Building Iron-Sulfur clusters: from ancestral simple systems to current diversity
Published in Ecology & Evolution
In living cells, many enzymes need cofactors to work properly. There are thousands of types of cofactors, that can be either organic or non-organic. Among them, one is particularly interesting because of its simplicity and ubiquity among all living organisms, the Iron-Sulfur cluster. As its name indicates, it is basically composed of Iron (Fe) and Sulfur (S) that can be organized in different ways, typically in a square shape [2Fe-2S] or cubic shape [4Fe-4S] (Figure 1). Fe-S clusters have a particularity that is very useful in biology: they can trap and release electrons. Thus, they can act as real electronic capacitors or even wires when they form a chain inside proteins. They participate to different oxidoreduction reactions in many important biological processes, such as respiration, photosynthesis, metabolic pathways, stress response, …
But wait, how are Fe-S clusters built? Well, two options are possible. First, the clusters can be spontaneously produced, only by chemistry. This is what happens when you simply mix Iron and Sulfur in a tube. The second option is biochemical synthesis. Indeed, some proteins can build Fe-S clusters in a controlled way (Figure 1). First, the protein needs to grab sulfur, iron, and electrons. Then a scaffold protein builds the cluster and eventually transmits it transiently to a carrier protein. Finally, the cluster is delivered to the target protein. Sadly, for our Fe-S clusters, they are sensitive to chemical stresses, and particularly to oxygen and derived species. In this context, the use of biosynthesis machineries helps to protect the clusters by hiding them in the buried regions of the proteins.
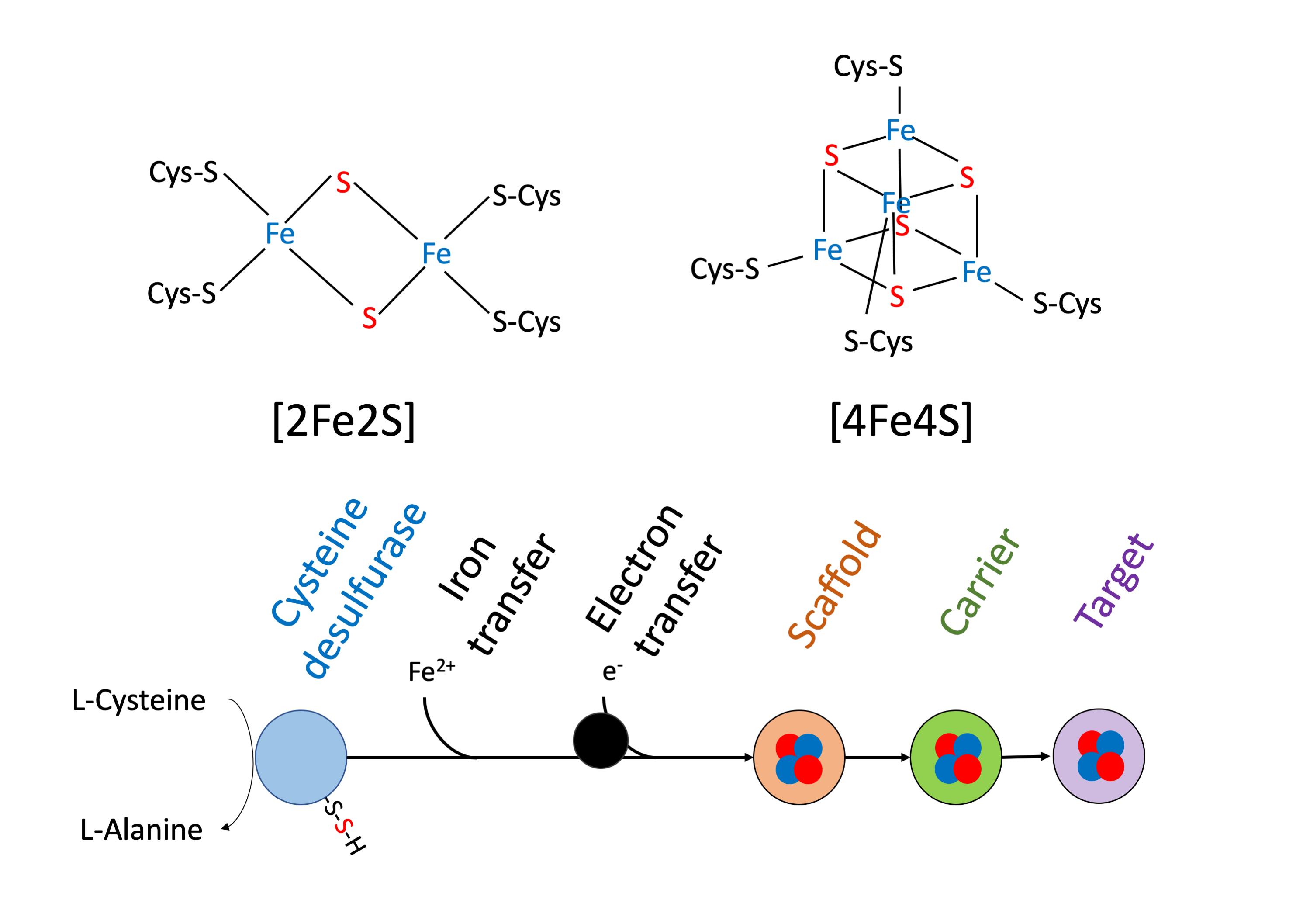
Structure of Iron-Sulfur clusters and general pathway for their biosynthesis.
Three different machineries that carry out Fe-S cluster synthesis have been described, the ISC, SUF and NIF machineries. ISC and SUF are the major systems that have been widely described in the model bacterium Escherichia coli. ISC is the most important one, and syntheses Fe-S clusters in normal conditions, while SUF works only as backup, especially under oxidative stress. The different systems share common components (i.e. cysteine desulfurases) but also have numerous specific components.
This diversity in term of Fe-S cluster biogenesis systems raises the question of the origin and evolutionary history that shaped them. A widespread idea is that the first forms of life used the Fe-S clusters that were spontaneously synthesized in the ancient anaerobic environment. Then, the concentration of oxygen increased dramatically circa 2.4 By ago, because of the emergence of Cyanobacteria and oxygenic photosynthesis. The resulting increasing oxidative stress is largely supposed to have been the selective pressure responsible for the emergence of Fe-S cluster biosynthesis machineries to protect the cluster. This is an attractive hypothesis, but until now, no data had clearly showed it. The rare studies that attempted to investigate the evolution of Fe-S machineries were quite old, used a limited taxonomic sampling, and did not consider all the components and all the systems at once. In addition, the evolutionary relationship between the different Fe-S machineries has never been explored. This is what motivated us to start this project. However, things are never as simple as they seem at the start!
First, I needed a database of proteomes that would be representative of the entire diversity of prokaryotes, in contrast to the previous studies. I therefore put a lot of efforts into finding the best way to build such database. On the NCBI, one could find around 200,000 prokaryotic genomes, all very heterogeneous in term of quality, with missing files, and with an over representativeness of some clades/species (i.e. E. coli). I developed different pipelines to progressively reduce the number of genomes, keeping the best quality ones and reducing the disparity of clade representativeness. The next step was to obtain reliable datasets of homologues of all components of the different Fe-S cluster biosynthesis machineries (SUF, ISC, and NIF). This step has been particularly time consuming, because several of these components belong to huge multigenic protein families, including many paralogues. In such case, analysis of the genomic organization of the different components helped me to understand “who is who” in this mess.
It was clearly worth the effort, as once I had all my clean datasets of homologues, I got the first striking result. Together with SUF, ISC and NIF, two additional minimal systems clearly stood out. They have in fact only two components and are clearly distinct from the other three machineries because they form their own clades in phylogenetic analyses. These systems were loosely described in the past literature but misidentified as SUF or NIF in the absence of a thorough bioinformatics analysis. Names are important in the description of biological diversity and future characterizations, and so to underline the distinctiveness of these two minimal systems, we proposed to rename them SMS (SUF minimal system) and MIS (Minimal Iron-Sulfur). But the mere presence of homologues does not tell you if they are really involved in Fe-S cluster biosynthesis. Here, our experimentalist collaborators jumped in and were able to show for the first time that SMS is a bona-fide Fe-S system, nicely confirming my bioinformatics predictions. These results already show that the diversity of Fe-S cluster biogenesis is higher than previously thought. But another surprise was waiting around the corner…
It came from the taxonomic distribution of the five systems in thousands of genomes from Bacteria and Archaea. Indeed, I found that SMS and MIS are not exotic exceptions as previously described but are instead widespread among prokaryotes. The SMS system is largely found in Archaea while the MIS system is widespread in both Archaea and Bacteria. In comparison, the well-studied ISC system is actually present in only a few bacterial clades. Furthermore, I found that most bacteria possess only one system, indicating that they are not redundant machineries. In contrast, the presence in E. coli of both SUF and ISC stands more as an exception than the rule, even among Gammaproteobacteria. So, our vision of Fe-S biogenesis systems in terms of importance in life was largely biased by what we know in E. coli!
The next step was to understand how the different Fe-S systems originated and evolved during the diversification of Bacteria and Archaea. By comparing the phylogenetic trees of all the components of the different systems with the reference bacterial and archaeal phylogenies, I was able to precisely dissect their evolutionary trajectories. I inferred that the Last Universal Common Ancestor (LUCA) already possessed SMS and MIS. This means that the earliest organisms were already synthesizing Fe-S clusters biochemically with dedicated machineries way before the apparition of oxygen in the atmosphere. This is a paradigm changing result as it implies that the arrival of oxygen was not the driving force that led to the emergence of Fe-S machineries. Also surprisingly, it appears that Cyanobacteria replaced their original SUF machinery by one obtained from Proteobacteria and likely more adapted to cope with oxidative stress, which may have helped the very emergence of oxygenic photosynthesis.
Wrapping up all these results, and after multiple brainstorming sessions (and heated discussions!) together with my two supervisors, an evolutionary biologist (Simonetta) and a specialist of Fe-S (Frédéric), we were able to reconstruct a global scenario for the evolution of Fe-S machineries after LUCA (Figure 2). While Archaea largely kept both SMS and MIS, Bacteria developed more complex Fe-S machineries by adding and combining new components. More precisely, the SMS system was complexified giving rise to the SUF system. The MIS system was also modified twice to give rise to the ISC and the NIF systems.
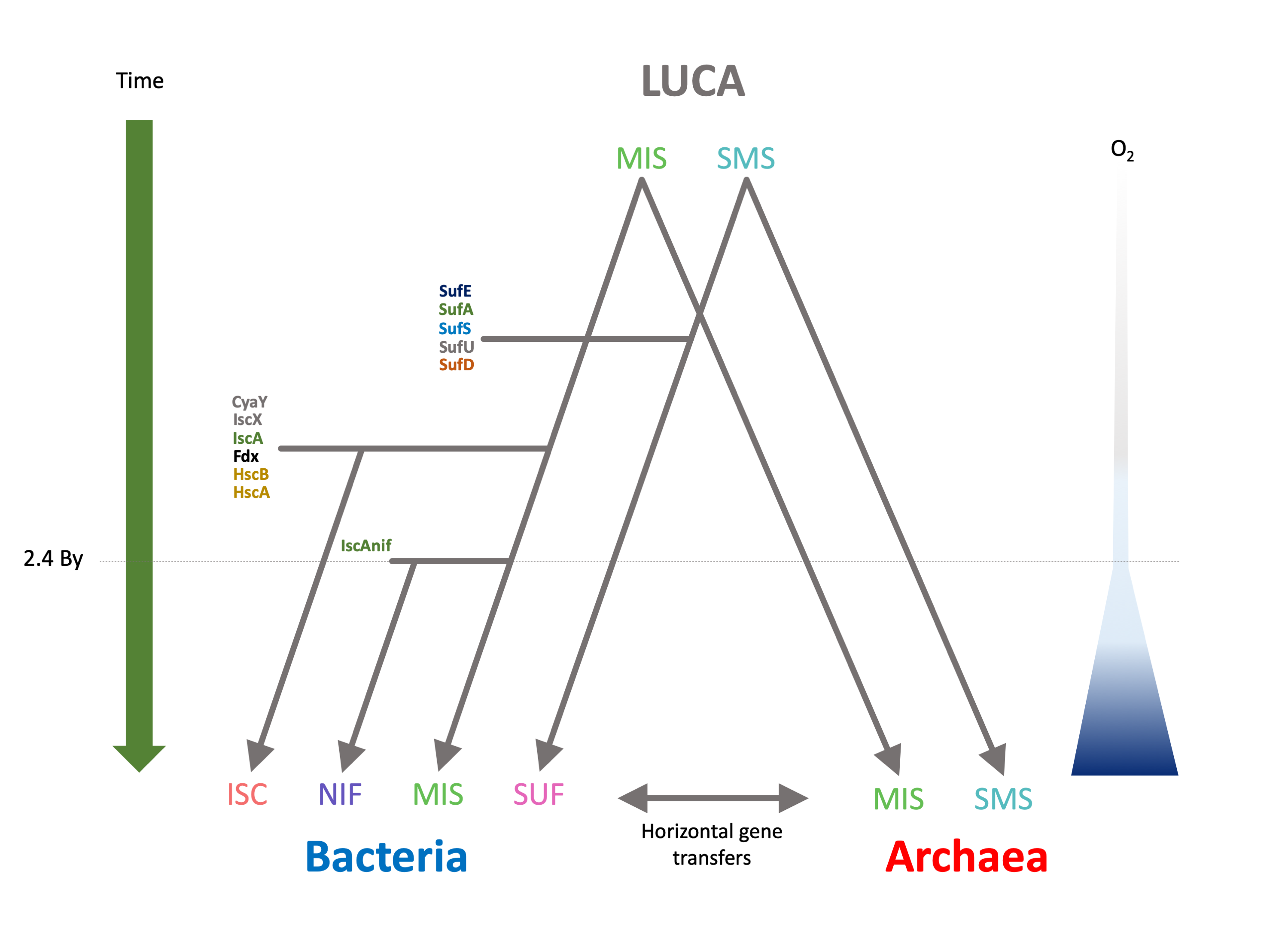
The selective pressure driving these evolutionary events remains to be determined and worth some speculation. As a matter of fact, it is clear that, if oxygen did not play a role in the very emergence of the ancestral Fe-S systems, it certainly did in their later evolution. I observed in fact that several archaea replaced their original SMS machineries by SUF systems acquired from bacteria, and this correlates with multiple adaptation to aerobic environments. Conversely, I observed that some bacteria replaced their original SUF systems by SMS acquired from archaea, potentially allowing them to adapt to stricter anaerobic niches. These results suggest that bacterial SUF may be more resilient to the presence of oxygen than archaeal SMS.
In conclusion, reconstructing the ancient evolution of biochemical processes is not a trivial task, but it can lead to challenging assumptions often largely anchored in the scientific community. Certainly, it was made possible by exploiting the current vast diversity of genomes that are now increasingly available from a large taxonomic spectrum. This work sets an important step in our understanding of the diversity and evolution of one of the most ancient and important biological processes, and what are the ecological forces that led to such diversity. It also provides an exhaustive and robust annotation of all components of Fe-S machineries across the whole diversity of Archaea and Bacteria, which will be of great use to the scientific community, and pave the way for future characterization. It will be especially important to obtain a better understanding of how the minimal SMS and MIS systems function, and notably in the Archaea, were Fe-S biology is still largely a mystery. More to come in the next few years!
Follow the Topic
-
Nature Ecology & Evolution

This journal is interested in the full spectrum of ecological and evolutionary biology, encompassing approaches at the molecular, organismal, population, community and ecosystem levels, as well as relevant parts of the social sciences.
Related Collections
With Collections, you can get published faster and increase your visibility.
Biodiversity and ecosystem functioning of global peatlands
Publishing Model: Hybrid
Deadline: Jul 27, 2026
Understanding species redistributions under global climate change
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in