Exploring native RNA diversity with nanopore microscopy and RNA nanotechnology
Published in Chemistry
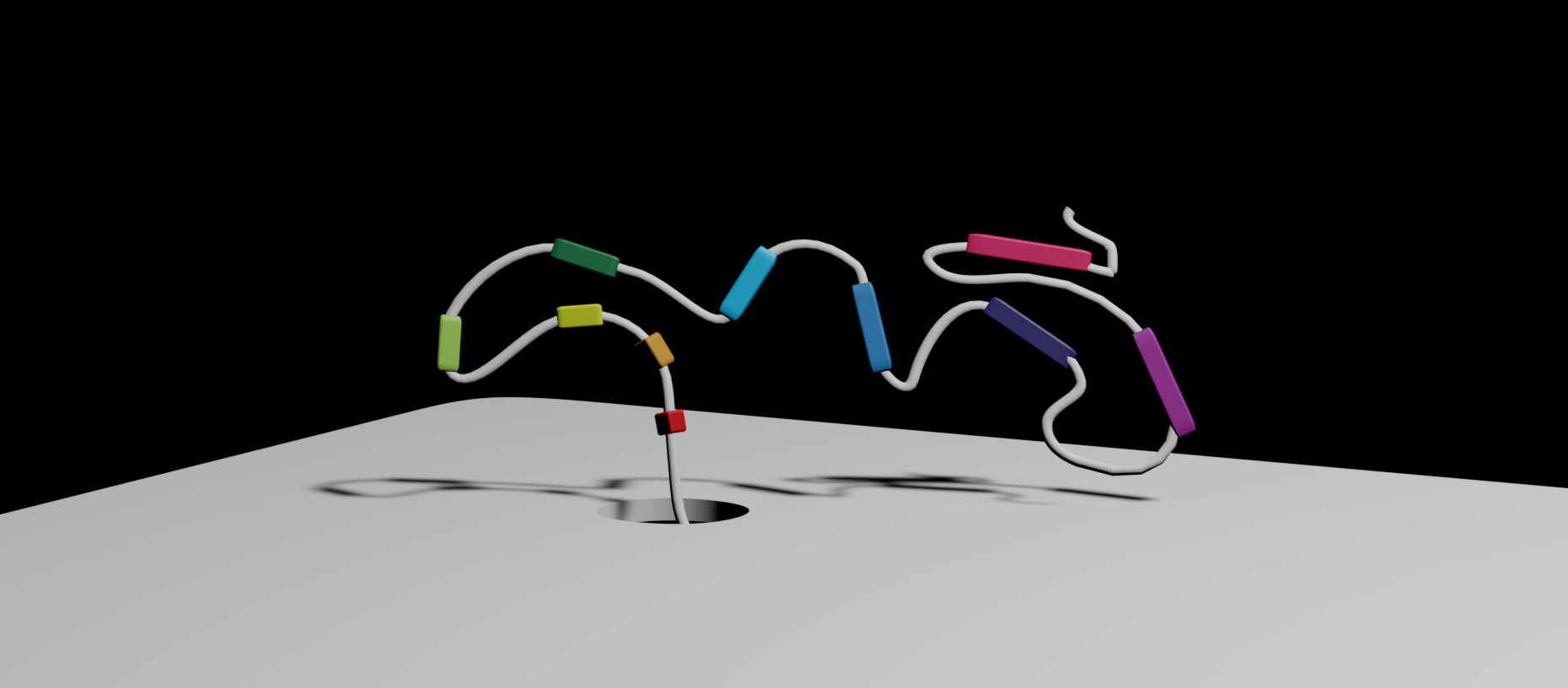
RNA is a key player in the transfer of information and regulation of biological processes underlined by the immense potential for RNA-based therapeutics and functional RNA nanostructures. Identifying RNA requires intricate protocols that suffer from various enzymatic biases and lead to irreversible loss of native information about RNA presence and quantity. Up until now, we could not detect native RNA ‘makeup’, including RNA structures spanning kilobases. The mere order of bases in RNA could not tell us how that RNA looked.
We have recently developed a new method - Amplification-free RNA TargEt Multiplex Isoform Sensing (ARTEMIS) that has made possible the identification of multiple RNA isoforms by their length, order of elements, and circularity in parallel.
Here we design three-dimensional molecular constructs that enable identification of native RNA at the single-molecule level using solid-state nanopore microscopy [1]. Target RNA is refolded into RNA origami codes with designed sets of complementary DNA strands. Each RNA code has a set of bits that correspond to the size of uncomplemented RNA structures. Nanopore microscope identification works via voltage-driven translocation of negatively charged RNA codes through a small orifice towards a positively charged electrode in an electrolyte solution [2-3]. It translates RNA code into a current signal with a spatial resolution comparable to complex optical microscopies with higher throughput and straightforward origami assembly. ARTEMIS discriminates between RNA isoforms, RNA circularity, and RNA structures. We employed RNA codes and identified multiple RNA variants of messenger RNA and long non-coding RNA in human cervical adenocarcinoma. Our approach has the potential to discriminate up to 10 billion unique RNAs for multiplexed pathogen detection, vaccine byproducts, cancer diagnosis, and as an ultrarapid native RNA characterization technique.
Besides multiplexed identification, we envision various applications for studying RNA intramolecular and intermolecular interactions with proteins and small molecules. One application of great interest that has emerged is ribosomal RNA-based pathogen identification. Ribosomal RNA (rRNA) gene (DNA) has been extensively used for bacterial strain identification. However, by using rRNA itself, a much more abundant molecule, instead of the gene which encodes it, enables a 104-106 times fold increase in sensitivity. Future directions would include native transcriptomics at the single-cell and single molecule levels. The single-cell native transcriptomics with ARTEMIS is the clear next step, where one can combine droplet-based handling of cells with ARTEMIS. We can achieve absolute quantification of the native transcriptome with ARTEMIS, allowing the potential discovery of novel biomarkers.
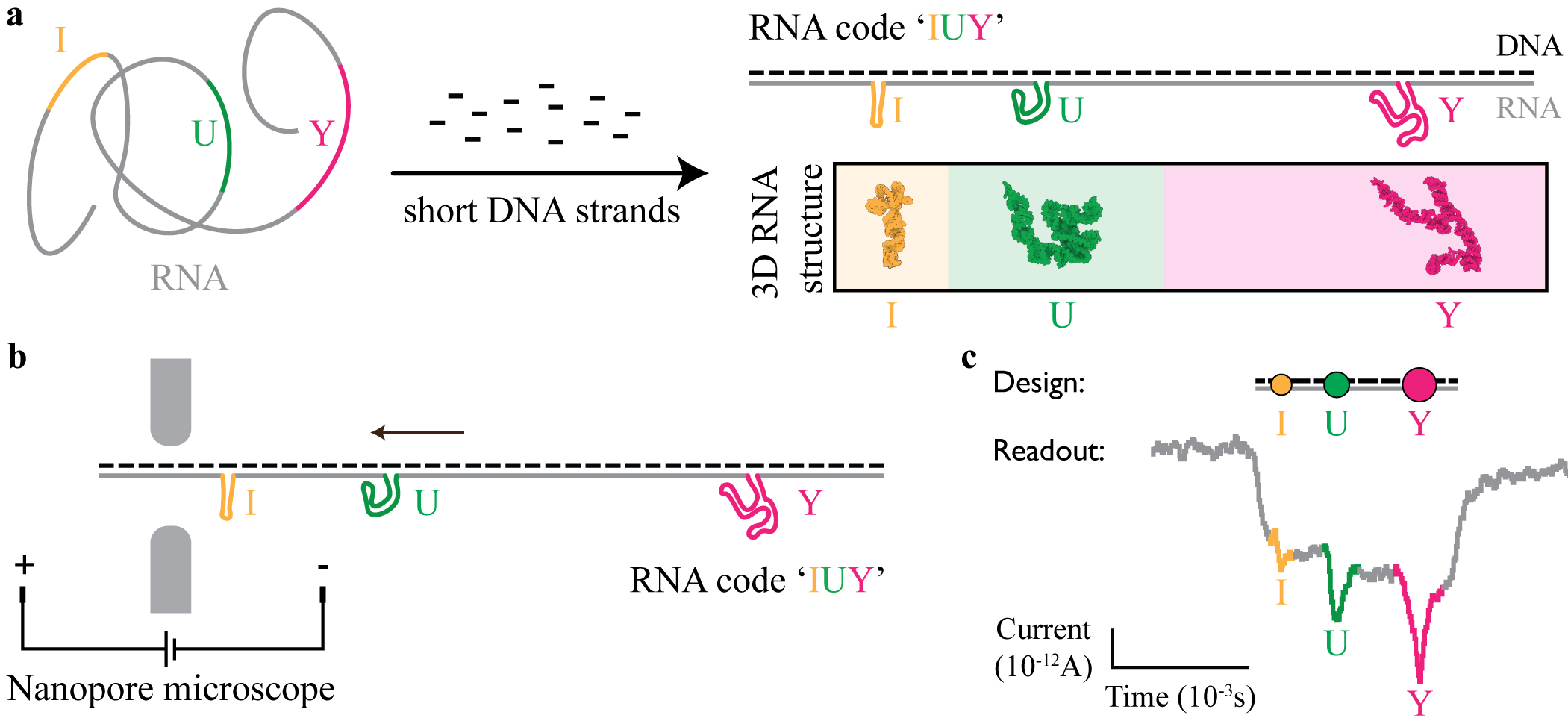
Fig. 1 Self-assembled RNA origami-based codes readout with solid-state nanopore microscopy. a) RNA is complemented with short DNA strands. Colored parts are left unpaired to self-assemble in RNA bits (‘I’, ‘U’, and ‘Y’) that together create a unique RNA code (‘IUY’). The Three-dimensional (3D) RNA structure of each bit is designed to resemble the shape of a letter. b) RNA code ‘IUY’ is voltage-driven through the nanopore microscope. c) Designed RNA codes read with nanopore microscope results in the ionic current signal where downward spikes correspond to designed RNA bits.
[1] Bošković, F., Keyser, U.F. Nanopore microscope identifies RNA isoforms with structural colours. Nature Chemistry (2022). https://doi.org/10.1038/s41557-022-01037-5
[2] Wanunu, M., Dadosh, T., Ray, V., Jin, J., McReynolds, L., Drndić, M. Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nature Nanotechnology 5, 807–814 (2010). https://doi.org/10.1038/nnano.2010.202
[3] Bošković F. and Keyser U.F., Science, 374, 6574, 1443-1444 (2021). https://doi.org/10.1126/science.abn0001
Follow the Topic
-
Nature Chemistry
A monthly journal dedicated to publishing high-quality papers that describe the most significant and cutting-edge research in all areas of chemistry, reflecting the traditional core subjects of analytical, inorganic, organic and physical chemistry.






Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in