Natural ‘killer’ cells (NK) are the first responders in the human body’s arsenal of immune cells. As their name suggests, to ‘kill’ or to simply eliminate enemies, they first need to recognize and connect with them. To do so, they use a range of activating and inhibitory receptors. DNAM-1, an activating receptor, is already well-known for its role in cancer immunity and autoimmune diseases. Interestingly, no information is currently available regarding its role in fungal infections. We started with a simple hypothesis: DNAM-1 may also play a role in fungal immunity. And that is how the story began!
The first hints
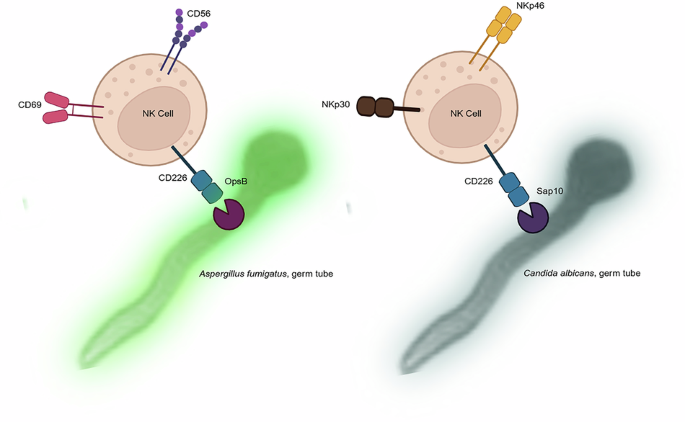
To learn more about the receptor itself, we investigated how DNAM-1 is distributed on primary human NK cells. Our weapon was super-resolution fluorescence microscopy, which revealed a uniform distribution of DNAM-1 on isolated primary NK cells. Using CRISPR-Cas9 technology, we knocked out DNAM-1 in primary NK cells and visualized NCAM-1, another activating receptor on NK cells, for comparative analysis. This encouraged us to examine how DNAM-1 is distributed or affected when NK cells are co-cultured with fungal cells. We chose Aspergillus fumigatus and Candida albicans, two of the most medically important fungi that cause invasive fungal infections. NCAM-1 is a well-known pathogen recognition receptor (PRR) for Aspergillus fumigatus. We visualized DNAM-1 and NCAM-1 on NK cells in co-culture with Aspergillus fumigatus. While NCAM-1 was completely relocalized after 6 hours of co-incubation, DNAM-1 was not. The accumulation of NCAM-1 was also observed in DNAM-1-deleted NK cells. These microscopic observations were further supported by flow cytometry measurements, which showed a slight reduction in mean fluorescence intensity (MFI). This is important because reduced MFI is an indirect measure of immunological synapse (IS) formation. Additionally, when NK cells were challenged with C. albicans hyphae, DNAM-1 distribution was similar to that observed with A. fumigatus.
The surprise
Following the interaction of A. fumigatus and C. albicans with DNAM-1, we investigated whether this interaction was direct or indirect. To test this, we coincubated soluble DNAM-1 protein with the hyphae of both fungi and visualized the interaction by microscopy as well as flow cytometry. In both cases, direct binding of DNAM-1 to the fungal cell wall was observed. The immunobiological consequences of these interactions were also assessed by exposing C. albicans hyphae to NK cells in the presence or absence of a DNAM-1 D1 domain-blocking antibody. The presence of the blocking antibody significantly reduced the percentage of damaged hyphae, indicating that DNAM-1 contributes to NK-cell cytotoxicity in response to fungal hyphae. As DNAM-1 is recognized as a co-stimulatory molecule, we then assessed the roles of DNAM-1, CD96, and TIGIT in antifungal activity using a viability assay. NK cells treated with blocking antibodies against these receptors showed effects on the viability of C. albicans after co-incubation. While DNAM-1 blockade partially reversed the reduction in viability, TIGIT blockade and the corresponding isotype controls did not consistently affect viability. Of note, CD96 blockade also increased viability, similar to DNAM-1 blockade.
From in silico prediction to biological evidence
After confirming the interaction of DNAM-1 with the fungal surface, we aimed to identify potential fungal interaction partners for DNAM-1 computationally. We searched the existing proteomes of both A. fumigatus and C. albicans using a domain-pair-based approach, taking into account protein localization and their potential involvement in pathogenicity-related processes. Although several proteins scored highly, we chose Sap10, a cell wall protease of C. albicans, as the focus of our study because of its accessibility and previous experimental data. High-affinity binding between DNAM-1 and Sap10 was also predicted by AlphaFold 3 and protein-protein docking. A protein from A. fumigatus, OpsB, was also predicted as a homologue of Sap10. However, this protein has not yet been analyzed in detail in A. fumigatus.
Following the domain–domain interaction forecast, we experimentally demonstrated a potential interaction between DNAM-1 and Sap10. For this purpose, a truncated excreted version of Sap10 was used. A modified immunoprecipitation assay was employed to analyze the interaction between soluble DNAM-1 and fluorescently labeled Sap10, followed by flow cytometric analysis. This experiment clearly indicated the high affinity of Sap10 for DNAM-1. We also analyzed the dose dependency of DNAM-1 binding to fluorescently labeled Sap10 by fluorescence correlation spectroscopy. This further indicated that Sap10 is an interaction partner of DNAM-1 and binds with high affinity. We then challenged NK cells with purified Sap10 protein and visualized DNAM-1. While the distribution of Sap10 was similar to that of DNAM-1 in primary NK cells, the fluorescence signal was absent in DDNAM-1 NK cells, again showing that DNAM-1 specifically promotes the binding of Sap10 to the cell surface.
Sap10 triggers NK-cell activation
Next, we investigated whether Sap10 can modulate NK cell activation. CD69 surface expression (MFI) was measured by flow cytometry as an indicator of NK-cell activation, with or without blocking antibody against DNAM-1. The highest level of activation was achieved at higher Sap10 concentrations and longer incubation times. To further confirm Sap10-induced NK-cell activation via DNAM-1, secretion of the cytotoxic effector molecule perforin and the chemokine CCL3 was measured using ELISA. Indeed, Sap10 exposure triggered robust secretion of both perforin and CCL3 in a dose-dependent manner after 24 hours of incubation. The relative amounts of both perforin and CCL3 released by Sap10-treated NK cells were significantly reduced when the blocking antibody was used. These findings clearly indicate that NK-cell activation occurs via Sap10 binding to DNAM-1.
Why does it matter?
Altogether, our data highlight DNAM-1 as a novel fungal recognition receptor and suggest that targeting the DNAM-1–Sap10 axis, alone or in combination with modulation of TIGIT/CD96 or NKG2D pathways, holds promise for antifungal immunotherapy. These concepts warrant evaluation in C. albicans and A. fumigatus preclinical models to assess both efficacy and safety.
Final thoughts
Our work has its limitations, and many questions remain open. But we should not forget this: even the moon has its scars, so why expect perfection in anything, even science? Like many stories in science, this one was built not only on experiments and analysis, but also on countless cups of coffee and endless discussions that kept the ideas moving.
Follow the Topic
-
Communications Biology
An open access journal from Nature Portfolio publishing high-quality research, reviews and commentary in all areas of the biological sciences, representing significant advances and bringing new biological insight to a specialized area of research.
Ask the Editor - Immunology, Pathogenesis, Inflammation and Innate Immunity
Got a question for the editor about the complement system in health and disease? Ask it here!
Continue reading announcementRelated Collections
With Collections, you can get published faster and increase your visibility.
Mechanistic insights into human host and microbiome interactions
Publishing Model: Open Access
Deadline: May 31, 2026
Advances in neurodegenerative diseases
Publishing Model: Hybrid
Deadline: Jun 30, 2026



Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in