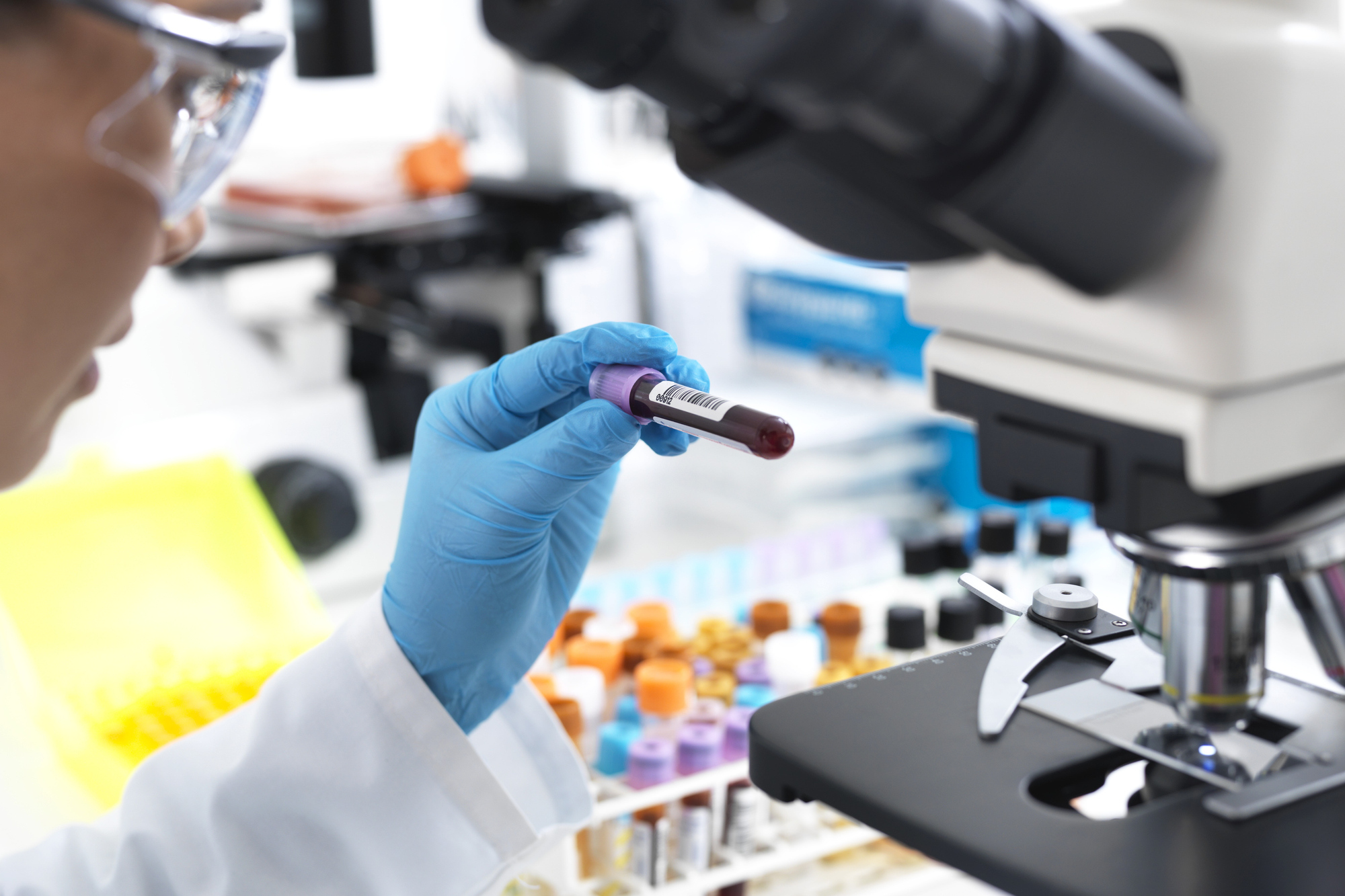

Explore the Research
Infectious disease diagnostics
This Collection welcomes original research into current challenges and advances within the field of infectious disease diagnostics.
Infectious diseases are responsible for millions of deaths every year, with the global impact of the COVID-19 pandemic still felt by many. While many lessons have been learnt, there is still concern that some of the vulnerabilities that allowed COVID-19 to spread remain, particularly in those most at risk from the disease.
As we creep closer to 2030, the United Nations’ Sustainable Development Goals are more relevant than ever, and while some goals seem within reach, it is becoming clear that some may be unattainable. Target 3.3 aims to end epidemics of communicable diseases including AIDS, Tuberculosis and Malaria. In 2024, Malaria alone accounted for an estimated 610,000 deaths worldwide.
Why are diagnostic tools so important?
Over the last decade, diagnostics for infectious diseases have advanced, with a particular focus on early diagnosis. Tools that focus on early diagnosis have positive implications for patient outcome and morbidity rates, by ensuring proper treatment is provided as soon as possible.
Additionally, early diagnosis can prevent infectious diseases from spreading and provides important information for public health surveillance. An example of this is the recent Meningitis B outbreak in the UK.
Meningitis can occur from different pathogens (including bacteria, viruses and fungi), meaning specific differential diagnosis is vital. Suspected Meningitis is usually confirmed through PCR testing on a sample of cerebrospinal fluid (CSF), taken via a lumbar puncture. Although it has not been specifically confirmed for the UK outbreak, we can assume that unless contradicting symptoms were shown, any patient with suspected bacterial Meningitis received a lumbar puncture on admittance. This protocol enables healthcare professionals to provide the correct treatment as soon as possible after a Meningitis patient enters the hospital.
Additionally, anyone who visited the site of infection has been encouraged to undergo preventative treatment, including vaccines and antibiotics, before showing symptoms. The fast response to this outbreak has meant the spread is contained, with no new cases being confirmed in recent updates.
By improving diagnostic tools, healthcare providers, doctors and researchers can continue to detect infectious diseases early, reducing their spread and impact and potentially saving many lives before a disease can progress.
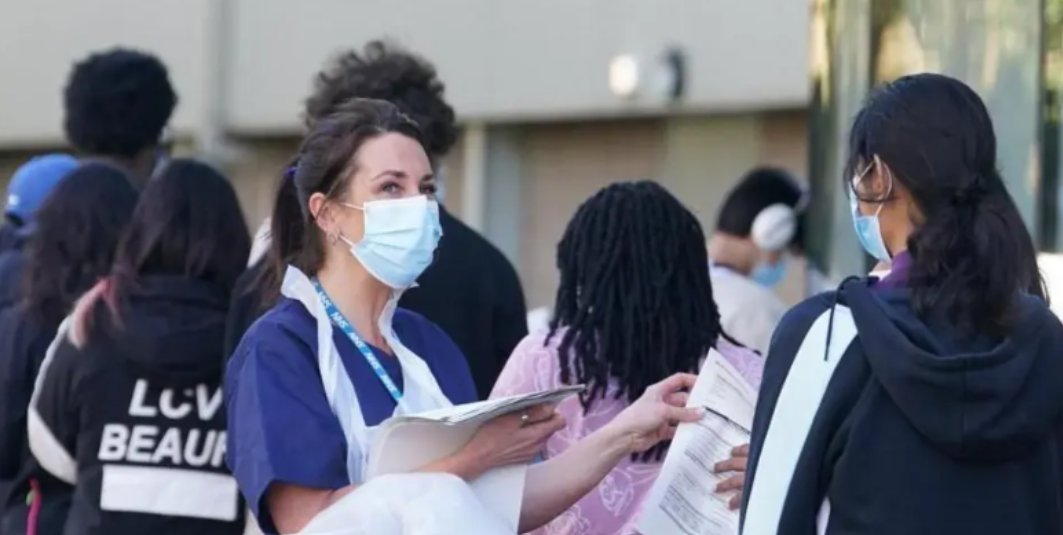

Students waiting for Meningitis B vaccines in Kent.
How have diagnostic tools evolved?
For many years, culture-based assays were considered the gold standard. This method involves growing and detecting specific microorganisms from clinical samples (blood, urine, etc.), to discern whether an infectious disease is present. They are robust methods, relatively simple to assemble/use, and can sometimes be low in cost, making them a valuable tool in low- and middle- income countries. More advanced diagnostic tools can be difficult to implement here due to infrastructural and financial challenges. However, culturing certain pathogens is expensive, and the risk of contamination is high. Although culture-based methods remain essential, especially for antimicrobial susceptibility testing, many settings now complement or partially replace them with faster molecular tools.
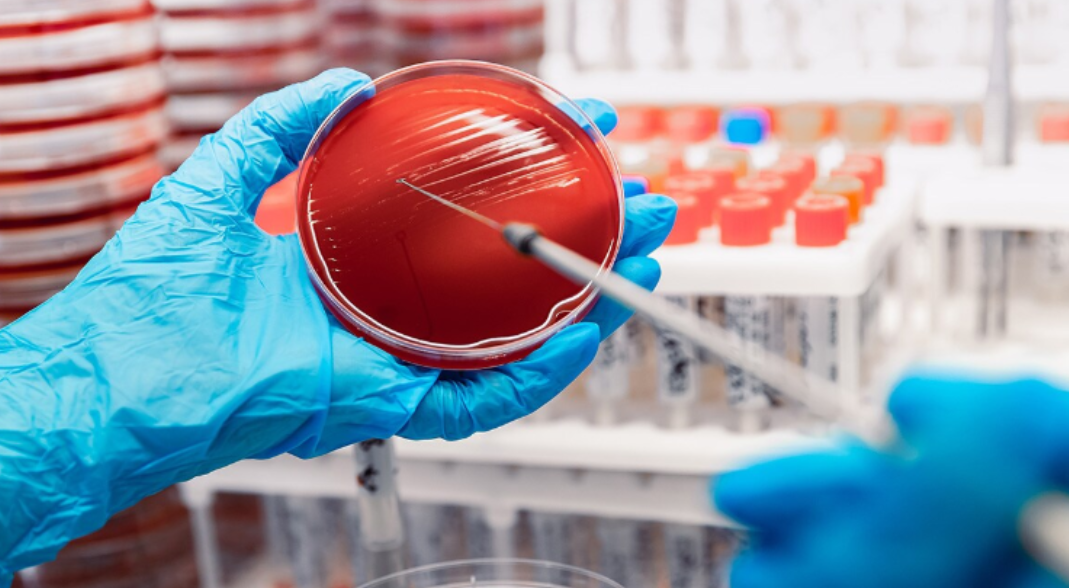

In recent decades, molecular biology advances have taken precedence as ‘diagnostic tools for the new millennium’. One example is the Polymerase Chain Reaction (PCR), involving targeted amplification using DNA templates, or cDNA generated from RNA templates, to detect pathogenic organisms or viruses. PCR has transformed infectious disease diagnostics but relies on specialised equipment and trained staff, which limits its use in low-resource settings. During the COVID-19 pandemic, PCR testing was heavily relied upon for disease surveillance. Drive-in settings for swab testing were hugely popular and had a big impact on detecting and containing infections early. Waiting 1-2 days for a result via text was a process familiar to most of us, and though rapid antigen tests were also available, it was generally accepted that the PCR test was much more reliable. PCR is a lot faster than culture-based assays and can detect extremely low levels of DNA/RNA. However, the high sensitivity that allows it to be so specific does also mean only a tiny fragment of contaminating DNA can lead to false-positive results.
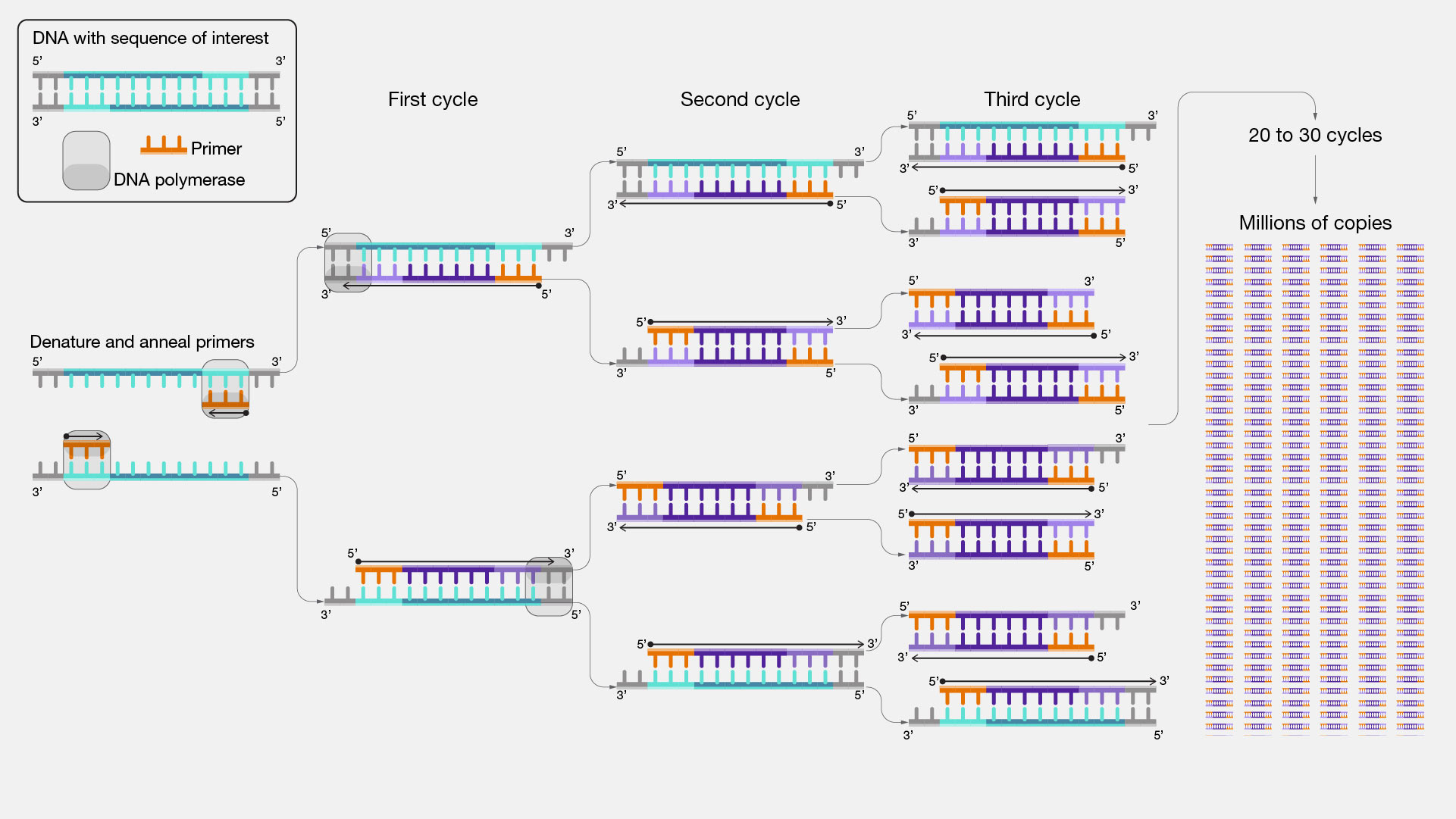
Polymerase Chain Reaction (PCR).
Multiplex PCR, where multiple target sequences can be amplified simultaneously, has also risen in popularity in recent years. Results from these tests can be obtained much sooner than from culture, and they allow simultaneous detection of multiple pathogens from a single sample. The ability to test for more than one disease at once has a cost-effective appeal to many laboratories, and promises savings on dye, reagents, dNTPs, and other essential requirements. It also gives doctors the ability to rule out diseases, avoiding mistreatment and unnecessary antibiotic use. Furthermore, time is a vital factor in effective disease surveillance, to ensure infected patients can be quarantined as soon as possible to mitigate spread. An additional benefit of multiplex PCR is to identify whether the pathogen is viral or bacterial quickly – reducing the prescription of unnecessary antibiotics in some cases.
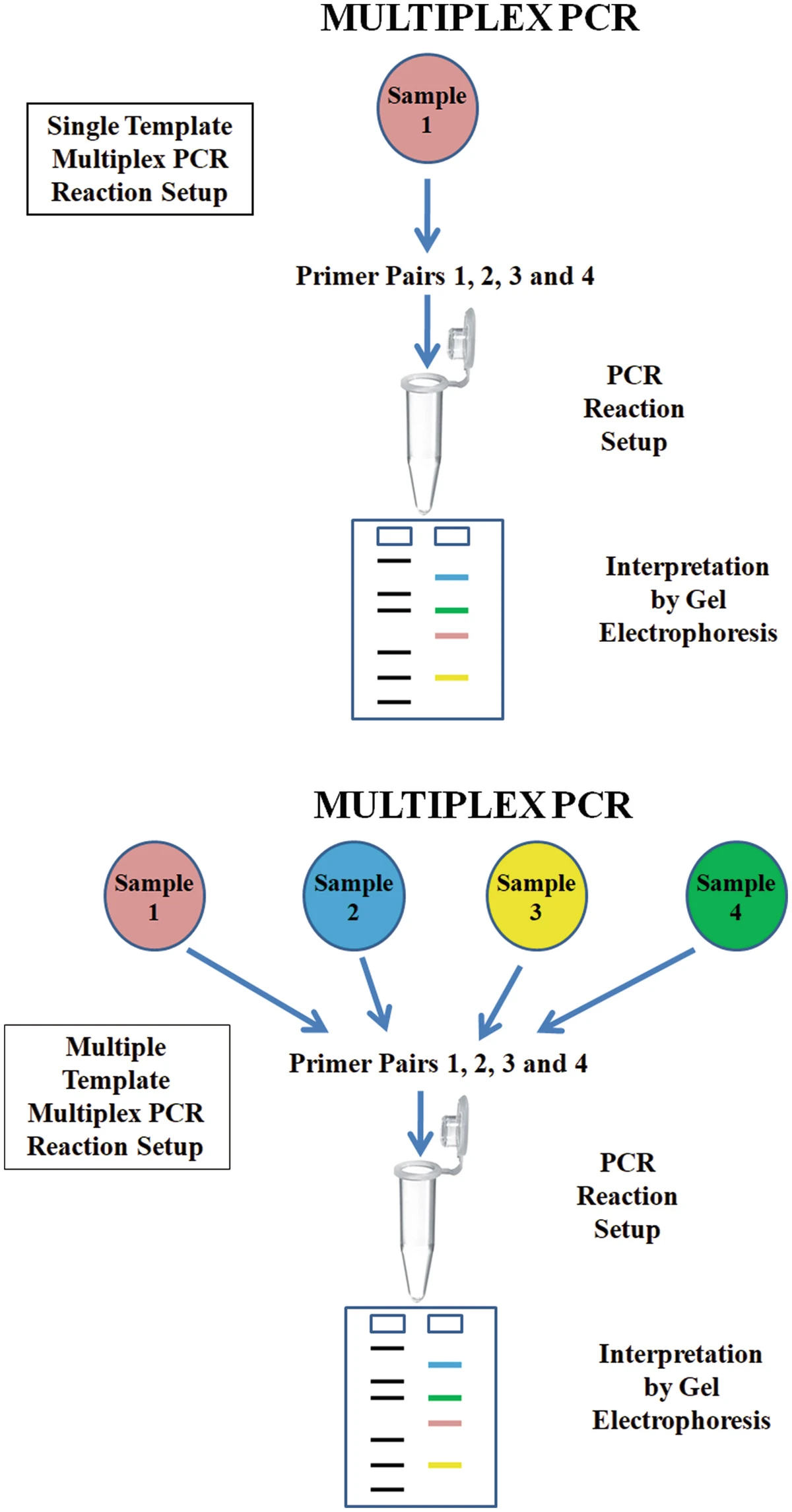
Multiplex PCR for diagnosis of porcine diseases.
Infectious diseases are also diagnosed using metagenomic next-generation sequencing (mNGS). This method works by sequencing the nucleic acids present in a sample, without requiring amplification of specific targets. The diagnosis of Neuroleptospirosis using mNGS cemented the method’s status in modern diagnostic methods in 2014. Using mNGS allows for a broader detection range, using a wider range of specimens, picking up pathogens that can be missed by standard methods. Studies suggest that “ultra-rapid” mNGS could soon be implemented more widely in clinical settings. However, there are of course limitations, namely cost and bioinformatics complexity. There can also be challenges in interpreting results as it is not always possible to distinguish contamination from infection.
Using metagenomic sequencing to evaluate the composition of microbiome in patients with Covid-19.
But which method is best?
Despite the downfalls of many of these methods, most of them still have their place in modern medical settings. Laboratory based methods promise higher sensitivity and accuracy in identifying pathogens, whereas point-of-care testing can produce results in a matter of minutes, although usually at a much higher cost and with implementation challenges. Although home testing is fast and easy, it can be expensive to distribute and there is always a risk of improper use or result misinterpretation. Factors such as location, available resources, patient condition, and public health risk can all influence the diagnostic tool utilised. Even when equipment is available, limited laboratory capacity and shortages of trained scientists can delay adoption of advanced molecular tools.
Having a range of tools to use when diagnosing infectious disease is vital to work towards SDG goal 3.3. There is no one-size-fits-all solution that can address this global challenge. Improving clinical turnaround times and increasing test specificity and sensitivity is important, to allow pathogens to be identified quickly before they can spread further. However, developing portable sequencing tools that can be used in low-resource settings is also vital. Without investment in healthcare access, new diagnostic technologies risk widening existing gaps in global health outcomes.
Innovative new methods are exciting – but implementing them on a wider scale can be a challenge. By making screening methods cost-effective and widely available, health services can gain a more accurate understanding of where outbreaks are starting and take measures to prevent them reoccurring. Many health policy leaders are shifting the focus to community testing, considering price, longevity of testing devices, and test simplicity to avoid misuse. Promoting healthcare access will enable patients to be prioritized for treatment early on in their infection.
Combatting and ultimately ending infectious disease epidemics is a nuanced and long-term challenge, but by investing in diagnostic tools healthcare systems can continue to learn from experiences and develop optimized responses to any new pathogen that emerges.
Scientific Reports is calling for papers on Infectious disease diagnostics as part of a Collection. This Collection welcomes original research into current challenges and advances within the field of infectious disease diagnostics. Submissions are welcome until 23rd June 2026. Collection manuscripts typically see higher citations, downloads, and Altmetric scores and provide a one-stop-shop on a cutting-edge topic of interest.
To submit an article, please visit https://www.nature.com/collections/ddaahffjdf
Follow the Topic
-
Scientific Reports
An open access journal publishing original research from across all areas of the natural sciences, psychology, medicine and engineering.
-
A Collection of original research articles on current challenges and advances within the field of infectious disease diagnostics.
Related Collections
With Collections, you can get published faster and increase your visibility.
Phytochemicals and health
Publishing Model: Open Access
Deadline: Jul 28, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in