Beyond Contamination: How Antibiotic Resistant Bacteria are Spreading in Nigeria's Drinking Water
Published in Microbiology
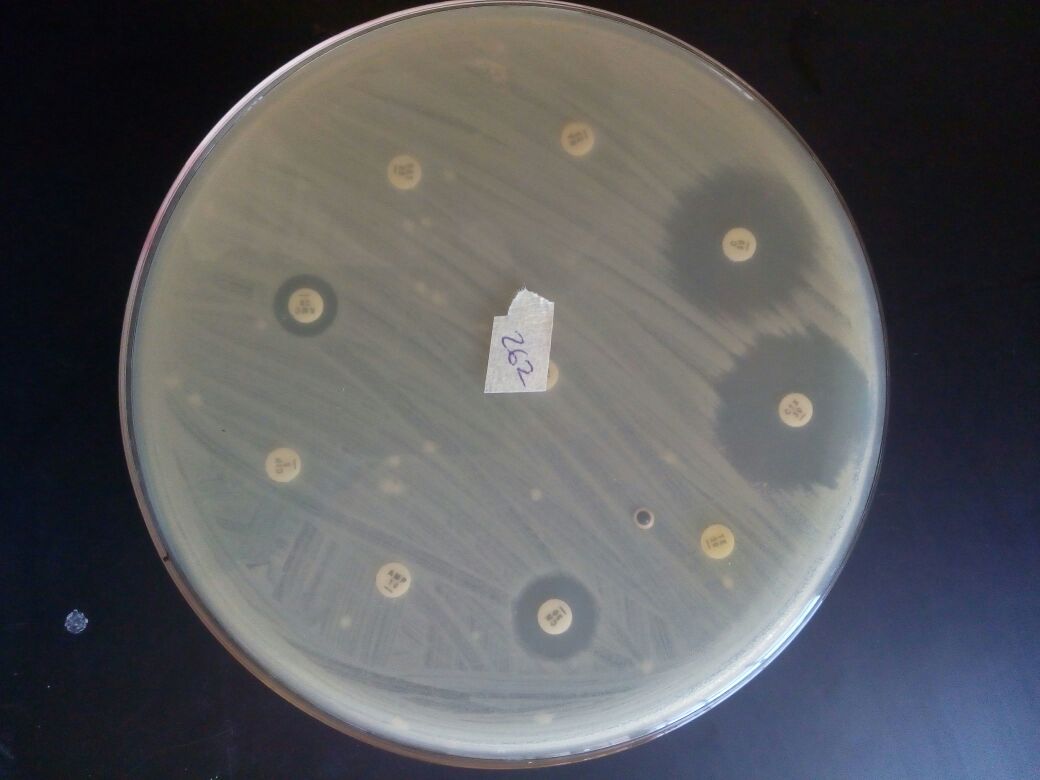
Water scarcity has been a challenge in Nigeria for decades, but beyond the issue of availability, there is a sinister threat that doesn't make headlines: antibiotic-resistant bacteria silently circulating through water. As with many parts of Africa, Kaduna state has its issues with adequate potable water for the population of around 9 million people. Where the municipal pipe-borne water is inadequate or unavailable, individuals depend on water from hand-dug wells, boreholes, streams and commercially available treated water sealed in polythene sachets, popularly known in local parlance as "pure water".
In previous years, the research on antibiotic- resistant bacteria in Nigeria focused majorly on bacteria isolated from clinical settings. This research however, which was carried out between 2014 and 2015, focused on detecting antibiotic resistant Salmonella enterica from various sources of drinking water in Kaduna state. This organism is well known to include serovars that cause diseases such as typhoid fever (which is endemic in Nigeria) and salmonellosis. The symptoms of these diseases can include fever and gastroenteritis, organ failure, shock and death.
The Quest for Answers
As part of her Ph.D research, Dr. Olajoke Alalade under the guidance of her supervisors from Ahmadu Bello University, Zaria collected 500 water samples from diverse sources: packaged water in sachets, municipal treated tap water, wells, boreholes, and streams across six different local government areas across Kaduna State. They were specifically hunting for Salmonella enterica and more importantly, they wanted to know if these bacteria were resistant to the antibiotics we rely on to treat infections. Those that showed resistance were assayed for the presence of some genes responsible for the resistance.
What They Found
Only six samples contained Salmonella enterica, giving an isolation rate of 1.2 %. This may seem low, but it is significant because this bacteria should not be present in water used for drinking. The sources of water where these resistant bacteria were obtained was troubling. Five isolates came from hand-dug wells and boreholes, which are common water sources for many rural and urban families. Most alarming, was finding resistant Salmonella in treated municipal water, the very supply that should be safest after undergoing purification processes. This discovery suggested that either the treatment was inadequate or the ageing distribution pipes had a source of post-treatment contamination.
Four isolates were resistant to multiple antibiotics, showing particular resistance to tetracycline, which is an affordable, widely available antibiotic used across Nigeria. Two-thirds of the isolates were resistant to this drug, along with nalidixic acid and sulfamethoxazole-trimethoprim, which are all front-line antibiotics that are available for purchase over the counter.
The Genetic Evidence
What makes this study particularly significant is that the researchers also found the genetic back up that explained the resistance. Two of the bacterial isolates harbored genes tetA and sul1. These carry genetic information that enable bacteria resist the actions of tetracycline and sulfamethoxazole- trimethoprim. The tetA gene gives bacteria a pump system that pushes tetracycline out of their cells before it can kill them, while the sul1 gene allows bacteria to continue essential functions even when sulfonamide antibiotics try to interfere. What makes this particularly worrying, is that these genes can be horizontally transferred to other bacteria. They can be copied and shared between bacteria, spreading resistance like a viral video on social media.
Why This Matters to You
You might wonder why findings from 2014-2015 in Nigeria matter today. The answer lies in understanding how antibiotic resistance works and spreads. When bacteria in water sources carry resistance genes, they can end up in the food chain through various pathways. For instance, when contaminated water is used for irrigation and animal husbandry, they ultimately end up in our bodies when we eat such food. This creates a vicious cycle where resistance spreads across environmental, animal, and human health domains. There are other implications such as higher treatment costs for ill individuals, longer hospital stays, increased chance of mortality, and greater economic burden on families.
The One Health Connection
This research exemplifies the "One Health" approach, which recognizes that human, animal, and environmental health are connected inextricably. The same antibiotics used carelessly in human medicine, animal farming, and even agriculture create selection pressure that allows resistant bacteria to thrive everywhere.
In Kaduna State, some communities' water sources contained bacteria susceptible to all antibiotics, while others harbored multi-drug resistant strains. These differences likely reflect local patterns of antibiotic use, sanitation infrastructure quality, and environmental contamination levels.
Practical Solutions
The researchers propose several actionable solutions that communities can implement. They include boiling drinking water since Salmonella is heat-sensitive, protection of water sources such as the proper construction of wells far away from latrines, and improved sanitation. Open defecation must be discouraged as it is a major way such bacteria get into water bodies and antibiotic stewardship must also be taken seriously. The populace must be educated on the use of antibiotics only on professional advice, and completing the course of treatment as these help reduce spread of resistance among bacteria. Water sources should also be tested periodically for contamination and resistant bacteria.
Looking Forward
While this data is now a decade old, it provides crucial baseline information. The presence of resistance genes in 2014-2015 means these genes were already circulating in environmental reservoirs long before many current interventions began. The researchers call for urgent follow-up surveillance to assess whether resistance patterns have intensified and whether any interventions implemented over the past decade have made a difference. This historical data becomes invaluable for tracking trends and evaluating the effectiveness of public health measures.
The Bigger Picture
This study from Kaduna State is a microcosm of a global crisis. The World Health Organization projects nearly 2 million deaths annually from bacterial antibiotic resistance by 2050. Water sources serve as both reservoirs and highways for resistance genes, making environmental monitoring essential to any comprehensive response strategy. Addressing antibiotic resistance requires coordinated action across healthcare, agriculture, environmental management, and public education. The genes detected in these water samples are warnings we cannot afford to ignore.
Follow the Topic
-
BMC Microbiology
This is an open access, peer-reviewed journal that considers articles on all microorganisms - bacteria, archaea, algae and fungi, viruses, unicellular parasites and helminths.
Related Collections
With Collections, you can get published faster and increase your visibility.
Mycobacteria: advances in physiology and mycobacterial diseases research
Mycobacteria are a large and diverse group of microorganisms, known for their adaptability and resistance to environmental stresses, particularly due to their unique thick, hydrophobic cell wall rich in mycolic acids. While most mycobacteria are not pathogenic, some of them (e.g. Mycobacterium tuberculosis or Mycobacterium leprae) have evolved to become very successful human pathogens, with complex pathogen-host interactions that are still poorly understood. Often because of their unique physiological properties that facilitate survival in different environments, persistence in a latent state within the host and evasion of host immune responses, mycobacteria pose significant challenges to global public health. A greater understanding of the physiology and mechanisms underlying mycobacterial infections is essential to develop effective treatments, diagnostics and preventive measures, especially considering the rise of drug-resistant mycobacterial strains, and the global burden of tuberculosis- and non-tuberculosis-related diseases. Recent advances in understanding mycobacterial basic biology and physiology, pathogenesis, and immune evasion are paving the way for novel therapeutic strategies, including the development of new vaccines, as well as new molecular diagnostics.
In support of United Nations’ SDG 3 (Good health and well-being), BMC Microbiology presents the Collection Mycobacteria: advances in physiology and mycobacterial diseases research. This Collection invites research on mycobacteria, focusing on advances in understanding their physiology, pathogenesis, and the diseases they cause. We welcome submissions that explore host-pathogen interactions, immune system responses, drug-resistance mechanisms, and innovative approaches for the prevention, diagnosis and treatment of mycobacterial infections. Research without a clear focus on microorganisms or pathogen-host interactions will not be considered. We will consider research articles that investigate, but are not limited to, the following topics:
- Physiology, metabolism, stress response mechanisms, adaptation to environmental stresses, sources of transmission and environmental persistence of mycobacteria
- Mycobacterial pathogenesis and immune evasion
- Innate and adaptive cellular immune responses to M. tuberculosis and non-tuberculous mycobacteria (NTM) infections
- Drug-resistance mechanisms, persistence and dormancy in mycobacteria
- Mycobacterial reactivation and transmission
- Host-pathogen interactions in mycobacterial diseases
- Advances for combating mycobacterial co-infections
- Conserved and specialized functions of Type VII secretion systems (T7SS) (ESX or ESAT-6 systems) of mycobacteria
- Advances in molecular diagnostics and biomarkers for the detection of mycobacterial infections, disease progression and cure
- Vaccine development for mycobacterial infections
- Novel drug targets and therapeutic strategies against mycobacterial infections
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Jul 17, 2026
Microbes in built environments
The microbial communities (e.g. bacteria, fungi, viruses) that inhabit human-built environments are diverse, dynamic and complex, and they play a significant role in human health and wellbeing. From indoor air quality to the surfaces and building materials we touch daily, we interact with a range of microbes that come from a variety of sources such as other humans, pets, plants, pests, and other outdoor sources. Investigating and understanding these interactions require interdisciplinary approaches that integrate microbiology, ecology, and engineering principles, among other disciplines. As our understanding of microbial dynamics within built environments advances, it becomes increasingly clear that built-environment microbes and microbiomes can strongly affect human health, either positively or negatively.
Recent research has highlighted the potential importance of indoor microbial diversity in influencing human health (e.g. respiratory health, allergies, and the spread of multidrug-resistant organisms, particularly in healthcare settings). Innovations in monitoring and modelling microbial communities are opening new avenues for designing healthier indoor environments and mitigating risks from unwanted microbial exposures. Ongoing research in the field has the potential to yield novel insights into how these microbes respond metabolically to certain stressors and how we can leverage that knowledge. Additionally, a better understanding of microbe-host interactions could aid in developing novel approaches in building design and public health initiatives. Adaptive antimicrobial methods based on real-world conditions (such as occupancy, ventilation, and microbial indicators) offer a promising pathway to control microbial risks while minimizing unintended impacts.
In support of the United Nations’ Sustainable Development Goal 3 (SDG 3, Good Health and Well-Being), BMC Microbiology launches the Collection Microbes in built environments. This Collection invites research articles that contribute to the understanding of microbes and microbiomes in human-built environments, encompassing microbial ecology, diversity, metabolism, and their implications for human health. Submissions are welcome on topics ranging from indoor microbial communities to the impact of design on microbe-host interactions in the built environment. Research without a clear focus on microorganisms, microbial communities, or microbiomes in built environments will not be considered. We invite researchers and experts in the field to submit research articles covering a broad range of topics, including, but not limited to:
- Interactions between microorganisms/viruses and hosts in built environments
- Microbial physiology, metabolism, adaptation and diversity of the built environment
- Impact of microbes in the built environment on human health and diseases
- Antimicrobial resistance of the built environment
- Holistic pathogen defence in the built environment
- Microbial ecology of the built environment
- Microbial adaptation and evolution in response to anthropogenic disturbances in built environments
- Viruses in the built environment
- Food surfaces and disinfection in the built environment
- Role of indoor air quality, ventilation and HVAC systems in microbial community dynamics
- Impact of surface and building materials on microbial colonisation
- Bioremediation and microbial management in built environments
- Interventions to promote/discourage microbial exposures in built environments
- Environmental monitoring methods of microbes in built environments
- Adaptive antimicrobial strategies in built environments
All manuscripts submitted to this journal, including those submitted to collections and special issues, are assessed in line with our editorial policies and the journal’s peer-review process. Reviewers and editors are required to declare competing interests and can be excluded from the peer review process if a competing interest exists.
Publishing Model: Open Access
Deadline: Oct 22, 2026


Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in